Abstract
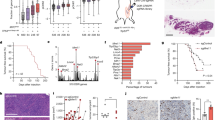
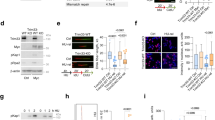
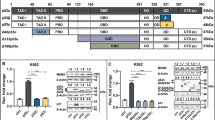
The RNA helicase p68 (DDX5) is an established co-activator of the p53 tumour suppressor that itself has a pivotal role in orchestrating the cellular response to DNA damage. Although several factors influence the biological outcome of p53 activation, the mechanisms governing the choice between cell-cycle arrest and apoptosis remain to be elucidated. In the present study, we show that, while p68 is critical for p53-mediated transactivation of the cell-cycle arrest gene p21WAF1/CIP1, it is dispensable for induction of several pro-apoptotic genes in response to DNA damage. Moreover, p68 depletion results in a striking inhibition of recruitment of p53 and RNA Pol II to the p21 promoter but not to the Bax or PUMA promoters, providing an explanation for the selective effect on p21 induction. Importantly, these findings are mirrored in a novel inducible p68 knockout mouse model in which p68 depletion results in a selective inhibition of p21 induction in several tissues. Moreover, in the bone marrow, p68 depletion results in an increased sensitivity to γ-irradiation, consistent with an increased level of apoptosis. These data highlight a novel function of p68 as a modulator of the decision between p53-mediated growth arrest and apoptosis in vitro and in vivo.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Vousden KH, Prives C . Blinded by the light: the growing complexity of p53. Cell 2009; 137: 413–431.
Espinosa JM . Mechanisms of regulatory diversity within the p53 transcriptional network. Oncogene 2008; 27: 4013–4023.
Murray-Zmijewski F, Slee EA, Lu X . A complex barcode underlies the heterogeneous response of p53 to stress. Nat Rev Mol Cell Biol 2008; 9: 702–712.
Linder P . Dead-box proteins: a family affair—active and passive players in RNP-remodeling. Nucleic Acids Res 2006; 34: 4168–4180.
Fuller-Pace FV, Moore HC . RNA helicases p68 and p72: multifunctional proteins with important implications for cancer development. Future Oncol 2011; 7: 239–251.
Endoh H, Maruyama K, Masuhiro Y, Kobayashi Y, Goto M, Tai H et al. Purification and identification of p68 RNA helicase acting as a transcriptional coactivator specific for the activation function 1 of human estrogen receptor alpha. Mol Cell Biol 1999; 19: 5363–5372.
Clark EL, Coulson A, Dalgliesh C, Rajan P, Nicol SM, Fleming S et al. The RNA helicase p68 is a novel androgen receptor coactivator involved in splicing and is overexpressed in prostate cancer. Cancer Res 2008; 68: 7938–7946.
Bates GJ, Nicol SM, Wilson BJ, Jacobs AM, Bourdon JC, Wardrop J et al. The DEAD box protein p68: a novel transcriptional coactivator of the p53 tumour suppressor. EMBO J 2005; 24: 543–553.
Metivier R, Penot G, Hubner MR, Reid G, Brand H, Kos M et al. Estrogen receptor-alpha directs ordered, cyclical, and combinatorial recruitment of cofactors on a natural target promoter. Cell 2003; 115: 751–763.
Wang XW, Zhan Q, Coursen JD, Khan MA, Kontny HU, Yu L et al. GADD45 induction of a G2/M cell cycle checkpoint. Proc Natl Acad Sci U S A 1999; 96: 3706–3711.
Caretti G, Schiltz RL, Dilworth FJ, Di Padova M, Zhao P, Ogryzko V et al. The RNA helicases p68/p72 and the noncoding RNA SRA are coregulators of MyoD and skeletal muscle differentiation. Dev Cell 2006; 11: 547–560.
Fukuda T, Yamagata K, Fujiyama S, Matsumoto T, Koshida I, Yoshimura K et al. DEAD-box RNA helicase subunits of the Drosha complex are required for processing of rRNA and a subset of microRNAs. Nat Cell Biol 2007; 9: 604–611.
Wallace M, Coates PJ, Wright EG, Ball KL . Differential post-translational modification of the tumour suppressor proteins Rb and p53 modulate the rates of radiation-induced apoptosis in vivo. Oncogene 2001; 20: 3597–3608.
Coates PJ, Lorimore SA, Lindsay KJ, Wright EG . Tissue-specific p53 responses to ionizing radiation and their genetic modification: the key to tissue-specific tumour susceptibility? J Pathol 2003; 201: 377–388.
Midgley CA, Owens B, Briscoe CV, Thomas DB, Lane DP, Hall PA . Coupling between gamma irradiation, p53 induction and the apoptotic response depends upon cell type in vivo. J Cell Sci 1995; 108: 1843–1848.
MacCallum DE, Hupp TR, Midgley CA, Stuart D, Campbell SJ, Harper A et al. The p53 response to ionising radiation in adult and developing murine tissues. Oncogene 1996; 13: 2575–2587.
Fei P, Bernhard EJ, El-Deiry WS . Tissue-specific induction of p53 targets in vivo. Cancer Res 2002; 62: 7316–7327.
