Key Points
-
Since the identification in 1991 of repeat instability as a disease-causing mutation, gene-specific repeat instability is now known to be the mutational cause of at least 40 neurological, neurodegenerative and neuromuscular diseases.
-
Repeat instability is a dynamic mutation and is defined by both cis-elements and trans-factors.
-
Both germline (parent-to-offspring) and tissue-specific somatic instability occurs.
-
There are unique and common effectors for the instability of different repeat sequences, although each disease or locus is unique.
-
Replication, repair and recombination contribute to repeat instability, either separately or in combination.
-
Extrapolation of repeat instability data from model systems must take into account the limitations of the model and observations from human patients.
Abstract
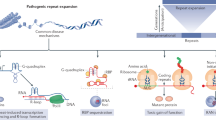
Disease-causing repeat instability is an important and unique form of mutation that is linked to more than 40 neurological, neurodegenerative and neuromuscular disorders. DNA repeat expansion mutations are dynamic and ongoing within tissues and across generations. The patterns of inherited and tissue-specific instability are determined by both gene-specific cis-elements and trans-acting DNA metabolic proteins. Repeat instability probably involves the formation of unusual DNA structures during DNA replication, repair and recombination. Experimental advances towards explaining the mechanisms of repeat instability have broadened our understanding of this mutational process. They have revealed surprising ways in which metabolic pathways can drive or protect from repeat instability.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Pearson, C. E. Slipping while sleeping? Trinucleotide repeat expansions in germ cells. Trends Mol. Med. 9, 490–495 (2003).
Yoon, S. R., Dubeau, L., de Young, M., Wexler, N. S. & Arnheim, N. Huntington disease expansion mutations in humans can occur before meiosis is completed. Proc. Natl Acad. Sci. USA 100, 8834–8838 (2003). By using laser-capture microdissection and repeat-length analysis by single-cell PCR the authors advanced our understanding of the timing (pre-meiotic) and mechanism of CAG expansions and parent-of-origin (paternal) expansion biases.
Mangiarini, L. et al. Instability of highly expanded CAG repeats in mice transgenic for the Huntington's disease mutation. Nature Genet. 15, 197–200 (1997).
Zhang, Y., Monckton, D. G., Siciliano, M. J., Connor, T. H. & Meistrich, M. L. Age and insertion site dependence of repeat number instability of a human DM1 transgene in individual mouse sperm. Hum. Mol. Genet. 11, 791–798 (2002).
Savouret, C. et al. CTG repeat instability and size variation timing in DNA repair-deficient mice. EMBO J. 22, 2264–2273 (2003).
Savouret, C. et al. MSH2-dependent germinal CTG repeat expansions are produced continuously in spermatogonia from DM1 transgenic mice. Mol. Cell. Biol. 24, 629–637 (2004). An important study on the timing (pre-meiotic) of male germline expansions/contractions and the involvement of the mismatch repair protein MSH2.
Kaytor, M. D., Burright, E. N., Duvick, L. A., Zoghbi, H. Y. & Orr, H. T. Increased trinucleotide repeat instability with advanced maternal age. Hum. Mol. Genet. 6, 2135–2139 (1997).
Malter, H. E. et al. Characterization of the full fragile X syndrome mutation in fetal gametes. Nature Genet. 15, 165–169 (1997). The authors provide an insight into the timing of the parent-of-origin effect, including maternal expansion bias and paternal contraction bias, and the potential role of CpG methylation in CGG instability.
Moutou, C., Vincent, M. C., Biancalana, V. & Mandel, J. L. Transition from premutation to full mutation in fragile X syndrome is likely to be prezygotic. Hum. Mol. Genet. 6, 971–979 (1997).
De Temmerman, N. et al. Intergenerational instability of the expanded CTG repeat in the DMPK gene: studies in human gametes and preimplantation embryos. Am. J. Hum. Genet. 75, 325–329 (2004). This paper gives an insight into the timing and mechanism of parent-of-origin maternal expansion bias of a CTG repeat and the timing of post-fertilization CTG instability.
Lenzi, M. L. et al. Extreme heterogeneity in the molecular events leading to the establishment of chiasmata during meiosis I in human oocytes. Am. J. Hum. Genet. 76, 112–127 (2005).
Martorell, L., Johnson, K., Boucher, C. A. & Baiget, M. Somatic instability of the myotonic dystrophy (CTG)n repeat during human fetal development. Hum. Mol. Genet. 6, 877–880 (1997). An important insight into the timing of somatic CTG instability and the tissue specificity of this instability, some of which can continue throughout the lifetime of the adult.
Reyniers, E. et al. Postmortem examination of two fragile X brothers with an FMR1 full mutation. Am. J. Med. Genet. 84, 245–249 (1999).
Nichol Edamura, K., Leonard, M. R. & Pearson, C. E. Role of replication and CpG methylation in fragile X syndrome CGG deletions in primate cells. Am. J. Hum. Genet. 76, 302–311 (2005).
Carbonell, P. et al. FRAXE mutation analysis in three Spanish families. Am. J. Med. Genet. 64, 434–440 (1996).
Moseley, M. L. et al. SCA8 CTG repeat: en masse contractions in sperm and intergenerational sequence changes may play a role in reduced penetrance. Hum. Mol. Genet. 9, 2125–2130 (2000).
