Abstract
Fas ligand (FasL) is a type II transmembrane protein belonging to the tumor necrosis factor family. Its binding to the cognate Fas receptor triggers the apoptosis that plays a pivotal role in the maintenance of immune system homeostasis. The cell death-inducing property of FasL has been associated with its extracellular domain, which can be cleaved off by metalloprotease activity to produce soluble FasL. The fate of the remaining membrane-anchored N-terminal part of the FasL molecule has not been determined. Here we show that post-translational processing of overexpressed and endogenous FasL in T-cells by the disintegrin and metalloprotease ADAM10 generates a 17-kDa N-terminal fragment, which lacks the receptor-binding extracellular domain. This FasL remnant is membrane anchored and further processed by SPPL2a, a member of the signal peptide peptidase-like family of intramembrane-cleaving proteases. SPPL2a cleavage liberates a smaller and highly unstable fragment mainly containing the intracellular FasL domain (FasL ICD). We show that this fragment translocates to the nucleus and is capable of inhibiting gene transcription. With ADAM10 and SPPL2a we have identified two proteases implicated in FasL processing and release of the FasL ICD, which has been shown to be important for retrograde FasL signaling.
Similar content being viewed by others
Main
Fas ligand (FasL; CD95L; CD178; tumor necrosis factor (TNF)SF6) is a glycosylated 40 kDa type II transmembrane protein belonging to the TNF superfamily of cytokines. Binding to its cognate Fas receptor triggers cell death in target cells,1 which is crucial for immune system function and has also been discussed in the context of immune privilege and tumorigenesis.2 Whereas many tissues constitutively express Fas receptor, cellular FasL expression is tightly regulated at transcriptional and post-translational levels.3 In hematopoietic cells, the protein is stored intracellularly and becomes externalized only in response to activation signals (e.g. target cell contact4). This regulated discharge can also lead to the production of enclosed membranous structures, termed microvesicles, which carry FasL on their surface.4, 5
FasL protein is efficiently proteolyzed by a poorly defined metalloprotease activity. This results in shedding of the receptor-binding FasL ectodomain (referred to as soluble Fas ligand (sFasL)), the biological role of which is still disputed (reviewed in1). sFasL can either cause apoptosis at a distant site6, 7 or alternatively dampen the apoptotic response by blocking Fas receptors.8, 9 The fate of the remaining membrane-anchored N-terminal portion of FasL, which comprises a transmembrane region and the proline-rich FasL intracellular domain (FasL ICD), has not been studied so far.
Several reports have demonstrated that FasL can transduce co-stimulatory/inhibitory signals into FasL-bearing cells.10, 11, 12, 13 For example, crosslinking cell surface FasL during T-cell receptor activation led to maximal CD8+ T-cell proliferation, indicating an ancillary role of FasL signaling in T-cell activation.11, 12, 13 This phenomenon, known as ‘reverse’ or ‘retrograde’ signaling, has also been associated with other members of the TNF superfamily. Its common outcomes are changes in proliferation and cytokine production profiles of the affected cells.1 Interestingly, FasL reverse signaling leading to T-cell co-stimulation is induced by the expression of the FasL cytoplasmic domain alone.11
The primary structure of the FasL protein suggests the presence of several signaling motifs within its intracellular portion that are well conserved across mammalian species (14 and references therein), for example, a casein kinase I (CKI) consensus site and a unique polyproline stretch. Different Src homology 3 (SH3)- or WW-domain-containing proteins have been reported to bind to the FasL proline-rich domain in various assays,1 among them members of the Fes/CIP4 homology/SH3 protein family, which regulate subcellular FasL distribution.14, 15 Nonetheless, the functional significance of the reported interactions for FasL reverse signaling remains unclear.
Here, we describe how FasL is processed by the disintegrin and metalloprotease ADAM1016 and by the intramembrane protease SPPL2a,17 ultimately leading to the liberation of the FasL ICD into the cytosol. The released FasL fragment is highly unstable and capable of translocating into the nucleus and influencing gene transcription.
Results
Endogenous FasL is upregulated and proteolytically processed on T-cell stimulation
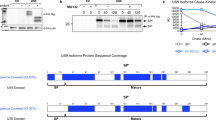
To investigate FasL processing in detail and to study the fate of the N-terminal remnant after shedding of the extracellular sFasL fragment, we employed two different antibodies, recognizing either the extracellular C-terminus (G247) or the intracellular N-terminus (Ab-3) of FasL (see cartoon in Supplementary Figure 1a). First, we used human embryonic kidney 293 (HEK293) cells stably transfected with an N-terminally FLAG-tagged full-length hFasL construct to confirm the specificity of the antibodies. Western blot analysis of the cell lysates revealed several glycosylated FasL full-length isoforms18 around 37–40 kDa (Figure 1a, left panel). The full-length FasL form was clearly detected with the extracellular domain-specific G247 antibody, but seemed also visible with the Ab-3 antibody and the anti-FLAG antibody M2. However, the surprisingly low intensity of the bands in the size range of 40 kDa disclosed by Ab-3 and M2 (in comparison to G247), and the apparent inability of Ab-3 to identify full-length FasL in stimulated human T-cells (see Supplementary Figure 1b) led us to the conclusion that Ab-3 is not capable to detect full-length FasL in Western blot analysis. With both the Ab-3 and the M2 antibodies (but not with G247), we observed smaller FasL fragments containing the N-terminal ICD, in particular a 17-kDa peptide, which we later called APL (for ADAM10-processed FasL form; Figure 1a).
