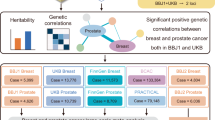
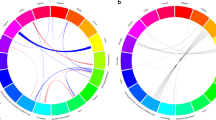
Abstract
Genome-wide association studies have recently identified ten common genetic variants associated with colorectal cancer susceptibility, several suggesting the involvement of components of the transforming growth factor beta (TGFβ) superfamily signalling pathway. To date, no causal sequence variants have been identified, and risk seems to be mediated through effects on gene regulation. Several markers are located close to poorly characterized genes or in gene deserts, raising challenges for elucidating mechanisms of susceptibility. Disease-associated common genetic variation offers the potential to refine risk stratification within populations and enable more targeted disease prevention strategies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
World Health Organization.World Cancer Report (eds Stewart B. W. & Kleihues P.) 13 (IARC, Lyon, 2003).
Barnetson, R. A. et al. Identification and survival of carriers of mutations in DNA mismatch-repair genes in colon cancer. N. Engl. J. Med. 354, 2751–2763 (2006).
Towler, B. P., Irwig, L., Glasziou, P., Weller, D. & Kewenter, J. Screening for colorectal cancer using the faecal occult blood test, hemoccult. Cochrane Database Syst. Rev. 2007, CD001216 (2000).
Jarvinen, H. J. et al. Controlled 15-year trial on screening for colorectal cancer in families with hereditary nonpolyposis colorectal cancer. Gastroenterology 118, 829–834 (2000).
Bost, B., de Vienne, D., Hospital, F., Moreau, L. & Dillmann, C. Genetic and nongenetic bases for the L-shaped distribution of quantitative trait loci effects. Genetics 157, 1773–1787 (2001).
Mackay, T. F. C. The genetic architecture of quantitative traits. Annu. Rev. Genet. 35, 303–339 (2001).
Foulkes, W. D. Inherited susceptibility to common cancers. N. Engl. J. Med. 359, 2143–2153 (2008).
Lichtenstein, P. et al. Environmental and heritable factors in the causation of cancer — analyses of cohorts of twins from Sweden, Denmark, and Finland. N. Engl. J. Med. 343, 78–85 (2000).
Yingling, J. M., Blanchard, K. L. & Sawyer, J. S. Development of TGF-β signalling inhibitors for cancer therapy. Nature Rev. Drug Discov. 3, 1011–1022 (2004).
The International HapMap Consortium.The International HapMap Project. Nature 426, 789–796 (2003).
Kruglyak, L. The road to genome-wide association studies. Nature Rev. Genet. 9, 314–318 (2008).
McCarthy, M. I. et al. Genome-wide association studies for complex traits: consensus, uncertainty and challenges. Nature Rev. Genet. 9, 356–369 (2008).
Tenesa, A. et al. Genome-wide association scan identifies a colorectal cancer susceptibility locus on 11q23 and replicates risk loci at 8q24 and 18q21. Nature Genet. 40, 631–637 (2008).
Antoniou, A. C. & Easton, D. F. Polygenic inheritance of breast cancer: implications for design of association studies. Genet. Epidemiol. 25, 190–202 (2003).
Tomlinson, I. et al. A genome-wide association scan of tag SNPs identifies a susceptibility variant for colorectal cancer at 8q24.21. Nature Genet. 39, 984–988 (2007).
Zanke, B. W. et al. Genome-wide association scan identifies a colorectal cancer susceptibility locus on chromosome 8q24. Nature Genet. 39, 989–994 (2007).
Broderick, P. et al. A genome-wide association study shows that common alleles of SMAD7 influence colorectal cancer risk. Nature Genet. 39, 1315–1317 (2007).
Jaeger, E. et al. Common genetic variants at the CRAC1 (HMPS) locus on chromosome 15q13.3 influence colorectal cancer risk. Nature Genet. 40, 26–28 (2008).
Tomlinson, I. P. et al. A genome-wide association study identifies colorectal cancer susceptibility loci on chromosomes 10p14 and 8q23.3. Nature Genet. 40, 623–630 (2008).
Houlston, R. S. et al. Meta-analysis of genome-wide association data identifies four new susceptibility loci for colorectal cancer. Nature Genet. 40, 1426–1435 (2008).
Howe, J. R. et al. The prevalence of MADH4 and BMPR1A mutations in juvenile polyposis and absence of BMPR2, BMPR1B, and ACVR1 mutations. J. Med. Genet. 41, 484–491 (2004).
Valle, L. et al. Germline allele-specific expression of TGFBR1 confers an increased risk of colorectal cancer. Science 321, 1361–1365 (2008).
Blobe, G. C., Schiemann, W. P. & Lodish, H. F. Role of transforming growth factor beta in human disease. N. Engl. J. Med. 342, 1350–1358 (2000).
Guilford, P. et al. E-cadherin germline mutations in familial gastric cancer. Nature 392, 402–405 (1998).
Okamoto, H., Yasui, K., Zhao, C., Arii, S. & Inazawa, J. PTK2 and EIF3S3 genes may be amplification targets at 8q23-q24 and are associated with large hepatocellular carcinomas. Hepatology 38, 1242–1249 (2003).
Savinainen, K. J. et al. Expression and copy number analysis of TRPS1, EIF3S3 and MYC genes in breast and prostate cancer. Br. J. Cancer 90, 1041–1046 (2004).
Shima, H. et al. Loss of heterozygosity on chromosome 10p14-p15 in colorectal carcinoma. Pathobiology 72, 220–224 (2005).