Monckton, D. G., Wong, L. J., Ashizawa, T. & Caskey, C. T. Somatic mosaicism, germline expansions, germline reversions and intergenerational reductions in myotonic dystrophy males: small pool PCR analyses. Hum. Mol. Genet. 4, 1–8 (1995). This study uses the ultra-sensitive technique of small pool PCR to qualitatively and quantitatively assess CTG-length heterogeneity, allowing the detection of the products of rare events. It also provides an insight into germline contraction events.
De Michele, G. et al. Parental gender, age at birth and expansion length influence GAA repeat intergenerational instability in the X25 gene: pedigree studies and analysis of sperm from patients with Friedreich's ataxia. Hum. Mol. Genet. 7, 1901–1906 (1998).
Lemmers, R. J. et al. Mechanism and timing of mitotic rearrangements in the subtelomeric D4Z4 repeat involved in facioscapulohumeral muscular dystrophy. Am. J. Hum. Genet. 75, 44–53 (2004).
Benitez, J. et al. Somatic stability in chorionic villi samples and other Huntington fetal tissues. Hum. Genet. 96, 229–232 (1995).
Jedele, K. B. et al. Spinal and bulbar muscular atrophy (SBMA): somatic stability of an expanded CAG repeat in fetal tissues. Clin. Genet. 54, 148–151 (1998).
Devys, D. et al. Analysis of full fragile X mutations in fetal tissues and monozygotic twins indicate that abnormal methylation and somatic heterogeneity are established early in development. Am. J. Med. Genet. 43, 208–216 (1992).
Taylor, A. K. et al. Tissue heterogeneity of the FMR1 mutation in a high-functioning male with fragile X syndrome. Am. J. Med. Genet. 84, 233–239 (1999). Together with reference 13, this study contributed to the understanding of the timing and tissue-specificity of germline and somatic mosaicism and CpG methylation mosaicism. It indicates a role of CpG methylation in CGG instability.
Pollard, L. M. et al. Replication-mediated instability of the GAA triplet repeat mutation in Friedreich ataxia. Nucleic Acids Res. 32, 5962–5971 (2004).
Wohrle, D. et al. Heterogeneity of DM kinase repeat expansion in different fetal tissues and further expansion during cell proliferation in vitro: evidence for a casual involvement of methyl-directed DNA mismatch repair in triplet repeat stability. Hum. Mol. Genet. 4, 1147–1153 (1995).
Thornton, C. A., Johnson, K. & Moxley, R. T. 3rd. Myotonic dystrophy patients have larger CTG expansions in skeletal muscle than in leukocytes. Ann. Neurol. 35, 104–107 (1994).
Zatz, M. et al. Analysis of the CTG repeat in skeletal muscle of young and adult myotonic dystrophy patients: when does the expansion occur? Hum. Mol. Genet. 4, 401–406 (1995).
Martorell, L., Martinez, J. M., Carey, N., Johnson, K. & Baiget, M. Comparison of CTG repeat length expansion and clinical progression of myotonic dystrophy over a five year period. J. Med. Genet. 32, 593–596 (1995).
Kinoshita, M. et al. A patient with myotonic dystrophy type 1 (DM1) accompanied by laryngeal and renal cell carcinomas had a small CTG triplet repeat expansion but no somatic instability in normal tissues. Intern. Med. 41, 312–318 (2002).
Osanai, R., Kinoshita, M., Hirose, K., Homma, T. & Kawabata, I. CTG triplet repeat expansion in a laryngeal carcinoma from a patient with myotonic dystrophy. Muscle Nerve 23, 804–806 (2000).
van Den Broek, W. J. et al. Somatic expansion behaviour of the (CTG)n repeat in myotonic dystrophy knock-in mice is differentially affected by Msh3 and Msh6 mismatch-repair proteins. Hum. Mol. Genet. 11, 191–198 (2002).
Lia, A. S. et al. Somatic instability of the CTG repeat in mice transgenic for the myotonic dystrophy region is age dependent but not correlated to the relative intertissue transcription levels and proliferative capacities. Hum. Mol. Genet. 7, 1285–1291 (1998).
Gomes-Pereira, M., Fortune, M. T. & Monckton, D. G. Mouse tissue culture models of unstable triplet repeats: in vitro selection for larger alleles, mutational expansion bias and tissue specificity, but no association with cell division rates. Hum. Mol. Genet. 10, 845–854 (2001).
Nouspikel, T. & Hanawalt, P. C. DNA repair in terminally differenetiated cells. DNA Repair 1, 59–75 (2002).
Vinson, R. K. & Hales, B. F. DNA repair during organogenesis. Mutat. Res. 509, 79–91 (2002).
Cleary, J. D. & Pearson, C. E. The contribution of cis-elements to disease-associated repeat instability: Clinical and experimental evidence. Cytogenet. Genome Res. 100, 25–55 (2003).
Watase, K., Venken, K. J., Sun, Y., Orr, H. T. & Zoghbi, H. Y. Regional differences of somatic CAG repeat instability do not account for selective neuronal vulnerability in a knock-in mouse model of SCA1. Hum. Mol. Genet. 12, 2789–2795 (2003).
Sato, T. et al. Transgenic mice harboring a full-length human mutant DRPLA gene exhibit age-dependent intergenerational and somatic instabilities of CAG repeats comparable with those in DRPLA patients. Hum. Mol. Genet. 8, 99–106 (1999).
Fortune, M. T., Vassilopoulos, C., Coolbaugh, M. I., Siciliano, M. J. & Monckton, D. G. Dramatic, expansion-biased, age-dependent, tissue-specific somatic mosaicism in a transgenic mouse model of triplet repeat instability. Hum. Mol. Genet. 9, 439–445 (2000).