FasL processing leads to the generation of different N-terminal fragments containing the intracellular FasL domain. (a) Cell lysates from 293 cells stably transfected with FLAG-hFasL or with empty vector were analyzed for FasL expression by immunoblotting using the G247 antibody (which recognizes the extracellular C-terminus of FasL; left panel), the Ab-3 antibody (which recognizes the intracellular N-terminus of FasL; middle panel) or the M2 anti-FLAG antibody (which recognizes the N-terminal FLAG-tag fused to the intracellular FasL domain; right panel). Whereas the G247 antibody exclusively detects several differently glycosylated full-length FasL bands between 37 and 40 kDa, the Ab-3 and the M2 antibodies recognize distinct truncated N-terminal FasL fragments between 14 and 18 kDa. Absence of these fragments in the cell lysates transfected with empty vector confirms the specificity of the detected signals. (b) PHA/IL-2 preactivated human T-cells were left inactivated (‘Ø’) or were activated with anti-CD3 (1 μg/ml) or anti-CD3/anti-CD28 antibodies (1 μg/ml each). After 24 h, the cells were collected and subjected to SDS-PAGE and Western blot analysis with anti-FasL antibodies. G247 identifies the full-length form of endogenous FasL (‘FL’), and Ab-3 detects both a 17-kDa fragment (APL for ADAM10-processed FasL fragment) and a smaller 11-kDa band (SPA for SPPL2a-processed APL). APL (strong signal) and SPA (weak signal) are detected simultaneously by Ab-3 on long exposure of the film (right panel). In the long exposure, additional unspecific bands above 30 kDa are appearing, which do not vary in intensity despite different FasL expression levels. Ezrin detection is used as a loading control
We then investigated whether similar processing can be seen with endogenous FasL. Lysates were prepared from activated and nonactivated primary human T-cells and analyzed by immunoblotting with the G247 and Ab-3 antibodies. As shown in Figure 1b, stimulation of the T-cells with either anti-CD3 antibody alone or with anti-CD3/-CD28 antibodies leads to an increase in the amount of full-length FasL protein detected by G247. This observation is in accordance with the upregulation of FasL mRNA on T-cell activation19 (also our own unpublished data). Interestingly, staining with the Ab-3 antibody revealed the same APL FasL fragment observed with overexpressed FasL. The APL fragment was already visible in nonactivated T-cells, and its production was boosted on anti-mCD3 stimulation. In contrast to 293 cells overexpressing FasL, only one FasL fragment (APL) was recognized in the size region of 17 kDa in T-cells, presumably because overexpressed FasL is extensively ‘trimmed’ in 293 cells (compare Figure 1a with Figure 1b).
We also observed a small amount of a low-abundance 11 kDa FasL peptide in T-cells, which we later called SPA (for SPPL2a-processed APL). The SPA peptide was not detectable in resting T-cells and was produced on anti-mCD3 antibody stimulation. T-cell activation with anti-mCD28 antibody alone does not lead to FasL upregulation and has no influence on APL and SPA production (data not shown).
Collectively, these data demonstrate that overexpressed FasL in 293 cells and endogenous FasL in T-cells are processed, constitutively or under CD3 stimulation, respectively, into different smaller fragments lacking the extracellular cell death-inducing FasL domain, but still containing the cytoplasmic FasL tail.
Processing of FasL by ADAM10 leads to the generation of the 17-kDa APL fragment
The ectodomain of a high proportion of FasL molecules is shed from the cell surface.1 Matrix metalloproteinases have been reported to be involved in the release of sFasL, and several matrix metalloprotease (MMP)7 (matrilysin) cleavage sites have been identified in the human and murine FasL sequences (20 and references therein). However, as the ADAM family members ADAM10 and ADAM17 have been implicated in TNFα processing,21 we examined a potential role for both proteases in FasL shedding and production of APL in 293 cells. In the first experiment, we used the metalloproteinase inhibitor GI254023X, which preferentially blocks ADAM10,22 on stable 293 transfectants overexpressing hFasL, and analyzed FasL processing by immunoblotting. Figure 2a demonstrates that incubation of the cells with the ADAM inhibitor increases the amount of full-length FasL (upper panel, G247 antibody), whereas production of the 17-kDa APL fragment declines (lower panel, M2 antibody). FACS analysis with the anti-FasL antibody Nok-1 (recognizing the extracellular C-terminus) confirmed increased cell surface expression of overexpressed full-length FasL on GI254023X-treated 293 cells.
ADAM10 cleaves overexpressed FasL in 293 cells and endogenous FasL in T-cells to produce the 17-kDa APL fragment. (a) 293 cells with stable FLAG-hFasL expression were incubated with or without the ADAM 10 inhibitor GI254023X (10 μ M; 24 h), before cell lysates were subjected to immunoblotting. G247 antibody was used to detect full-length FasL (‘FL’), and the anti-FLAG antibody M2 the N-terminal 17-kDa APL fragment. FACS analysis (bottom panel) was performed with the anti-FasL antibody NOK-1 (which recognizes the C-terminal part of the extracellular FasL domain) to quantify full-length FasL at the cell surface. The dashed curves represent cells either unstained or incubated only with secondary antibody (‘controls’), the solid curve stands for cells without (‘−GI254023X‘) and the thick line for cells with (‘+GI254023X‘) inhibitor. (b) 293 cells stably expressing FLAG-hFasL were transiently transfected with 6 μM ADAM10 (left column) or ADAM17 siRNA (right column). After 16 h, cell lysates were prepared and analyzed by Western blotting which revealed efficient knockdown of siRNA targets (upper panels). G247 and M2 antibodies were used again to detect full-length FasL (‘FL’) and the APL fragment. Cells transfected with unrelated control siRNA directed against the Drosophila SIMA RNA served as a negative control (‘−‘). FACS analysis with NOK-1 antibody was used as in (a) to quantify full-length FasL surface expression. (c, d) PHA/IL-2 preactivated human T-cells stimulated for 24 h with anti-CD3 antibody (1 μg/ml) were pretreated (‘+’) with GI254023X (10 μ M; 24 h) or left untreated (‘−‘; c) or transiently transfected with 6 μM of either ADAM10/17 (‘+’) or unrelated (‘−‘) siRNA (d), before cell lysates were prepared for Western blot analysis with G247 and Ab-3 antibodies. In (d), the efficiency of ADAM 10/17 silencing by siRNA transfection was confirmed by using anti-ADAM10/17 antibodies. Ezrin served again as a loading control. In (a–d), inhibition of ADAM10, but not of ADAM17 activity, leads to an increase in full-length FasL (‘FL’) and diminished levels of APL in FasL overexpressing 293 cells and in T-cells
We also incubated FasL-expressing 293 cells with other protease inhibitors, among them very specific small molecule ADAM10 and/or ADAM17 inhibitors which have been described recently.23 Like GI254023X and the broad MMP and TNF-α-converting enzyme (TACE) inhibitor TAPI-2, the ADAM10 inhibitors INCB-3619 (which suppresses both ADAM10 and ADAM17 activity) and INCB-8765 (specifically inhibiting ADAM10) prevented production of APL, leading to an accumulation of full-length FasL at the cell surface (see Supplementary Figure 2). In contrast, the ADAM17-specific inhibitor INCB-12881 had no such effect.
We then applied small interfering RNA (siRNA) to downregulate endogenous ADAM10 and ADAM17 in 293 cells overexpressing FasL. Transfection with ADAM10-specific siRNA resulted in efficient impairment of ADAM10 protein translation (Figure 2b, left column, upper panel) and decreased production of the APL FasL fragment (bottom panel), as observed with the ADAM10 inhibitor. Simultaneously, the amount of full-length FasL at the cell surface increased, as judged by immunoblotting (middle panel) and FACS analysis. On the other hand, ADAM17 downregulation did not influence the extent of APL formation and FasL processing (Figure 2b, right column), again suggesting that FasL is specifically cleaved by ADAM10 to produce the APL fragment.