Lorimore SA, Pragnell IB, Eckmann L, Wright EG . Synergistic interactions allow colony formation in vitro by murine haemopoietic stem cells. Leuk Res 1990; 14: 481–489.
Lotem J, Sachs L . Hematopoietic cells from mice deficient in wild-type p53 are more resistant to induction of apoptosis by some agents. Blood 1993; 82: 1092–1096.
Lorimore SA, Goodhead DT, Wright EG . The effect of p53 status on the radiosensitivity of haemopoietic stem cells. Cell Death Differ 1995; 2: 233–234.
Bergamaschi D, Samuels Y, O'Neil NJ, Trigiante G, Crook T, Hsieh JK et al. iASPP oncoprotein is a key inhibitor of p53 conserved from worm to human. Nat Genet 2003; 33: 162–167.
Budram-Mahadeo V, Morris PJ, Latchman DS . The Brn-3a transcription factor inhibits the pro-apoptotic effect of p53 and enhances cell cycle arrest by differentially regulating the activity of the p53 target genes encoding Bax and p21(CIP1/Waf1). Oncogene 2002; 21: 6123–6131.
Hudson CD, Morris PJ, Latchman DS, Budhram-Mahadeo VS . Brn-3a transcription factor blocks p53-mediated activation of proapoptotic target genes Noxa and Bax in vitro and in vivo to determine cell fate. J Biol Chem 2005; 280: 11851–11858.
Das S, Raj L, Zhao B, Kimura Y, Bernstein A, Aaronson SA et al. Hzf Determines cell survival upon genotoxic stress by modulating p53 transactivation. Cell 2007; 130: 624–637.
Aylon Y, Oren M . Living with p53, dying of p53. Cell 2007; 130: 597–600.
Espinosa JM, Verdun RE, Emerson BM . p53 functions through stress- and promoter-specific recruitment of transcription initiation components before and after DNA damage. Mol Cell 2003; 12: 1015–1027.
Morachis JM, Murawsky CM, Emerson BM . Regulation of the p53 transcriptional response by structurally diverse core promoters. Genes Dev 2010; 24: 135–147.
Lu X, Burbidge SA, Griffin S, Smith HM . Discordance between accumulated p53 protein level and its transcriptional activity in response to u.v. radiation. Oncogene 1996; 13: 413–418.
Chene P, Fuchs J, Bohn J, Garcia-Echeverria C, Furet P, Fabbro D . A small synthetic peptide, which inhibits the p53-hdm2 interaction, stimulates the p53 pathway in tumour cell lines. J Mol Biol 2000; 299: 245–253.
Bouvard V, Zaitchouk T, Vacher M, Duthu A, Canivet M, Choisy-Rossi C et al. Tissue and cell-specific expression of the p53-target genes: bax, fas, mdm2 and waf1/p21, before and following ionising irradiation in mice. Oncogene 2000; 19: 649–660.
Causevic M, Hislop RG, Kernohan NM, Carey FA, Kay RA, Steele RJ et al. Overexpression and poly-ubiquitylation of the DEAD-box RNA helicase p68 in colorectal tumours. Oncogene 2001; 20: 7734–7743.
Yang L, Lin C, Liu ZR . Phosphorylations of DEAD box p68 RNA helicase are associated with cancer development and cell proliferation. Mol Cancer Res 2005; 3: 355–363.
Wortham NC, Ahamed E, Nicol SM, Thomas RS, Periyasamy M, Jiang J et al. The DEAD-box protein p72 regulates ERalpha-/oestrogen-dependent transcription and cell growth, and is associated with improved survival in ERalpha-positive breast cancer. Oncogene 2009; 28: 4053–4064.
Hoy CA, Carswell C, Schimke RT . Bromodeoxyuridine/DNA analysis of replication in CHO cells after exposure to UV light. Mutat Res 1993; 290: 217–230.
Acknowledgements
We thank Colin Henderson for helpful discussions. This work was supported by grants from Cancer Research UK (C8745/A11216) and the Association for International Cancer Research (06–613).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Oncogene website
Supplementary information
Rights and permissions
About this article
Cite this article
Nicol, S., Bray, S., Derek Black, H. et al. The RNA helicase p68 (DDX5) is selectively required for the induction of p53-dependent p21 expression and cell-cycle arrest after DNA damage. Oncogene 32, 3461–3469 (2013). https://doi.org/10.1038/onc.2012.426
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2012.426
Keywords
This article is cited by
-
Role of the DEAD-box RNA helicase DDX5 (p68) in cancer DNA repair, immune suppression, cancer metabolic control, virus infection promotion, and human microbiome (microbiota) negative influence
Journal of Experimental & Clinical Cancer Research (2023)
-
The DEAD-box protein family of RNA helicases: sentinels for a myriad of cellular functions with emerging roles in tumorigenesis
International Journal of Clinical Oncology (2021)
-
DDX5 plays essential transcriptional and post-transcriptional roles in the maintenance and function of spermatogonia
Nature Communications (2019)
-
The DEAD-box RNA helicase 51 controls non-small cell lung cancer proliferation by regulating cell cycle progression via multiple pathways
Scientific Reports (2016)
-
DNA damage and the balance between survival and death in cancer biology
Nature Reviews Cancer (2016)