Al-Mahdawi, S. et al. GAA repeat instability in Friedreich ataxia YAC transgenic mice. Genomics 84, 301–310 (2004).
Wheeler, V. C. et al. Mismatch repair gene Msh2 modifies the timing of early disease in HdhQ111 striatum. Hum. Mol. Genet. 12, 273–281 (2003).
Dobbing, J. & Sands, J. Comparative aspects of the brain growth spurt. Early Hum. Dev. 3, 79–83 (1979).
Maciel, P., Lopes-Cendes, I., Kish, S., Sequeiros, J. & Rouleau, G. A. Mosaicism of the CAG repeat in CNS tissue in relation to age at death in spinocerebellar ataxia type 1 and Machado–Joseph disease patients. Am. J. Hum. Genet. 60, 993–996 (1997).
Kennedy, L. et al. Dramatic tissue-specific mutation length increases are an early molecular event in Huntington disease pathogenesis. Hum. Mol. Genet. 12, 3359–3367 (2003).
Takano, H. et al. Somatic mosaicism of expanded CAG repeats in brains of patients with dentatorubral-pallidoluysian atrophy: cellular population-dependent dynamics of mitotic instability. Am. J. Hum. Genet. 58, 1212–1222 (1996).
Hashida, H. et al. Single cell analysis of CAG repeat in brains of dentatorubral-pallidoluysian atrophy (DRPLA). J. Neurol. Sci. 190, 87–93 (2001).
Watanabe, H. et al. Differential somatic CAG repeat instability in variable brain cell lineage in dentatorubral pallidoluysian atrophy (DRPLA): a laser-captured microdissection (LCM)-based analysis. Hum. Genet. 107, 452–457 (2000). In references 46 and 47 the authors used high-selective laser-capture microdissection to study neuronal and non-neuronal somatic instability in patients' brains, and the timing and tissue-specificity of repeat expansions and contractions.
Curtis, M. A. et al. Increased cell proliferation and neurogenesis in the adult human Huntington's disease brain. Proc. Natl Acad. Sci. USA 100, 9023–9027 (2003).
Kahlem, P. & Djian, P. The expanded CAG repeat associated with juvenile Huntington disease shows a common origin of most or all neurons and glia in human cerebrum. Neurosci. Lett. 286, 203–207 (2000).
Kennedy, L. & Shelbourne, P. F. Dramatic mutation instability in HD mouse striatum: does polyglutamine load contribute to cell-specific vulnerability in Huntington's disease? Hum. Mol. Genet. 9, 2539–2544 (2000).
Lopes-Cendes, I. et al. Somatic mosaicism in the central nervous system in spinocerebellar ataxia type 1 and Machado–Joseph disease. Ann. Neurol. 40, 199–206 (1996).
Manley, K., Shirley, T. L., Flaherty, L. & Messer, A. Msh2 deficiency prevents in vivo somatic instability of the CAG repeat in Huntington disease transgenic mice. Nature Genet. 23, 471–473 (1999). The authors propose the active contribution of the MSH2 mismatch repair protein to the mutation of repeat sequences — a model that contrasts with this protein's role in maintaining genome integrity. They also reveal the effects of tissue-specificity of MSH2 on somatic CAG instability.
Panigrahi, G. B., Lau, R., Montgomery, S. E., Leonard, M. R. & Pearson, C. E. Slipped (CTG)*(CAG) repeats can be correctly repaired, escape repair or undergo error-prone repair. Nature Struct. Mol. Biol. 12, 654–662 (2005). This paper provides direct evidence for the processing of slipped DNAs, through a potential mutagenic intermediate of instability, and reveals mechanistic pathways through which this processing might lead to repeat stability, expansion and contraction.
Hashida, H., Goto, J., Kurisaki, H., Mizusawa, H. & Kanazawa, I. Brain regional differences in the expansion of a CAG repeat in the spinocerebellar ataxias: dentatorubral-pallidoluysian atrophy, Machado–Joseph disease, and spinocerebellar ataxia type 1. Ann. Neurol. 41, 505–511 (1997).
Tanaka, F. et al. Tissue-specific somatic mosaicism in spinal and bulbar muscular atrophy is dependent on CAG-repeat length and androgen receptor-gene expression level. Am. J. Hum. Genet. 65, 966–973 (1999).
Ansved, T., Lundin, A. & Anvret, M. Larger CAG expansions in skeletal muscle compared with lymphocytes in Kennedy disease but not in Huntington disease. Neurology 51, 1442–1444 (1998).
Montermini, L., Kish, S. J., Jiralerspong, S., Lamarche, J. B. & Pandolfo, M. Somatic mosaicism for Friedreich's ataxia GAA triplet repeat expansions in the central nervous system. Neurology 49, 606–610 (1997).
Tassone, F., Hagerman, R. J., Gane, L. W. & Taylor, A. K. Strong similarities of the FMR1 mutation in multiple tissues: postmortem studies of a male with a full mutation and a male carrier of a premutation. Am. J. Med. Genet. 84, 240–244 (1999).
Chong, S. S. et al. Gametic and somatic tissue-specific heterogeneity of the expanded SCA1 CAG repeat in spinocerebellar ataxia type 1. Nature Genet. 10, 344–350 (1995).
Pearson, C. E. et al. Interruptions in the triplet repeats of SCA1 and FRAXA reduce the propensity and complexity of slipped strand DNA (S-DNA) formation. Biochemistry 37, 2701–2708 (1998).