As a next step, we analyzed endogenous FasL processing by ADAM10 in activated primary T-cells. Again, incubation with the ADAM10 inhibitor GI254023X led to an accumulation of full-length FasL and diminished generation of the 17-kDa APL fragment (Figure 2c). Interestingly, a slightly larger band could be detected instead (see also Figure 4c). This band corresponds in size to a recombinant 129 residue FasL peptide (data not shown) that is possibly cleaved by a member of the ADAM family. Further experiments with in vitro-translated N-terminal FasL peptides showed that APL itself corresponds in size to a 126 residue FasL fragment containing the ICD and the transmembrane region plus a short stretch of extracellular amino acids (data not shown).
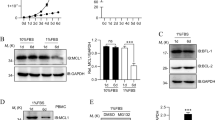
Shedding of endogenous FasL by ADAM10 in T-cells is necessary for subsequent intramembrane cleavage of APL and SPA production by SPPL2a. (a) Human peripheral blood T-cells were transfected with 4 μg each of a mock construct (‘Ø’), with wild-type SPPL2a (SPPL2a WT) or with the SPPL2aD412A active site mutant (SPPL2a DN). After 24 h, cell lysates were analyzed for the production of the endogenous FasL APL fragment by Western blot using the Ab-3 antibody. Ezrin detection was used as a loading control. (b) PHA/IL-2 pre-activated human T-cells, stimulated with anti-CD3 antibody (24 h, 1 μg/ml) and incubated with lactacystin (10 μM) to prevent protein degradation, were treated without additions (‘Ø’), or with DMSO alone, or with the SPP inhibitor (Z-LL)2 (100 μ M; 5 h) to block SPPL2a activity. Cell lysates were analyzed for generation of the endogenous APL and SPA fragments with Ab-3 antibody. G247 antibody detects full-length FasL (‘FL’). (c) PHA/IL-2 pre-activated primary human T-cells were left untreated (‘Ø’), or were incubated with the indicated protease inhibitors (each 10 μM). 1.5 h later, cells were activated with anti-CD3 antibody (1 μg/ml), and 3 h after the beginning of anti-CD3 stimulation, lactacystin was added at 10 μM for 6 h. At the end of the incubation time, cells were collected and subjected to SDS-PAGE and Western blot analysis with anti-FasL antibodies (G247 to detect full-length FasL (‘FL’) and Ab-3 to identify both the APL and the SPA fragments). Ezrin detection was used as a loading control
We also incubated activated T-cells with the small molecule ADAM10 and ADAM17 inhibitors. As can be seen in Figure 4c, suppressing ADAM10 activity led to the disappearance of APL and an increase in the amount of full-length FasL, whereas specific inhibition of ADAM17 had no influence on APL production. This result is in accordance with our data obtained in 293 cells.
In an alternative approach, we transfected stimulated primary T-cells with ADAM10 or ADAM-17-specific siRNA or with an unrelated control siRNA. Figure 2d illustrates a reduction in endogenous ADAM10 and ADAM17 protein levels on transfection with the appropriate siRNAs. Again, downregulation of ADAM10 (but not of ADAM17) led to an increase in the amount of unprocessed full-length FasL, whereas production of the APL fragment ceased. Apparently, ADAM10 is the protease responsible for the removal of the FasL ectodomain and for simultaneous generation of the remnant APL fragment.
The intramembrane protease SPPL2a cleaves the FasL APL fragment to produce an 11-kDa SPA peptide.
Transmembrane proteins that undergo proteolytic removal of their ectodomains are frequently further processed. This proteolysis, catalyzed by a unique group of intramembrane proteases liberates cytoplasmic domains with a potential signaling role24 (e.g., presenilin (PS)-dependent γ-secretase involved in Notch signaling25). Using chemical inhibitors and PS-deficient cells, we tested whether FasL might serve as a substrate for PS. However, no PS-dependent FasL processing was detectable (data not shown). This may be explained by the type I transmembrane protein-specificity of PS.26 Recently, a new group of proteases that belong to the signal peptide peptidase-like (SPPL) family of intramembrane-cleaving proteases (I-CliPs) has been described.17 They display no particular sequence homology to PS, but all contain the same active-site motifs, YD and GxGD. The crucial difference, however, is the orientation of their catalytic domain, which suggests their propensity for type II transmembrane protein substrates.17, 26
To test the potential involvement of SPPL2a protease in FasL intramembrane cleavage, we co-expressed SPPL2a together with FLAG-FasL in 293T cells. Intriguingly, we observed that the 17-kDa FasL APL fragment is further cleaved by SPPL2a, generating a 13-kDa SPA fragment (Figure 3a, lane 2; the SPA fragment produced from overexpressed FasL contains an additional N-terminal FLAG-tag; also note the decrease in the amount of APL in comparison to lane 1). SPA roughly corresponds in size to a recombinant FLAG-tagged fragment consisting of the ICD FasL (aa 1–80), expressed and immunoprecipitated in parallel as reference (Figure 3a, lane 7). However, when an active site mutant of SPPL2a (SPPL2aD412A) was transfected, SPA could not be detected. Instead, the 17-kDa APL fragment accumulated, indicating its continuous conversion through an endogenous SPPL2a activity in 293T cells, which is inhibited by SPPL2aD412A (Figure 3a, compare lane 3 with lane 1). FasL APL processing by SPPL2a appears to be specific, as overexpression of another member of the family, the SPP protein (wildtype or active site mutant SPPD265A), did not affect FasL proteolysis under the same experimental conditions (Figure 3a, lanes 5, 6). To further confirm that this cleavage is catalyzed by an SPPL family protease, we used the SPP inhibitor (Z-LL)2-ketone.27 As expected, it prevented SPPL2a-induced cleavage of FasL in 293T cells (Figure 3a, lane 4).
