Abstract
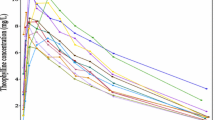
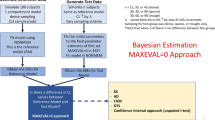
The purpose of this study was to address the question of whether the use of nonlinear mixed-effect models has an impact on the detection and characterization of nonlinear processes (pharmacokinetic and pharmacodynamic) in rich data obtained from a few subjects. Simulations were used to assess the difference between applying population analysis, ie, nonlinear mixed-effects models as implemented in NONMEM, and the standard 2-stage (STS) method as the data analysis method for detection and characterization of nonlinearities. Three situations were considered, 2 pharmacokinetic and 1 pharmacodynamic. Both the first-order (FO) and FO conditional estimation (FOCE) algorithms were used for the population analyses. Within each situation, rich data were simulated for 8 subjects at multiple dose levels. The true nonlinear model and a simpler linear model were fit to each data set using each of the STS, FO, and FOCE methods. Criteria were prespecified to determine when each data analysis method detected the true nonlinear model. For all 3 simulated situations, the application of population analysis with the FOCE algorithm enabled the detection and characterization of the true nonlinear models in at least a 4-fold lower dose level than the STS approach. For both of the pharmacokinetic settings, population analysis with the FO algorithm performed much more poorly than the STS approach. The superior detection and characterization of nonlinearities provided by population analysis with the FOCE algorithm should allow drug developers to better predict and define how a drug should be used in clinical practice in such situations.
Similar content being viewed by others
References
Ludden TM. Nonlinear pharmacokinetics. Clinical Implications. Clin Pharmacokinet. 1991;20:429–446
Dutta S, Matsumoto Y, Ebling WF. Is it possible to estimate the parameters of the sigmoid Emax model with truncated data typical of clinical studies? J Pharm Sci. 1996;85232–239.
Spigset O. The selective serotonin reuptake inhibitor fluvoxamine. Ume niversity medical dissertations, 1997.
Blackledge GRP. High-dose bicalutamide monotherapy for the treatment of prostrate cancer. Urology 1996;47(suppl 1A):44–47
Sedman AJ, Wagner JG. Importance of the use of the appropriate pharmacokinetic model to analyse in vivo enzyme constants J Pharmacokin Biopharm. 1974;2:161–173
Sheiner LB, Rosenberg B, Marathe VV. Estimation of population characteristics of pharmacokinetic parameters from routine clinical data J Pharmacokin Biopharm. 1977;5:445–479.
Sheiner LB, Beal SL. Evaluation of methods for estimating population pharmacokinetic parameters. ii. Biexponential model and experimental pharmacokinetic data. J Pharmacokin. Biopharm. 1981;9:635–651.
Graves DA, Chang I. Application of NONMEM to routine bioavailability data. J Pharmacokin Biopharm. 1990;18:145–160.
Aarons O, Mandema JW, Danhof M A population analysis of the pharmacokinetics and pharmacodynamics of midazolam in the rat. J Pharmacokin Biopharm. 1991;19:485–496.
Beal SL, Sheiner LB. NONMEM Users Guides. NONMEM Project Group, University of California at San Francisco, 1992.
Schoemaker RC, Cohen AF. Estimating impossible curves using NONMEM. B J Clin Pharmacol. 1996;42:283–290.
Hashimoto Y, Mori S, Hama N, Nakao K, Imura H, Yamaguchi M, Yasuhara M, Hori R. Nonlinear mixed effect modelling of the pharmacodynamics of natriuretic peptides in rats. J Pharmacokin Biopharm. 1993;21:281–297.
Hashimoto Y, Odani A, Tanigawara Y, Yasuhara M, Okuno T, and Hori R. Population analysis of the dose-dependent pharmacokinetics of zonisamide in epileptic patients. Biol Pharm Bull. 1994;17:323–326.
Hashimoto Y, Ozaki J, Koue T, Odani A, Yasuhara M, Hori R. Simulation for the analysis of distorted pharmacodynamic data. Pharm Res. 1994;11:545–548.
Beal SL, Sheiner LB. NONMEM Users Guide—Part VII. Conditional estimation methods. University of California at San Francisco, November 1992.
Rodman JH, Silverstein K. Comparison the two stage (TS) and first order (FO) methods for estimation of population parameters in an intensive pharmacokinetic study. Clin Pharmacol Ther. 1990;47:151.
Godfrey KR, Fitch WR. The deterministic identifiability of nonlinear pharmacokinetic models. J. Pharmacokin Biopharm. 1984;12:177–191.
Godfrey KR, Fitch WR. On the identification of Michaelis-Menton elimination parameters from a single dose-response curve. J Pharmacokin Biopharm. 1984;12:193–221.
Godfrey KR, Chapman MJ, Vajda S. Identifiability and indistinguishability of nonlinear pharmacokinetic models. J Pharmacokin Biopharm. 1994;22:229–251.
Hashimoto Y, Koue T, Otsuki Y, Yasuhara M, Hori R, Inui K. Simulation for population analysis of Michaelis-Menton kinetics. J Pharmacokin Biopharm. 1995;23:205–216.
Author information
Authors and Affiliations
Corresponding author
Additional information
Published: September 25, 2000
Rights and permissions
About this article
Cite this article
Jonsson, E.N., Karlsson, M.O. & Wade, J.R. Nonlinearity detection: Advantages of nonlinear mixed-effects modeling. AAPS PharmSci 2, 32 (2000). https://doi.org/10.1208/ps020332
Received:
Accepted:
Published:
DOI: https://doi.org/10.1208/ps020332