Brock, G. J., Anderson, N. H. & Monckton, D. G. Cis-acting modifiers of expanded CAG/CTG triplet repeat expandability: associations with flanking GC content and proximity to CpG islands. Hum. Mol. Genet. 8, 1061–1067 (1999).
Gourdon, G., Dessen, P., Lia, A. S., Junien, C. & Hofmann-Radvanyi, H. Intriguing association between disease associated unstable trinucleotide repeat and CpG island. Ann. Genet. 40, 73–77 (1997).
Monckton, D. G., Coolbaugh, M. I., Ashizawa, K. T., Siciliano, M. J. & Caskey, C. T. Hypermutable myotonic dystrophy CTG repeats in transgenic mice. Nature Genet. 15, 193–196 (1997).
Gourdon, G. et al. Moderate intergenerational and somatic instability of a 55-CTG repeat in transgenic mice. Nature Genet. 15, 190–192 (1997).
La Spada, A. R. et al. Androgen receptor YAC transgenic mice carrying CAG 45 alleles show trinucleotide repeat instability. Hum. Mol. Genet. 7, 959–967 (1998).
Libby, R. T. et al. Genomic context drives SCA7 CAG repeat instability, while expressed SCA7 cDNAs are intergenerationally and somatically stable in transgenic mice. Hum. Mol. Genet. 12, 41–50 (2003). This article demonstrates the need for flanking sequence for the active instability to occur in both germline and somatic tissues and confirms the implication (from haplotype analysis) that cis -elements contribute to repeat instability.
Seznec, H. et al. Transgenic mice carrying large human genomic sequences with expanded CTG repeat mimic closely the DM CTG repeat intergenerational and somatic instability. Hum. Mol. Genet. 9, 1185–1194 (2000).
Sutcliffe, J. S. et al. DNA methylation represses FMR-1 transcription in fragile X syndrome. Hum. Mol. Genet. 1, 397–400 (1992).
Willemsen, R., Bontekoe, C. J., Severijnen, L. A. & Oostra, B. A. Timing of the absence of FMR1 expression in full mutation chorionic villi. Hum. Genet. 110, 601–605 (2002).
Wohrle, D. et al. Unusual mutations in high functioning fragile X males: apparent instability of expanded unmethylated CGG repeats. J. Med. Genet. 35, 103–111 (1998).
Burman, R. W., Popovich, B. W., Jacky, P. B. & Turker, M. S. Fully expanded FMR1 CGG repeats exhibit a length- and differentiation-dependent instability in cell hybrids that is independent of DNA methylation. Hum. Mol. Genet. 8, 2293–2302 (1999).
Salat, U., Bardoni, B., Wohrle, D. & Steinbach, P. Increase of FMRP expression, raised levels of FMR1 mRNA, and clonal selection in proliferating cells with unmethylated fragile X repeat expansions: a clue to the sex bias in the transmission of full mutations? J. Med. Genet. 37, 842–850 (2000).
Nichol, K. & Pearson, C. E. CpG methylation modifies the genetic stability of cloned repeat sequences. Genome Res. 12, 1246–1256 (2002).
Gorbunova, V., Seluanov, A., Mittelman, D. & Wilson, J. H. Genome-wide demethylation destabilizes CTG. CAG trinucleotide repeats in mammalian cells. Hum. Mol. Genet. 13, 2979–2989 (2004).
Wang, Y. H., Amirhaeri, S., Kang, S., Wells, R. D. & Griffith, J. D. Preferential nucleosome assembly at DNA triplet repeats from the myotonic dystrophy gene. Science 265, 669–671 (1994).
Mulvihill, D. J., Edamura, K. N., Hagerman, K. A., Pearson, C. E. & Wang, Y. H. Effect of CAT or AGG interruptions and CpG methylation on nucleosome assembly upon trinucleotide repeats on spinocerebellar ataxia type 1 and fragile X syndrome. J. Biol. Chem. 280, 4498–4503 (2005).
Otten, A. D. & Tapscott, S. J. Triplet repeat expansion in myotonic dystrophy alters the adjacent chromatin structure. Proc. Natl Acad. Sci. USA 92, 5465–5469 (1995).
Filippova, G. N. et al. CTCF-binding sites flank CTG/CAG repeats and form a methylation-sensitive insulator at the DM1 locus. Nature Genet. 28, 335–343 (2001).
Cleary J. D., Nichol Edamura, K. & Pearson C. E. The complex nature of trinucleotide repeat instability. Chemtracts Biochem. Mol. Biol. 17, 663–676 (2004).
Martorell, L. et al. Progression of somatic CTG repeat length heterogeneity in the blood cells of myotonic dystrophy patients. Hum. Mol. Genet. 7, 307–312 (1998).
Cleary, J. D., Nichol, K., Wang, Y. H. & Pearson, C. E. Evidence of cis-acting factors in replication-mediated trinucleotide repeat instability in primate cells. Nature Genet. 31, 37–46 (2002). This work reveals the complex influence of DNA replication on CTG instability. Along with works such as reference 87, it highlights the importance of replication-fork perturbation.
Yang, Z., Lau, R., Marcadier, J. L., Chitayat, D. & Pearson, C. E. Replication inhibitors modulate instability of an expanded trinucleotide repeat at the myotonic dystrophy type 1 disease locus in human cells. Am. J. Hum. Genet. 73, 1092–1105 (2003).