Processing of the FasL APL fragment by the intramembrane protease SPPL2a generates the 11-kDa SPA peptide in 293 cells. (a) 293T cells were transfected with 5 μg each of the following constructs: lane 1, Flag-hFasL alone; lanes 2 and 4, Flag-hFasL and wild-type (WT) SPPL2a; lane 3, Flag-hFasL and the SPPL2aD412A active site mutant (here referred to as dominant-negative; DN); lane 5, hFlag-FasL and wildtype (WT) SPP; lane 6, Flag-FasL and the SPPD265A DN mutant; lane 7, the Flag-tagged intracellular FasL domain (aa 1–80). The sample in lane 4 was derived from cells treated with the SPP inhibitor (Z-LL)2 (6 h, 100 μM). Flag-FasL was immunoprecipitated from cell lysates and analyzed by Western blotting with HRP-conjugated M2 anti-Flag antibody. The same blot is shown after different exposure times. (b) 293T cells transiently transfected with 5 μg of Flag-FasL were either left untreated (‘Ø’) or were incubated with the protease inhibitors lactacystin or lactacystin plus (Z-LL)2 ketone. Flag-FasL was immunoprecipitated from cell lysates and analyzed by immunoblotting with M2 anti-Flag antibody, followed by incubation with HRP-conjugated secondary anti-mouse antibody. The arrow indicates liberated 11-kDa FasL SPA fragment (which is only seen on proteasome inhibition), the asterisk shows the light chain of the IP antibody. (c) 293 cells stably expressing FLAG-hFasL were transiently transfected with 5 μg of SPPL2a-HA (left column) or SPPL2b-HA (right column) cDNA. In addition, siRNA (6 μM) specific for SPPL2a or SPPL2b (‘+’), or, as a control, unrelated SIMA siRNA (‘−‘) were co-transfected. Cell lysates were analyzed by Western blotting with G247 (full-length FasL; ‘FL’) and M2 (FasL APL fragment) antibodies. Immunoblotting with anti-HA tag antibody revealed efficient knockdown of SPPL2a and SPPL2b. Only the specific downregulation of SPPL2a leads to accumulation of APL
Despite endogenous SPPL2a activity, the released 13-kDa FLAG-tagged SPA fragment remained undetectable in 293T cells expressing FasL in the absence of SPPL2a overexpression (see Figure 3a, lane 1). Proteolytically released ICDs of many other transmembrane proteins are short-lived and removed by the proteasome.28, 29, 30 Indeed, treatment of FasL-expressing 293T cells with the proteasome inhibitor lactacystin facilitated detection of soluble SPA generated by endogenous SPPL2a activity (Figure 3b, lane 2). The use of (Z-LL)2 ketone abolished SPA production, again confirming the dependence of SPA generation on SPPL2a (Figure 3b, lane 3).
We also applied siRNA as an alternative approach to interfere with SPPL2a function. 293 cells stably expressing hFasL were transiently transfected with SPPL2a, together with either SPPL2a-specific or control siRNA (Figure 3c, left column). Western blot analysis revealed the efficient siRNA knockdown of overexpressed SPPL2a (top panel) and a strong accumulation of the APL fragment (bottom panel), indicating that its further intramembrane cleavage by SPPL2a was prevented. This result is in accordance with the data obtained using the chemical SPP inhibitor (Z-LL)2 (Figure 3a, b). In a similar experiment, transfection of SPPL2b-specific siRNA did not lead to APL accumulation (Figure 3c, right column), although we had observed production of a small amount of SPA fragment in 293T cells on transient co-overexpression of FasL and SPPL2b (data not shown). The siRNA knockdown experiments strongly suggest specific processing of the FasL APL fragment by endogenous SPPL2a.
To analyze SPPL2a cleavage of endogenous FasL in T-cells, we transfected wild-type SPPL2a or the SPPL2a active site mutant into stimulated peripheral blood cells. As in 293T cells, overexpression of SPPL2a led to the disappearance of the ADAM10 cleavage product APL, whereas APL levels slightly increased when the active site mutant SPPL2a was transfected (Figure 4a). The relatively small extent of APL accumulation on SPPL2aD412A overexpression is probably due to the low transfection efficiency currently achievable with primary T-cells.
We then again used the SPP inhibitor (Z-LL)2-ketone to block endogenous SPPL2a activity in primary T-cells and observed a profound accumulation of the (endogenous) FasL APL fragment. When we applied lactacystin to prevent protein degradation, we could observe production of the 11-kDa SPA peptide, which was abolished when the cells were treated with (Z-LL)2-ketone, emphasizing the notion that APL is a physiological target of the intramembrane protease SPPL2a (Figure 4b).
We wanted to find out whether FasL processing and production of APL by ADAM10 is required for the subsequent generation of SPA by SPPL2a. Primary human T-cells were activated with anti-mCD3 antibody and treated simultaneously with lactacystin to allow detection of SPA. The cells were then incubated with different protease inhibitors, including GI254023X, TAPI-2 and several small molecule ADAM10/17 inhibitors. The upper panel of Figure 4c shows the increase in the amount of endogenous full-length FasL on ADAM10 inhibition by several of the inhibitors, including the very specific ADAM10 inhibitor INCB-8765. Consequently, the ADAM10 cleavage product APL disappeared (instead, a different, slightly larger band could be detected, as already seen in Figure 2c, which is absent in 293 cells). Specific inhibition of ADAM17 (by INCB-12881) did not affect APL production. Strikingly, not only APL, but also the SPPL2a cleavage product SPA was no longer visible after ADAM10 inhibition, strongly suggesting that FasL shedding by ADAM10 is a prerequisite for subsequent intramembrane cleavage by SPPL2a in T-cells. Interestingly, such disappearance of SPA after ADAM10 inhibition is not observed in stably transfected 293 cells overexpressing FasL (see Supplementary Figure 2a).
After processing by SPPL2a, the released intracellular FasL domain translocates to the nucleus
Proteolytically liberated intracellular domains of various transmembrane proteins translocate to the nucleus.31 Analysis of transfected COS-7 cells by confocal microscopy revealed strong nuclear localization of a peptide comprising the FasLICD (aa 1–80; Figure 5a, lower panel). In contrast, N-terminally FLAG-tagged full-length FasL exhibited only membrane and cytoplasmic staining, lacking any obvious nuclear localization due to the low abundance of liberated SPA (Figure 5a, upper panel).
The FasL SPA fragment is released by SPPL2a cleavage and translocates to the nucleus. (a) COS-7 cells were transiently transfected with 2 μg of full-length Flag-hFasL or with a FLAG-tagged construct consisting of the intracellular FasL domain (aa 1–80), which roughly corresponds to the FasL SPA fragment (Flag-SPA). Fixed cells were permeabilized, stained with anti-FLAG antibody and analyzed by laser scanning confocal microscopy. Nuclei were visualized with the DNA stain DRAQ5. (b) COS-7 cells were transiently transfected with 2 μg of N-terminally EGFP-tagged full-length hFasL (EGFP-FasL) together with wild-type HA-tagged human SPPL2a (SPPL2a-HA) or with the active site mutant SPPL2aD412A (SPPL2a(DN)-HA). After 24 h, cells were treated overnight with lactacystin (10 μM) to stabilize the intracellular FasL fragment. Fixed cells were processed for confocal imaging as described above
To demonstrate nuclear localization of the SPA fragment released by intramembrane proteolysis, we co-expressed FasL (with EGFP fused to its N-terminus) with SPPL2a in COS-7 cells. Lactacystin was used to increase the stability of SPA after its generation. Under these conditions, we observed a high proportion of released SPA localizing to the nucleus (Figure 5b, upper panel). Importantly, co-expression of the active site mutant of SPPL2a did not produce a similar effect (Figure 5b, lower panel). The confocal localization studies also demonstrated that FasL colocalizes with SPPL2a protease, which strongly supports the notion of FasL/SPPL2a interaction (Figure 5b).