Haiman, C. A. et al. A common genetic risk factor for colorectal and prostate cancer. Nature Genet. 39, 954–956 (2007).
Ghoussaini, M. et al. Multiple loci with different cancer specificities within the 8q24 gene desert. J. Natl Cancer Inst. 100, 962–966 (2008).
Falconer, D. S. & Mackay, T. F. C. Introduction to Quantitative Genetics (Longman, 1996).
Wray, N. R., Goddard, M. E. & Visscher, P. M. Prediction of individual genetic risk to disease from genome-wide association studies. Genome Res. 17, 1520–1528 (2007).
Wang, E. T. et al. Alternative isoform regulation in human tissue transcriptomes. Nature 456, 470–476 (2008).
Bodmer, W. & Bonilla, C. Common and rare variants in multifactorial susceptibility to common diseases. Nature Genet. 40, 695–701 (2008).
Derynck, R. & Zhang, Y. E. Smad-dependent and Smad-independent pathways in TGF-β family signalling. Nature 425, 577–584 (2003).
Peck, J. W., Oberst, M., Bouker, K. B., Bowden, E. & Burbelo, P. D. The RhoA-binding protein, rhophilin-2, regulates actin cytoskeleton organization. J. Biol. Chem. 277, 43924–43932 (2002).
Chang, Y. W., Marlin, J. W., Chance, T. W. & Jakobi, R. RhoA mediates cyclooxygenase-2 signaling to disrupt the formation of adherens junctions and increase cellmotility. Cancer Res. 66, 11700–11708 (2006).
Howe, J. R. et al. Mutations in the SMAD4/DPC4 gene in juvenile polyposis. Science 280, 1086–1088 (1998).
Huang, S. C. et al. Genetic heterogeneity in familial juvenile polyposis. Cancer Res. 60, 6882–6885 (2000).
Woodford-Richens, K. L. et al. SMAD4 mutations in colorectal cancer probably occur before chromosomal instability, but after divergence of the microsatellite instability pathway. Proc. Natl Acad. Sci. USA 98, 9719–9723 (2001).
Kodach, L. L. et al. The bone morphogenetic protein pathway is inactivated in the majority of sporadic colorectal cancers. Gastroenterology 134, 1332–1341 (2008).
Parsons, R. et al. Microsatellite instability and mutations of the transforming growth factor β type II receptor gene in colorectal cancer. Cancer Res. 55, 5548–5550 (1995).
Pasche, B. et al. Somatic acquisition and signaling of TGFBR1*6A in cancer. JAMA 294, 1634–1646 (2005).
Visscher, P. M., Hill, W. G. & Wray, N. R. Heritability in the genomics era — concepts and misconceptions. Nature Rev. Genet. 9, 255–266 (2008).
Acknowledgements
Work that forms the basis of this discussion is supported by Cancer Research UK (C348/A8896, C48/A6361), and the Scottish Executive Chief Scientist's Office (CZB/4/449) — a centre grant from CORE as part of the Digestive Cancer Campaign. We acknowledge those in the Colon Cancer Genetics Group who have contributed to the work reviewed, particularly S. Farrington and H. Campbell, as well as our collaborators R. Houlston and I. Tomlinson and their groups. We thank R. Wilson and N. Cartwright, all of who worked on the COGS and SOCCS administrative teams, R. Cetnarskyj and the research nurse teams who recruited in Scotland, and all clinicians throughout Scotland at collaborating centres. We acknowledge N. Wray and P. Visscher for comments on BOX 1 and for sharing unpublished data on prediction models of genetic risk.
Author information
Authors and Affiliations
Related links
Glossary
- Excess familial risk
-
The increased risk of developing the disease in a relative of an affected individual. It is usually, and more appropriately, referred to for a specific type of relative. For example, the full-sibling relative risk (s) is the increased risk of developing a disease for a full sibling of an affected person compared with the risk of a person from the general population.
- Heritability
-
The proportion of phenotypic variance that is explained by inherited genetic factors.
- Liability scale
-
The assumed and unobserved normally distributed risk scale. In the case of a disease, those individuals with a liability score above a specific threshold will have the disease.
- Observed scale
-
For a disease trait, the observed scale of the phenotype is either disease or non-disease. The heritability in the observed scale depends on the disease prevalence.
- Odds ratio
-
A measure of the effect size. It is the odds of exposure (that is, a specific allele) among the cases divided by the odds of exposure among the controls. In casecontrol studies, the odds ratio is used as an approximation to the relative risk.
Rights and permissions
About this article
Cite this article
Tenesa, A., Dunlop, M. New insights into the aetiology of colorectal cancer from genome-wide association studies. Nat Rev Genet 10, 353–358 (2009). https://doi.org/10.1038/nrg2574
Issue Date:
DOI: https://doi.org/10.1038/nrg2574
This article is cited by
-
Kinome-wide analysis of the effect of statins in colorectal cancer
British Journal of Cancer (2021)
-
Comparison of natural orifice specimen extraction surgery and conventional laparoscopic-assisted resection in the treatment effects of low rectal cancer
Scientific Reports (2021)
-
TUFT1 Facilitates Metastasis, Stemness, and Vincristine Resistance in Colorectal Cancer via Activation of PI3K/AKT Pathway
Biochemical Genetics (2021)
-
Associations among dietary seaweed intake, c-MYC rs6983267 polymorphism, and risk of colorectal cancer in a Korean population: a case–control study
European Journal of Nutrition (2020)
-
Syndecan-1 suppresses cell growth and migration via blocking JAK1/STAT3 and Ras/Raf/MEK/ERK pathways in human colorectal carcinoma cells
BMC Cancer (2019)