Freudenreich, C. H. & Lahiri, M. Structure-forming CAG/CTG repeat sequences are sensitive to breakage in the absence of Mrc1 checkpoint function and S-phase checkpoint signaling: implications for trinucleotide repeat expansion diseases. Cell Cycle 3, 1370–1374 (2004).
Nenguke, T., Aladjem, M. I., Gusella, J. F., Wexler, N. S. & Arnheim, N. Candidate DNA replication initiation regions at human trinucleotide repeat disease loci. Hum. Mol. Genet. 12, 1021–1028 (2003).
Cleary, J. D. & Pearson, C. E. Replication fork dynamics and dynamic mutations: the fork-shift model of repeat instability. Trends Genet. 21, 272–280 (2005).
Kang, S., Ohshima, K., Shimizu, M., Amirhaeri, S. & Wells, R. D. Pausing of DNA synthesis in vitro at specific loci in CTG and CGG triplet repeats from human hereditary disease genes. J. Biol. Chem. 270, 27014–27021 (1995).
Samadashwily, G. M., Raca, G. & Mirkin, S. M. Trinucleotide repeats affect DNA replication in vivo. Nature Genet. 17, 298–304 (1997).
Pelletier, R., Krasilnikova, M. M., Samadashwily, G. M., Lahue, R. & Mirkin, S. M. Replication and expansion of trinucleotide repeats in yeast. Mol. Cell. Biol. 23, 1349–1357 (2003).
Krasilnikova, M. M. & Mirkin, S. M. Replication stalling at Friedreich's ataxia (GAA)n repeats in vivo. Mol. Cell. Biol. 24, 2286–2295 (2004).
Freudenreich, C. H., Kantrow, S. M. & Zakian, V. A. Expansion and length-dependent fragility of CTG repeats in yeast. Science 279, 853–856 (1998).
Marcadier, J. L. & Pearson, C. E. Fidelity of primate cell repair of a double-strand break within a (CTG)•(CAG) tract. Effect of slipped DNA structures. J. Biol. Chem. 278, 33848–33856 (2003).
Hebert, M. L., Spitz, L. A. & Wells, R. D. DNA double-strand breaks induce deletion of CTG. CAG repeats in an orientation-dependent manner in Escherichia coli. J. Mol. Biol. 336, 655–672 (2004).
Spiro, C. et al. Inhibition of FEN-1 processing by DNA secondary structure at trinucleotide repeats. Mol. Cell 4, 1079–1085 (1999).
Henricksen, L. A., Tom, S., Liu, Y. & Bambara, R. A. Inhibition of flap endonuclease 1 by flap secondary structure and relevance to repeat sequence expansion. J. Biol. Chem. 275, 16420–16427 (2000).
Lee, S. & Park, M. S. Human FEN-1 can process the 5′-flap DNA of CTG/CAG triplet repeat derived from human genetic diseases by length and sequence dependent manner. Exp. Mol. Med. 34, 313–317 (2002).
Callahan, J. L., Andrews, K. J., Zakian, V. A. & Freudenreich, C. H. Mutations in yeast replication proteins that increase CAG/CTG expansions also increase repeat fragility. Mol. Cell. Biol. 23, 7849–7860 (2003).
Liu, Y., Zhang, H., Veeraraghavan, J., Bambara, R. A. & Freudenreich, C. H. Saccharomyces cerevisiae flap endonuclease 1 uses flap equilibration to maintain triplet repeat stability. Mol. Cell. Biol. 24, 4049–4064 (2004).
Spiro, C. & McMurray, C. T. Nuclease-deficient FEN-1 blocks Rad51/BRCA1-mediated repair and causes trinucleotide repeat instability. Mol. Cell. Biol. 23, 6063–6074 (2003).
Jackson, S. M. et al. A SCA7 CAG/CTG repeat expansion is stable in Drosophila melanogaster despite modulation of genomic context and gene dosage. Gene 347, 35–41 (2005).
Wang, W. & Bambara, R. A. Human Bloom protein stimulates flap endonuclease 1 activity by resolving DNA secondary structure. J. Biol. Chem. 280, 5391–5399 (2005).
Kao, H. I., Veeraraghavan, J., Polaczek, P., Campbell, J. L. & Bambara, R. A. On the roles of Saccharomyces cerevisiae Dna2p and Flap endonuclease 1 in Okazaki fragment processing. J Biol Chem 279, 15014–15024 (2004).
Ireland, M. J., Reinke, S. S. & Livingston, D. M. The impact of lagging strand replication mutations on the stability of CAG repeat tracts in yeast. Genetics 155, 1657–1665 (2000).
Hashem, V. I. et al. Chemotherapeutic deletion of CTG repeats in lymphoblast cells from DM1 patients. Nucleic Acids Res. 32, 6334–6346 (2004).
Otto, C. J., Almqvist, E., Hayden, M. R. & Andrew, S. E. The 'flap' endonuclease gene FEN1 is excluded as a candidate gene implicated in the CAG repeat expansion underlying Huntington disease. Clin. Genet. 59, 122–127 (2001).
Kramer, P. R., Pearson, C. E. & Sinden, R. R. Stability of triplet repeats of myotonic dystrophy and fragile X loci in human mutator mismatch repair cell lines. Hum. Genet. 98, 151–157 (1996).
Goellner, G. M. et al. Different mechanisms underlie DNA instability in Huntington disease and colorectal cancer. Am. J. Hum. Genet. 60, 879–890 (1997).
Grewal, R. P. Neurodegeneration in xeroderma pigmentosum: a trinucleotide repeat mutation analysis. J. Neurol. Sci. 163, 183–186 (1999).