The intracellular FasL domain inhibits Gal4-mediated gene transcription
The reverse signaling potential of FasL implies that it may affect gene expression. To explore the nuclear role of liberated SPA and to test whether it influences transcription, we employed a standard Gal4-based reporter assay. As shown in Figure 6, a Gal4 protein fused to the FasL ICD (aa 1–80) inhibited Gal4-responsive promoter activity in a dose-dependent manner. This Gal4-SPA-mediated transcriptional repression was comparable to the transrepression of a Gal4–nuclear receptor coreppressor (NcoR) fusion protein.32 An NCoR fragment lacking repression domains (aa 2174–2453) did not influence the reporter activity (data not shown). Thus, SPA-mediated transcription repression strongly suggests that this fragment acts as a transcriptional regulator in the nucleus.
The intracellular FasL domain inhibits Gal4-mediated gene transcription. 293T cells were transfected with 0.1 μg of a Gal4 reporter construct, together with 0.1 μg of a plasmid encoding either the Gal4 transactivation factor alone (Gal4), or 0.1–0.3 μg of Gal4 fused to the intracellular FasL domain (aa 1–80; Gal4-SPA), or 0.1 μg of the transcriptional repressor NCoR (Gal4-NCoR). After 48 h, cell lysates were assayed for Gal4-induced luciferase activity. RLU, relative luciferase units. The experiment was performed twice, each time in duplicates. Error bars indicate standard deviations
Discussion
FasL is well characterized as a potent inducer of apoptosis.33 Much less is known about the molecular mechanisms underlying its capacity to transduce nonapoptotic signals into the ligand-bearing cell. FasL reverse signaling has been shown to enhance T-cell proliferation,11, 12, 13 and a recent publication has demonstrated that expression of the cytoplasmic FasL domain alone is sufficient to costimulate T-cell receptor signals.11
Nevertheless, it remains unclear exactly how the FasL ICD is engaged in the transmission of such costimulatory signals. Our data reveal a novel regulated processing of endogenous FasL in T-cells by the metalloproteinase ADAM 10 and the intramembrane protease SPPL2a, which ultimately leads to liberation of the FasL ICD into the cytosol. Members of the ADAM protease family are involved in the proteolysis of various substrates, such as Notch, EGF ligands and others (34 and references therein). By using different inhibitors specific for the disintegrin-like metalloproteinase ADAM1022, 23 and by using a genetic siRNA knockdown approach, we have shown that ADAM10 is one of the proteases responsible for ectodomain shedding of overexpressed FasL in 293 cells and of endogenous FasL in T-cells. Blockage of ADAM10 (but not of ADAM17) activity results in an accumulation of full-length FasL and a decrease in the FasL ADAM10 cleavage product APL. Processing by the shedding protease ADAM10 is the first step of a regulated intramembrane proteolysis (RIP) of FasL in T-cells, which is executed in the second step by SPPL2a, a member of the SPPL family.35 In contrast to presenilin, SPPL proteases catalyze the intramembrane proteolysis of type II anchored membrane proteins.17 Our inhibitor and siRNA knockdown experiments demonstrated that intramembrane cleavage of the FasL remnant APL leads to the liberation of the 11-kDa SPA fragment, which consists of the FasL ICD. FasL processing by SPPL2a is further supported by co-immunoprecipitation experiments, which demonstrate a direct interaction between both molecules (data not shown). Our data suggest that, at least in T-cells, processing of FasL by SPPL2a requires production of APL by ADAM10. Surprisingly, we did not observe abrogation of SPA production on treatment of FasL-overexpressing 293 cells with ADAM10 inhibitors, as we had discovered in T cells. A possible explanation could be the unphysiological high expression level of transfected FasL, which may lead to direct processing of full-length FasL (instead of APL) by SPPL2a in 293 cells.
To our knowledge, FasL is only the second physiological target protein identified for members of the SPPL protease family. Two recent publications report TNFα intramembrane cleavage by SPPL2a and SPPL2b,36, 37 which triggers expression of the pro-inflammatory cytokine interleukin-12 (IL-12) by activated human dendritic cells.37 Together with our data on FasL processing by SPPL2a, these results indicate an emerging important role of SPPLs in RIP of TNF family members.
Several transmembrane proteins are subjected to intramembrane proteolysis, as a result of which the ICD is released for signaling to the nucleus31, 37 (also see references therein). Our findings of a similar FasL RIP by ADAM10 and SPPL2a prompted us to investigate whether the FasL ICD (liberated by SPPL2a cleavage) could translocate to the nucleus and influence gene transcription. Indeed, we were able detect the FasL SPA fragment in the nucleus of cells overexpressing FasL and SPPL2A, and an overexpressed peptide representing the cytoplasmic FasL domain accumulated in the nucleus. In addition, we observed large amounts of this overexpressed ICD in the nuclear fraction obtained in fractionation experiments (data not shown). We identified a putative nuclear localization signal (NLS) in the FasL ICD domain sequence close to the membrane (aa 69–76); however, the relatively small size of the liberated SPA fragment should allow NLS-independent passive transport into the nucleus.
Our difficulties in detecting the FasL SPA fragment as a product of endogenous SPPL2a activity and the experiments with the proteasome inhibitor lactacystin indicate that the released FasL ICD is highly unstable. Proteolytically released ICD of many other transmembrane proteins are typically short-lived and rapidly degraded by the proteasome,28, 29, 30 which may represent an effective way to downregulate any (nuclear) signal provided by the released peptides. The Deltex protein, for example, has been proposed to act as an E3 ligase responsible for ubiquitination of the Notch ICD ((N)ICD).38
Having established the regulated proteolysis of FasL by ADAM10 and SPPL2a, we wanted to test whether the FasL ICD can influence gene transcription, similar to the intracellular Notch domain. We used an artificial Gal4 reporter system in our experiments, which proved that the intracellular FasL domain can inhibit gene transcription in principle. Whether FasL reverse signaling under physiological conditions is indeed dependent on transcriptional changes induced by the released intracellular domain remains to be investigated. It will, for example, be interesting to see whether FasL RIP by ADAM10 and SPPL2a provides the molecular basis for the observed costimulatory function of FasL reverse signaling.
