Abstract
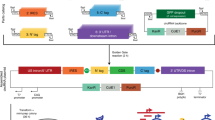
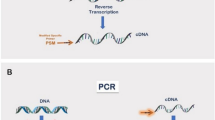
Having knowledge of the entire 3′ sequence of a cDNA is often important because the non-coding terminal region can contain signals that regulate the stability or subcellular localization of the mRNA. Also, some messages use alternative genomic sites for cleavage and polyadenylation that can alter the above properties, or change the encoded protein. Full-length cDNAs can be obtained from complex mixtures of cellular mRNA using rapid amplification of cDNA ends (RACE) PCR as long as part of the mRNA sequence is known; adding non-specific tags to the ends of the cDNA allows the regions between the known parts of the sequence and the ends to be amplified. In 3′ RACE, the poly(A) tail functions as a non-specific tag at the 3′ end of the mRNA. cDNA ends can be obtained in 1–3 days using this protocol.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Frohman, M.A., Dush, M.K. & Martin, G.R. Rapid production of full-length cDNAs from rare transcripts by amplification using a single gene-specific oligonucleotide primer. Proc. Natl Acad. Sci. USA 85, 8998–9002 (1988).
Scotto–Lavino, E., Du, G. & Frohman, M.A. 5′ End cDNA amplification using classic RACE. Nat. Protocols (2006). DOI: 10.1038/nprot.2006.480
Scotto–Lavino, E., Du, G. & Frohman, M.A. 5′ End cDNA amplification using new RACE. Nat. Protocols (2007). DOI: 10.1038/nprot.2006.479
Wickens, M. & Stephenson, P. Role of the conserved AAUAAA sequence: four AAUAAA point mutants prevent mRNA 3′ end formation. Science 226, 1045–1050 (1984).
Coleclough, C. Use of primer-restriction end adapters in cDNA cloning. Methods Enzymol. 154, 64–83 (1987).
Zhang, Y. In Generation of cDNA Libraries (ed. Ying, S.-Y.) 13–24 (Humana Press, Totowa, NJ, 2003).
Acknowledgements
The authors thank Yue Zhang and the publisher for permission to use and adapt material from “RACE all the way to the end” (ref. 6). This work was supported by awards NIHDDK 64166 and NIHGM71520 to M.A.F., and NIHGM071475 and a Scientist Development Grant from the American Heart Association to G.D. (0430096N).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Scotto–Lavino, E., Du, G. & Frohman, M. 3′ End cDNA amplification using classic RACE. Nat Protoc 1, 2742–2745 (2006). https://doi.org/10.1038/nprot.2006.481
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2006.481
This article is cited by
-
Molecular characterization of a novel mycovirus from binucleate Rhizoctonia AG-A strain A46
Archives of Virology (2024)
-
Downregulation of hepatic lncRNA Gm19619 improves gluconeogenesis and lipogenesis following vertical sleeve gastrectomy in mice
Communications Biology (2023)
-
Heterologous expression of Sesuvium portulacastrum SOS-related genes confer salt tolerance in yeast
Acta Physiologiae Plantarum (2023)
-
Molecular cloning and characterization of an alpha-amylase inhibitor (TkAAI) gene from Trichosanthes kirilowii Maxim.
Biotechnology Letters (2022)
-
Disruption of the odorant coreceptor Orco impairs foraging and host finding behaviors in the New World screwworm fly
Scientific Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.