Kovtun, I. V. & McMurray, C. T. Trinucleotide expansion in haploid germ cells by gap repair. Nature Genet. 27, 407–411 (2001).
Owen, B. A. et al. (CAG)n-hairpin DNA binds to Msh2–Msh3 and changes properties of mismatch recognition. Nature Struct. Mol. Biol. 12, 663–670 (2005).
Pearson, C. E., Ewel, A., Acharya, S., Fishel, R. A. & Sinden, R. R. Human MSH2 binds to trinucleotide repeat DNA structures associated with neurodegenerative diseases. Hum. Mol. Genet. 6, 1117–1123 (1997).
Wilson, T. E. & Lieber, M. R. Efficient processing of DNA ends during yeast nonhomologous end joining. Evidence for a DNA polymeraseβ (Pol4)-dependent pathway. J. Biol. Chem. 274, 23599–23609 (1999).
Lin, D. P. et al. An Msh2 point mutation uncouples DNA mismatch repair and apoptosis. Cancer Res. 64, 517–522 (2004).
Plotz, G., Raedle, J., Brieger, A., Trojan, J. & Zeuzem, S. hMutSα forms an ATP-dependent complex with hMutLα and hMutLβ on DNA. Nucleic Acids Res. 30, 711–718 (2002).
Gomes-Pereira, M., Fortune, M. T., Ingram, L., McAbney, J. P. & Monckton, D. G. Pms2 is a genetic enhancer of trinucleotide CAG•CTG repeat somatic mosaicism: implications for the mechanism of triplet repeat expansion. Hum. Mol. Genet. 13, 1815–1825 (2004).
Hartenstine, M. J., Goodman, M. F. & Petruska, J. Base stacking and even/odd behavior of hairpin loops in DNA triplet repeat slippage and expansion with DNA polymerase. J. Biol. Chem. 275, 18382–18390 (2000).
Gaudreault, I., Guay, D. & Lebel, M. YB-1 promotes strand separation in vitro of duplex DNA containing either mispaired bases or cisplatin modifications, exhibits endonucleolytic activities and binds several DNA repair proteins. Nucleic Acids Res. 32, 316–327 (2004).
Yang, Q. et al. The mismatch DNA repair heterodimer, hMSH2/6, regulates BLM helicase. Oncogene 23, 3749–3756 (2004).
Cheok, C. F., Wu, L., Garcia, P. L., Janscak, P. & Hickson, I. D. The Bloom's syndrome helicase promotes the annealing of complementary single-stranded DNA. Nucleic Acids Res. 33, 3932–3941 (2005).
Doherty, K. M. et al. RECQ1 helicase interacts with human mismatch repair factors that regulate genetic recombination. J. Biol. Chem. 280, 28085–28094 (2005).
Sharma, S. et al. Biochemical analysis of the DNA unwinding and strand annealing activities catalyzed by human RECQ1. J. Biol. Chem. 280, 28072–28084 (2005).
Schmidt, K. H., Abbott, C. M. & Leach, D. R. Two opposing effects of mismatch repair on CTG repeat instability in Escherichia coli. Mol. Microbiol. 35, 463–471 (2000).
Schweitzer, J. K. & Livingston, D. M. Destabilization of CAG trinucleotide repeat tracts by mismatch repair mutations in yeast. Hum. Mol. Genet. 6, 349–355 (1997).
Jaworski, A. et al. Mismatch repair in Escherichia coli enhances instability of (CTG)n triplet repeats from human hereditary diseases. Proc. Natl Acad. Sci. USA 92, 11019–11023 (1995).
Schweitzer, J. K. & Livingston, D. M. Expansions of CAG repeat tracts are frequent in a yeast mutant defective in Okazaki fragment maturation. Hum. Mol. Genet. 7, 69–74 (1998).
Jankowski, C., Nasar, F. & Nag, D. K. Meiotic instability of CAG repeat tracts occurs by double-strand break repair in yeast. Proc. Natl Acad. Sci. USA 97, 2134–2139 (2000).
Richard, G. F., Goellner, G. M., McMurray, C. T. & Haber, J. E. Recombination-induced CAG trinucleotide repeat expansions in yeast involve the MRE11–RAD50–XRS2 complex. EMBO J. 19, 2381–2390 (2000).
Meservy, J. L. et al. Long CTG tracts from the myotonic dystrophy gene induce deletions and rearrangements during recombination at the APRT locus in CHO cells. Mol. Cell. Biol. 23, 3152–3162 (2003).
Paques, F., Leung, W. Y. & Haber, J. E. Expansions and contractions in a tandem repeat induced by double-strand break repair. Mol. Cell. Biol. 18, 2045–2054 (1998).
Parniewski, P., Bacolla, A., Jaworski, A. & Wells, R. D. Nucleotide excision repair affects the stability of long transcribed (CTG*CAG) tracts in an orientation-dependent manner in Escherichia coli. Nucleic Acids Res. 27, 616–623 (1999).
Oussatcheva, E. A., Hashem, V. I., Zou, Y., Sinden, R. R. & Potaman, V. N. Involvement of the nucleotide excision repair protein UvrA in instability of CAG*CTG repeat sequences in Escherichia coli. J. Biol. Chem. 276, 30878–30884 (2001).
Bowater, R. P., Jaworski, A., Larson, J. E., Parniewski, P. & Wells, R. D. Transcription increases the deletion frequency of long CTG•CAG triplet repeats from plasmids in Escherichia coli. Nucleic Acids Res. 25, 2861–2868 (1997).