Materials and Methods
Antibodies and reagents
The antibodies used for Western blotting were as follows: anti-Ezrin (Zymed Laboratories), anti-FasL G247-4 (PharMingen; in the manuscript referred to as G247), anti-FasL Ab-3 (Calbiochem), anti-FLAG (M2 and its HRP-conjugate; Sigma), anti-HA (12CA5; Roche), anti-ADAM10 and anti-ADAM17 (Chemicon) and horseradish peroxidase-conjugated anti-rabbit and anti-mouse antibodies (Jackson ImmunoResearch). Phytohemagglutinin (PHA), the SPP protease inhibitor (Z-LL)2-ketone and clastolactacystin (for simplicity, referred to as lactacystin) were purchased from Calbiochem, and IL-2 from Roche Diagnostics (Mannheim, Germany). The ADAM10 inhibitor GI254023X ((2R,3S)-3-(formyl-hydroxyamino)-2-(3-phenyl-1-propyl) butanoic acid ((1S)-2,2-dimethyl-1-methylcarbamoyl-1-propyl) amide) has been described elsewhere,22 the MMP and TACE inhibitor TAPI-2 (N-(R)-(2-(Hydroxyaminocarbonyl)methyl)-4-methylpentanoyl-L-t-butyl-alanyl-L-alanine, 2-aminoethyl amid) were purchased from Calbiochem. The small molecule protease inhibitors INCB-3619, INCB-3420, INCB-8765 and INCB-3420 were obtained from P. Scherle (Incyte Corporation, Wilmington, Delaware, USA) and have been described elsewhere.23
Cell culture and transfection procedures
Human peripheral blood cells were prepared from buffy coat samples by density gradient centrifugation over a Ficoll solution. Cells (1.5 × 106 cells/ml) obtained from the interlayer were cultured in RPMI 1640 medium (Gibco-BRL), supplemented with 10% FCS, 2 mM l-glutamine, 100 U/ml penicillin and 100 μg/ml streptomycin at 37°C in 5% CO2. The medium was complemented for the first 3 days with PHA (5 μg/ml) and for an extra 8 days with IL-2 (10 U/ml), before being activated with anti-CD3/CD28 antibodies (PharMingen). T-cells were transfected using the Amaxa nucleofection technology™ (Amaxa GmbH, Cologne, Germany). Cells were resuspended in the solution from the nucleofector kit V, following the Amaxa guidelines for cell line transfection. Briefly, a suspension of 8 × 106 cells in 100 μl buffer solution was mixed with 4 μg cDNA or 4 μg siRNA, transferred to the provided cuvette and nucleofected with an Amaxa Nucleofector apparatus using the T23 program. Cells were immediately transferred into six-well plates containing 37°C pre-warmed culture medium and cultured for 24 h in the presence of IL-2 before analysis.
HEK293 and HEK293T cells (American Type Culture Collection (ATCC) no. CRL-1573), and COS-7 cells (ATCC no. CRL-1651) were cultured in Dulbecco's Modified Eaglés Medium (Invitrogen), supplemented with 10% FCS (Biotech GmbH) in a 5% CO2 atmosphere. 293 cells were transiently transfected with the indicated DNA constructs using calcium phosphate precipitation in 10 cm culture dishes. Alternatively, transient transfections of 293T and COS-7 cells were performed using the Fugene 6 transfection reagent (Roche).
Flow cytometry
FACS analysis was performed with the anti-FasL antibody NOK-1 (PharMingen) according to standard protocols on a BD FACS Calibur.
Plasmids
N-terminally FLAG-tagged human (h)FasL was produced by PCR from a plasmid template and cloned into the mammalian expression vector pCR33 (kindly provided by Dr. Hermann Eibel, University Hospital of Freiburg, Germany), resulting in pCR33-FLAG-hFasL. N-terminally EGFP-tagged FasL (pEGFP-C-FasL) was kindly provided by O. Janssen, University of Kiel, Germany. The FLAG-SPA construct used for confocal microscopy codes for aa 1–80 of human FasL and was cloned with an N-terminal FLAG-tag into pCR33. C-terminally HA-tagged human SPP, SPPL2a and SPPL2b, as well as the active site mutants SPPL2aD412A and SPPD265A, were cloned by PCR into the vector pCI (Promega). For Gal4 reporter assays, a fragment of the FasL cDNA (encoding the intracellular aa 1–80) was cloned in frame with the C-terminus of a Gal4 fragment (aa 1–147) into the vector pCMX (kindly provided by E. Pfitzner, Frankfurt/Main, Germany), resulting in Gal4-SPA. pCMX-Gal4-NCoR, UAS-tk-luc Gal4 reporter and β-Gal transfection control constructs were generously supplied by T. Heinzel, University of Jena, Germany.
Immunoprecipitation and Western blot analysis
Western blot analysis of proteins was performed according to standard protocols. Briefly, cells were collected and directly resuspended in Laemmli buffer. After sonication and denaturing (95°C, 5 min), the solubilized proteins were resolved by sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE), and transferred to a polyvinyliden difluoride (PVDF) membrane (Immobilon-P, Millipore, Molsheim, France) by electroblotting.
For immunoprecipitation experiments, 3 × 106 293T cells were seeded in 10-cm plates, allowed to adhere and transfected with appropriate constructs. 2 days later, cells were lysed in ice-cold lysis buffer (30 mM Tris-HCl (pH7.4), 150 mM NaCl, 1 mM EDTA, 0.5% Triton X-100, 0.5% sodium deoxycholate, freshly supplemented with Complete Protease Inhibitor Cocktail (Roche)) and homogenized using syringes fitted with 25G needles. After a clearing step (centrifugation at 13 000 rpm for 15 min), lysates were subjected to immunoprecipitation with 1–2 μg of anti-FLAG M2 antibody (2 h to overnight at 4°C). Antibody–antigen complexes were precipitated by adding Protein A/G PLUS agarose (Santa Cruz) for another 2 h at 4°C. The immunoprecipitates were washed three times in lysis buffer, resuspended in loading buffer and resolved on 15% SDS-PAGE gels. Proteins were transferred onto PVDF membranes and analyzed by Western blotting with anti-FLAG M2 antibody. Non-radioactive reference peptides were produced using the Rabbit Reticulocyte Lysate System from Promega according to the manufacturer's guidelines, mixed with lysis buffer and immunoprecipitated as described above.