Parniewski, P., Jaworski, A., Wells, R. D. & Bowater, R. P. Length of CTG•CAG repeats determines the influence of mismatch repair on genetic instability. J. Mol. Biol. 299, 865–874 (2000).
Schumacher, S., Pinet, I. & Bichara, M. Modulation of transcription reveals a new mechanism of triplet repeat instability in Escherichia coli. J. Mol. Biol. 307, 39–49 (2001).
Stojic, L., Brun, R. & Jiricny, J. Mismatch repair and DNA damage signalling. DNA Repair (Amst.) 3, 1091–1101 (2004).
Lee, G. S., Brandt, V. L. & Roth, D. B. B cell development leads off with a base hit: dU:dG mismatches in class switching and hypermutation. Mol. Cell 16, 505–508 (2004).
O'Hoy, K. L. et al. Reduction in size of the myotonic dystrophy trinucleotide repeat mutation during transmission. Science 259, 809–812 (1993).
van den Ouweland, A. M. et al. Loss of mutation at the FMR1 locus through multiple exchanges between maternal X chromosomes. Hum. Mol. Genet. 3, 1823–1827 (1994).
Miret, J. J., Pessoa-Brandao, L. & Lahue, R. S. Orientation-dependent and sequence-specific expansions of CTG/CAG trinucleotide repeats in Saccharomyces cerevisiae. Proc. Natl Acad. Sci. USA 95, 12438–12443 (1998).
Jakupciak, J. P. & Wells, R. D. Genetic instabilities in (CTG•CAG) repeats occur by recombination. J. Biol. Chem. 274, 23468–23479 (1999).
Cohen, H., Sears, D. D., Zenvirth, D., Hieter, P. & Simchen, G. Increased instability of human CTG repeat tracts on yeast artificial chromosomes during gametogenesis. Mol. Cell. Biol. 19, 4153–4158 (1999).
Moore, H., Greenwell, P. W., Liu, C. P., Arnheim, N. & Petes, T. D. Triplet repeats form secondary structures that escape DNA repair in yeast. Proc. Natl Acad. Sci. USA 96, 1504–1509 (1999).
Jankowski, C. & Nag, D. K. Most meiotic CAG repeat tract-length alterations in yeast are SPO11 dependent. Mol. Genet. Genomics 267, 64–70 (2002).
Brown, L. Y. & Brown, S. A. Alanine tracts: the expanding story of human illness and trinucleotide repeats. Trends Genet. 20, 51–58 (2004).
Stead, J. D. & Jeffreys, A. J. Allele diversity and germline mutation at the insulin minisatellite. Hum. Mol. Genet. 9, 713–723 (2000).
van der Maarel, S. M. et al. De novo facioscapulohumeral muscular dystrophy: frequent somatic mosaicism, sex-dependent phenotype, and the role of mitotic transchromosomal repeat interaction between chromosomes 4 and 10. Am. J. Hum. Genet. 66, 26–35 (2000).
Cancel, G. et al. Somatic mosaicism of the CAG repeat expansion in spinocerebellar ataxia type 3/Machado–Joseph disease. Hum. Mutat. 11, 23–27 (1998).
Telenius, H. et al. Molecular analysis of juvenile Huntington disease: the major influence on (CAG)n repeat length is the sex of the affected parent. Hum. Mol. Genet. 2, 1535–1540 (1993).
Onodera, O. et al. Progressive atrophy of cerebellum and brainstem as a function of age and the size of the expanded CAG repeats in the MJD1 gene in Machado–Joseph disease. Ann. Neurol. 43, 288–296 (1998).
Li, J. Y., Plomann, M. & Brundin, P. Huntington's disease: a synaptopathy? Trends Mol. Med. 9, 414–420 (2003).
Mhatre, A. N. et al. Reduced transcriptional regulatory competence of the androgen receptor in X-linked spinal and bulbar muscular atrophy. Nature Genet. 5, 184–188 (1993).
Seo, H., Sonntag, K. C. & Isacson, O. Generalized brain and skin proteasome inhibition in Huntington's disease. Ann. Neurol. 56, 319–328 (2004).
Sathasivam, K. et al. Centrosome disorganization in fibroblast cultures derived from R6/2 Huntington's disease (HD) transgenic mice and HD patients. Hum. Mol. Genet. 10, 2425–2435 (2001).
Lin, X., Antalffy, B., Kang, D., Orr, H. T. & Zoghbi, H. Y. Polyglutamine expansion down-regulates specific neuronal genes before pathologic changes in SCA1. Nature Neurosci. 3, 157–163 (2000).
Caramins, M., Halliday, G., McCusker, E. & Trent, R. J. Genetically confirmed clinical Huntington's disease with no observable cell loss. J. Neurol. Neurosurg. Psychiatry 74, 968–970 (2003).
Adachi, H. et al. Transgenic mice with an expanded CAG repeat controlled by the human AR promoter show polyglutamine nuclear inclusions and neuronal dysfunction without neuronal cell death. Hum. Mol. Genet. 10, 1039–1048 (2001).
Xuereb, J. H., MacMillan, J. C., Snell, R., Davies, P. & Harper, P. S. Neuropathological diagnosis and CAG repeat expansion in Huntington's disease. J. Neurol. Neurosurg. Psychiatry 60, 78–81 (1996).
Mizuno, H. et al. An autopsy case with clinically and molecular genetically diagnosed Huntington's disease with only minimal non-specific neuropathological findings. Clin. Neuropathol. 19, 94–103 (2000).
Chamberlain, N. L., Driver, E. D. & Miesfeld, R. L. The length and location of CAG trinucleotide repeats in the androgen receptor N-terminal domain affect transactivation function. Nucleic Acids Res. 22, 3181–3186 (1994).
Beilin, J., Ball, E. M., Favaloro, J. M. & Zajac, J. D. Effect of the androgen receptor CAG repeat polymorphism on transcriptional activity: specificity in prostate and non-prostate cell lines. J. Mol. Endocrinol. 25, 85–96 (2000).
Linja, M. J. & Visakorpi, T. Alterations of androgen receptor in prostate cancer. J. Steroid Biochem. Mol. Biol. 92, 255–264 (2004).
Tsujimoto, Y. et al. In situ shortening of CAG repeat length within the androgen receptor gene in prostatic cancer and its possible precursors. Prostate 58, 283–290 (2004).
Strachan, T. & Read, A. P. Human Molecular Genetics 2 2nd edn (John Wiley & Sons, New York, 1999).
Nichol Edamura, K. & Pearson, C. E. DNA methylation and replication: implications for the 'deletion hotspot' region of FMR1. Hum. Genet. (in the press).
Acknowledgements
The authors wish to thank the many scientists, too numerous to list individually, for discussions and comments on this manuscript. In addition, we wish to express our gratitude to the members of the Pearson laboratory (K. Hagerman, R. Lau, M. Leonard and G. Panigrahi) for their input into the manuscript. We apologize to those authors whose work we could not cite owing to space limitations. The Pearson laboratory is supported by funding from the Muscular Dystrophy Association, USA, the Canadian Institutes of Health Research (CIHR), the Fragile X Research Foundation of Canada and the University of Toronto Deans fund. J.D.C. is supported by a CIHR doctoral research award. C.E.P. is a CIHR Scholar and Canadian Genetic Disease Network Scholar.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Related links
Related links
DATABASES
Entrez Gene
OMIM
dentatorubral-pallidoluysian atrophy
facioscapulohumeral muscular dystrophy 1A
myoclonic epilepsy of Unverricht and Lundborg
spinal and bulbar muscular atrophy
FURTHER INFORMATION
Huntington's Disease Society of America
Muscular Dystrophy Association
Neuromuscular Disease Centre DNA Repeat Sequence and Disease page
Tandemly Repeated (Satellite) DNA
The National Fragile X Foundation
Glossary
- GENETIC ANTICIPATION
-
A phenomenon in which disease severity increases and/or age of onset of disease decreases from one generation to the next.
- SPERMATOGONIA
-
The mitotically dividing stem cells of the male germ line, the descendants of which ultimately become mature sperm.
- FULL MUTATION
-
The expanded repeat tract that is typically associated with disease. The term is often used to distinguish this event from an individual who has the shorter premutation or proto-mutation expansions that are not associated with disease.
- PREMUTATION
-
A repeat tract of a length that rarely leads to disease symptoms. However, the possibility for further repeat-length expansion to occur on transmission is high as a result of the longer repeat length. The term applies to disorders such as Huntington disease and fragile X syndrome. The term proto-mutation applies to dystrophia myotonica 1, for which individuals might (or might not) eventually develop symptoms.
- CpG ISLAND
-
A sequence of at least 200 bp with a greater number of CpG sites than expected for its GC content. These regions are often GC rich, associated with genes and typically undermethylated.
- PARENT-OF-ORIGIN EFFECT
-
The increased proportion of paternal or maternal disease-causing transmission to offspring. This effect is molecularly explained by a paternal or maternal repeat-expansion bias in the germ line.
- LINKAGE DISEQUILIBRIUM
-
A measure of genetic associations between alleles at different loci, which indicates whether allelic or marker associations on the same chromosome are more common than expected.
- OKAZAKI FRAGMENT
-
Short DNA fragments (∼140 nucleotides for primates) that are produced on the lagging strand of the replication fork during discontinuous DNA replication; these fragments are eventually processed (to remove the RNA primer) and ligated to form the mature full-length lagging nascent strand.
- GENE CONVERSION
-
A meiotic process that involves the non-reciprocal transfer of genetic information, in which one allele directs the conversion of its sister allele.
- SINGLE-STRAND ANNEALING
-
A process that is typically initiated after a double-strand break, in which regions of ssDNA are created to allow for complementary strands of sister chromatids or homologues to anneal to each other.
- XERODERMA PIGMENTOSUM
-
A human disease that is characterized by extraordinary sensitivity to sunlight, and is caused by a defect in the ultraviolet-mutation-repair system.
Rights and permissions
About this article
Cite this article
Pearson, C., Edamura, K. & Cleary, J. Repeat instability: mechanisms of dynamic mutations. Nat Rev Genet 6, 729–742 (2005). https://doi.org/10.1038/nrg1689
Issue Date:
DOI: https://doi.org/10.1038/nrg1689
This article is cited by
-
Huntington disease-like 2: insight into neurodegeneration from an African disease
Nature Reviews Neurology (2024)
-
LUSTR: a new customizable tool for calling genome-wide germline and somatic short tandem repeat variants
BMC Genomics (2024)
-
The sequence of the repetitive motif influences the frequency of multistep mutations in Short Tandem Repeats
Scientific Reports (2023)
-
Current advances in neuronal intranuclear inclusion disease
Neurological Sciences (2023)
-
Context dependency of nucleotide probabilities and variants in human DNA
BMC Genomics (2022)