Generation and transfection of small interfering RNA
The following siRNA sequences were synthesized by Eurogentec (Liege, Belgium) and annealed according to the manufacturer's protocol:
ADAM10 (nt 532–550 of human ADAM10 cDNA; GeneBank accession number NM_001110):
sense: 5′-AGACAUUAUGAAGGAUUAUTT-3′
antisense: 5′- AUAAUCCUUCAUAAUGUCUTT -3′
ADAM17 (nt 280–299 of human ADAM17 cDNA; GeneBank accession number NM_003183):
sense: 5′-GAG AAG CUU GAU UCU UUG CTT-3′
antisense: 5′-GCA AAG AAU CAA GCU UCU CTT-3
SPPL2a (nt 929–947 of human SPPL2a cDNA; GeneBank accession number NM_032802):
sense: 5′-CUG UCU UGC UGC ACU AAU UTT-3′
antisense: 5′-AAU UAG UGC AGC AAG ACA GTT-3
SPPL2b (nt 423–441 of human SPPL2b cDNA; GeneBank accession number NM_152988):
sense: 5′-GAC GCA GUA UGA UGA GAU UTT-3′
antisense: 5′-AAU CUC AUC AUA CUG CGU CTT-3
The oligonucleotide sequences were subjected to a BLAST search, and no significant identities to other sequences were detected. The siRNA duplex sequences used as a negative control targeted the SIMA protein (Drosophila HIF) and showed no significant homology to any known protein, as assessed by the BLAST and NCBI databases. For 293 cells, 6 μM of siRNA were transfected with CaPO4, and for T-cells, 6 μM of siRNA were electroporated using Amaxa nucleofection technology™.
Confocal microscopy
COS-7 cells were grown on glass cover slips and transfected with the constructs of interest. At 2 days post-transfection, cells were fixed for 15 min with 2% formaldehyde, washed and permeabilized for 5 min with 0.2% Triton X-100. After blocking for 30 min in 10% FCS, cells were stained with primary antibody (anti-FLAG M2 (Sigma) and anti-HA (12CA5; Roche), both used at 1 : 500) for 1 h. Washed cells were then stained for another hour with secondary fluorochrome-conjugated antibody (polyclonal anti-mouse Alexa-488; Molecular Probes, used at 1 : 1000). Cells were washed again and incubated with DRAQ5 (Alexa) to counterstain the nuclei. Cover slips were then mounted in Moviol, and cell staining was documented using a Leica TCS SL Laser Scanning Confocal Microscope.
Gal4 reporter assay
293T cells (1 × 105 cells/well, using 12-well plates) were transfected with a mixture of 0.1 μg UAS-tk-luc, 0.1 μg β-Gal and 0.1–0.3 μg of the appropriate Gal4 fusion plasmids. After 48 h, cells were lysed using luciferase lysis buffer (50 mM MES-Tris (pH7.8), 10% glycerol, 0.1% Triton X-100, 1 mM DTT and Complete Protease Inhibitor Cocktail). Cleared lysates were utilized to determine luciferase activity, which was normalized against the β-Gal activity measured in the same sample.
Abbreviations
- FasL:
-
Fas ligand
- sFasL:
-
soluble Fas ligand
- APL:
-
ADAM10-processed FasL form
- SPA:
-
SPPL2a-processed APL
- (N)ICD:
-
(Notch) intracellular domain
- PS:
-
presenilins
- MMP:
-
matrix metalloprotease
- TACE:
-
tumor necrosis factor-alpha (TNF-α)-converting enzyme
- ADAM:
-
a disintegrin and metalloproteinase
- SPPL:
-
signal peptide peptidase-like
- RIP:
-
regulated intramembrane proteolysis
- I-CliP:
-
intramembrane-cleaving protease
- kDa:
-
kilo Dalton
- CKI:
-
casein kinase I
- FCH domain:
-
Fes/CIP4 homology domain
- SH3 domain:
-
Src homology 3 domain
- NCoR:
-
nuclear receptor coreppressor
- NLS:
-
nuclear localization signal
- PHA:
-
phytohemagglutinin
- DMEM:
-
Dulbecco's Modified Eaglés Medium
- HEK293 cells:
-
human embryonic kidney 293 cells
- ATCC:
-
American Type Culture Collection
- SDS-PAGE:
-
sodium dodecyl sulfate-polyacrylamide gel electrophoresis
- PVDF:
-
polyvinyliden difluoride
References
Janssen O, Qian J, Linkermann A, Kabelitz D . CD95 ligand – death factor and costimulatory molecule? Cell Death Differ 2003; 10: 1215–1225.
Li-Weber M, Krammer PH . Function and regulation of the CD95 (APO-1/Fas) ligand in the immune system. Semin Immunol 2003; 15: 145–157.
Kavurma MM, Khachigian LM . Signaling and transcriptional control of Fas ligand gene expression. Cell Death Differ 2003; 10: 36–44.
Martinez-Lorenzo MJ, Anel A, Gamen S, Monle n I, Lasierra P, Larrad L et al. Activated human T cells release bioactive Fas ligand and APO2 ligand in microvesicles. J Immunol 1999; 163: 1274–1281.
Bohana-Kashtan O, Civin CI . Profiling tumor counterattack: do Fas ligand-containing microvesicles reduce anticancer immunity? Clin Cancer Res 2005; 11: 968–970.
Dhein J, Walczak H, Baumler C, Debatin KM, Krammer PH . Autocrine T-cell suicide mediated by APO-1/(Fas/CD95). Nature 1995; 373: 438–441.
Tanaka M, Suda T, Takahashi T, Nagata S . Expression of the functional soluble form of human fas ligand in activated lymphocytes. EMBO J 1995; 14: 1129–1135.
Knox PG, Milner AE, Green NK, Eliopoulos AG, Young LS . Inhibition of metalloproteinase cleavage enhances the cytotoxicity of Fas ligand. J Immunol 2003; 170: 677–685.
Schneider P, Holler N, Bodmer JL, Hahne M, Frei K, Fontana A et al. Conversion of membrane-bound Fas(CD95) ligand to its soluble form is associated with downregulation of its proapoptotic activity and loss of liver toxicity. J Exp Med 1998; 187: 1205–1213.
Desbarats J, Duke RC, Newell MK . Newly discovered role for Fas ligand in the cell-cycle arrest of CD4+ T cells. Nat Med 1998; 4: 1377–1382.
Sun M, Ames KT, Suzuki I, Fink PJ . The cytoplasmic domain of Fas ligand costimulates TCR signals. J Immunol 2006; 177: 1481–1491.
Suzuki I, Fink PJ . Maximal proliferation of cytotoxic T lymphocytes requires reverse signaling through Fas ligand. J Exp Med 1998; 187: 123–128.
Suzuki I, Martin S, Boursalian TE, Beers C, Fink PJ . Fas ligand costimulates the in vivo proliferation of CD8+ T cells. J Immunol 2000; 165: 5537–5543.
Baum W, Kirkin V, Fernandez SB, Pick R, Lettau M, Janssen O et al. Binding of the intracellular Fas ligand (FasL) domain to the adaptor protein PSTPIP results in a cytoplasmic localization of FasL. J Biol Chem 2005; 280: 40012–40024.
Qian J, Chen W, Lettau M, Podda G, Zornig M, Kabelitz D et al. Regulation of FasL expression: a SH3 domain containing protein family involved in the lysosomal association of FasL. Cell Signal 2006; 18: 1327–1337.
White JM . ADAMs: modulators of cell–cell and cell–matrix interactions. Curr Opin Cell Biol 2003; 15: 598–606.
Friedmann E, Lemberg MK, Weihofen A, Dev KK, Dengler U, Rovelli G et al. Consensus analysis of signal peptide peptidase and homologous human aspartic proteases reveals opposite topology of catalytic domains compared with presenilins. J Biol Chem 2004; 279: 50790–50798.
Schneider P, Bodmer JL, Holler N, Mattmann C, Scuderi P, Terskikh A et al. Characterization of Fas (Apo-1, CD95)-Fas ligand interaction. J Biol Chem 1997; 272: 18827–18833.
Eischen CM, Williams BL, Zhang W, Samelson LE, Lynch DH, Abraham RT et al. ZAP-70 tyrosine kinase is required for the up-regulation of Fas ligand in activation-induced T cell apoptosis. J Immunol 1997; 159: 1135–1139.
Vargo-Gogola T, Crawford HC, Fingleton B, Matrisian LM . Identification of novel matrix metalloproteinase-7 (matrilysin) cleavage sites in murine and human Fas ligand. Arch Biochem Biophys 2002; 408: 155–161.
Zheng Y, Saftig P, Hartmann D, Blobel C . Evaluation of the contribution of different ADAMs to tumor necrosis factor alpha (TNFalpha) shedding and of the function of the TNFalpha ectodomain in ensuring selective stimulated shedding by the TNFalpha convertase (TACE/ADAM17). J Biol Chem 2004; 279: 42898–42906.
Hundhausen C, Misztela D, Berkhout TA, Broadway N, Saftig P, Reiss K et al. The disintegrin-like metalloproteinase ADAM10 is involved in constitutive cleavage of CX3CL1 (fractalkine) and regulates CX3CL1-mediated cell–cell adhesion. Blood 2003; 102: 1186–1195.
Zhou BB, Peyton M, He B, Liu C, Girard L, Caudler E et al. Targeting ADAM-mediated ligand cleavage to inhibit HER3 and EGFR pathways in non-small cell lung cancer. Cancer Cell 2006; 10: 39–50.
Fortini ME . Gamma-secretase-mediated proteolysis in cell-surface-receptor signalling. Nat Rev Mol Cell Biol 2002; 3: 673–684.
Martinez Arias A, Zecchini V, Brennan K . CSL-independent Notch signalling: a checkpoint in cell fate decisions during development? Curr Opin Genet Dev 2002; 12: 524–533.
Weihofen A, Martoglio B . Intramembrane-cleaving proteases: controlled liberation of proteins and bioactive peptides. Trends Cell Biol 2003; 13: 71–78.
Weihofen A, Lemberg MK, Ploegh HL, Bogyo M, Martoglio B . Release of signal peptide fragments into the cytosol requires cleavage in the transmembrane region by a protease activity that is specifically blocked by a novel cysteine protease inhibitor. J Biol Chem 2000; 275: 30951–30956.
Cowan JW, Wang X, Guan R, He K, Jiang J, Baumann G et al. Growth hormone receptor is a target for presenilin-dependent gamma-secretase cleavage. J Biol Chem 2005; 280: 19331–19342.
Gupta-Rossi N, Le Bail O, Gonen H, Brou C, Logeat F, Six E et al. Functional interaction between SEL-10, an F-box protein, and the nuclear form of activated Notch1 receptor. J Biol Chem 2001; 276: 34371–34378.
Kimberly WT, Zheng JB, Guenette SY, Selkoe DJ . The intracellular domain of the beta-amyloid precursor protein is stabilized by Fe65 and translocates to the nucleus in a notch-like manner. J Biol Chem 2001; 276: 40288–40292.
Carpenter G . Nuclear localization and possible functions of receptor tyrosine kinases. Curr Opin Cell Biol 2003; 15: 143–148.
Tagami T, Madison LD, Nagaya T, Jameson JL . Nuclear receptor corepressors activate rather than suppress basal transcription of genes that are negatively regulated by thyroid hormone. Mol Cell Biol 1997; 17: 2642–2648.
Fas SC, Fritzsching B, Suri-Payer E, Krammer PH . Death receptor signaling and its function in the immune system. Curr Dir Autoimmun 2006; 9: 1–17.
Reiss K, Maretzky T, Haas IG, Schulte M, Ludwig A, Frank M et al. Regulated ADAM10-dependent ectodomain shedding of gamma-protocadherin C3 modulates cell–cell adhesion. J Biol Chem 2006; 281: 21735–21744.
Martoglio B, Golde TE . Intramembrane-cleaving aspartic proteases and disease: presenilins, signal peptide peptidase and their homologs. Hum Mol Genet 2003; 12 (Spec No 2): R201–R206.
Fluhrer R, Grammer G, Israel L, Condron MM, Haffner C, Friedmann E et al. A gamma-secretase-like intramembrane cleavage of TNFalpha by the GxGD aspartyl protease SPPL2b. Nat Cell Biol 2006; 8: 894–896.
Friedmann E, Hauben E, Maylandt K, Schleeger S, Vreugde S, Lichtenthaler SF et al. SPPL2a and SPPL2b promote intramembrane proteolysis of TNFalpha in activated dendritic cells to trigger IL-12 production. Nat Cell Biol 2006; 8: 843–848.
Miele L, Miao H, Nickoloff BJ . NOTCH signaling as a novel cancer therapeutic target. Curr Cancer Drug Targets 2006; 6: 313–323.
Acknowledgements
We thank S Bösser for excellent technical assistance, and P Scherle (Incyte Corporation, Wilmington, Delaware, USA) for providing the small molecule inhibitors INCB-3619, INCB-3420, INCB-8765 and INCB-12881. This work was supported in part by grants from the DFG (ZO 110/2; MZ), the German National Genome Research Network (NGFN project N1KR-S12T23; VK and MZ), the Centre National de la Recherche Scientifique (AOH), the Ligue Nationale contre le Cancer (AOH), the Association pour la Recherche contre le Cancer (AOH), The Emerald Foundation (AOH) and the Canceropole-PACA ACI 2004 (AOH).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Edited by PH Krammer
Supplementary Information accompanies the paper on Cell Death and Differentiation website (http://www.nature.com/cdd)
Supplementary information
Rights and permissions
About this article
Cite this article
Kirkin, V., Cahuzac, N., Guardiola-Serrano, F. et al. The Fas ligand intracellular domain is released by ADAM10 and SPPL2a cleavage in T-cells. Cell Death Differ 14, 1678–1687 (2007). https://doi.org/10.1038/sj.cdd.4402175
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.cdd.4402175