Abstract
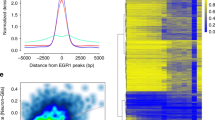
DNA methylation has been traditionally viewed as a highly stable epigenetic mark in postmitotic cells. However, postnatal brains appear to show stimulus-induced methylation changes, at least in a few identified CpG dinucleotides. How extensively the neuronal DNA methylome is regulated by neuronal activity is unknown. Using a next-generation sequencing–based method for genome-wide analysis at single-nucleotide resolution, we quantitatively compared the CpG methylation landscape of adult mouse dentate granule neurons in vivo before and after synchronous neuronal activation. About 1.4% of 219,991 CpGs measured showed rapid active demethylation or de novo methylation. Some modifications remained stable for at least 24 h. These activity-modified CpGs showed a broad genomic distribution with significant enrichment in low-CpG density regions, and were associated with brain-specific genes related to neuronal plasticity. Our study implicates modification of the neuronal DNA methylome as a previously underappreciated mechanism for activity-dependent epigenetic regulation in the adult nervous system.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
References
Reik, W. Stability and flexibility of epigenetic gene regulation in mammalian development. Nature 447, 425–432 (2007).
Suzuki, M.M. & Bird, A. DNA methylation landscapes: provocative insights from epigenomics. Nat. Rev. Genet. 9, 465–476 (2008).
Ma, D.K. et al. Epigenetic choreographers of neurogenesis in the adult mammalian brain. Nat. Neurosci. 13, 1338–1344 (2010).
Wu, S.C. & Zhang, Y. Active DNA demethylation: many roads lead to Rome. Nat. Rev. Mol. Cell Biol. 11, 607–620 (2010).
Zemach, A., McDaniel, I.E., Silva, P. & Zilberman, D. Genome-wide evolutionary analysis of eukaryotic DNA methylation. Science 328, 916–919 (2010).
Hellman, A. & Chess, A. Gene body-specific methylation on the active X chromosome. Science 315, 1141–1143 (2007).
Ball, M.P. et al. Targeted and genome-scale strategies reveal gene-body methylation signatures in human cells. Nat. Biotechnol. 27, 361–368 (2009).
Wu, H. et al. Dnmt3a-dependent nonpromoter DNA methylation facilitates transcription of neurogenic genes. Science 329, 444–448 (2010).
Guo, J.U., Su, Y., Zhong, C., Ming, G.L. & Song, H. Emerging roles of TET proteins and 5-hydroxymethylcytosines in active DNA demethylation and beyond. Cell Cycle 10, 2662–2668 (2011).
Tsankova, N., Renthal, W., Kumar, A. & Nestler, E.J. Epigenetic regulation in psychiatric disorders. Nat. Rev. Neurosci. 8, 355–367 (2007).
Borrelli, E., Nestler, E.J., Allis, C.D. & Sassone-Corsi, P. Decoding the epigenetic language of neuronal plasticity. Neuron 60, 961–974 (2008).
Szyf, M., McGowan, P. & Meaney, M.J. The social environment and the epigenome. Environ. Mol. Mutagen. 49, 46–60 (2008).
Day, J.J. & Sweatt, J.D. DNA methylation and memory formation. Nat. Neurosci. 13, 1319–1323 (2010).
Guan, J.S. et al. HDAC2 negatively regulates memory formation and synaptic plasticity. Nature 459, 55–60 (2009).
Doi, M., Hirayama, J. & Sassone-Corsi, P. Circadian regulator CLOCK is a histone acetyltransferase. Cell 125, 497–508 (2006).
Maze, I. et al. Essential role of the histone methyltransferase G9a in cocaine-induced plasticity. Science 327, 213–216 (2010).
Ma, D.K. et al. Neuronal activity-induced Gadd45b promotes epigenetic DNA demethylation and adult neurogenesis. Science 323, 1074–1077 (2009).
Nelson, E.D., Kavalali, E.T. & Monteggia, L.M. Activity-dependent suppression of miniature neurotransmission through the regulation of DNA methylation. J. Neurosci. 28, 395–406 (2008).
Miller, C.A. et al. Cortical DNA methylation maintains remote memory. Nat. Neurosci. 13, 664–666 (2010).
Feng, J. et al. Dnmt1 and Dnmt3a maintain DNA methylation and regulate synaptic function in adult forebrain neurons. Nat. Neurosci. 13, 423–430 (2010).
LaPlant, Q. et al. Dnmt3a regulates emotional behavior and spine plasticity in the nucleus accumbens. Nat. Neurosci. 13, 1137–1143 (2010).
Martinowich, K. et al. DNA methylation-related chromatin remodeling in activity-dependent BDNF gene regulation. Science 302, 890–893 (2003).
Weaver, I.C. et al. Epigenetic programming by maternal behavior. Nat. Neurosci. 7, 847–854 (2004).
Miller, C.A. & Sweatt, J.D. Covalent modification of DNA regulates memory formation. Neuron 53, 857–869 (2007).
Dong, E., Nelson, M., Grayson, D.R., Costa, E. & Guidotti, A. Clozapine and sulpiride but not haloperidol or olanzapine activate brain DNA demethylation. Proc. Natl. Acad. Sci. USA 105, 13614–13619 (2008).
Lubin, F.D., Roth, T.L. & Sweatt, J.D. Epigenetic regulation of BDNF gene transcription in the consolidation of fear memory. J. Neurosci. 28, 10576–10586 (2008).
Elliott, E., Ezra-Nevo, G., Regev, L., Neufeld-Cohen, A. & Chen, A. Resilience to social stress coincides with functional DNA methylation of the Crf gene in adult mice. Nat. Neurosci. 13, 1351–1353 (2010).
Murgatroyd, C. et al. Dynamic DNA methylation programs persistent adverse effects of early-life stress. Nat. Neurosci. 12, 1559–1566 (2009).
McGowan, P.O. et al. Epigenetic regulation of the glucocorticoid receptor in human brain associates with childhood abuse. Nat. Neurosci. 12, 342–348 (2009).
Lisanby, S.H. Electroconvulsive therapy for depression. N. Engl. J. Med. 357, 1939–1945 (2007).
Shock, L.S., Thakkar, P.V., Peterson, E.J., Moran, R.G. & Taylor, S.M. DNA methyltransferase 1, cytosine methylation, and cytosine hydroxymethylation in mammalian mitochondria. Proc. Natl. Acad. Sci. USA 108, 3630–3635 (2011).
Nass, M.M. Differential methylation of mitochondrial and nuclear DNA in cultured mouse, hamster and virus-transformed hamster cells. In vivo and in vitro methylation. J. Mol. Biol. 80, 155–175 (1973).
Laurent, L. et al. Dynamic changes in the human methylome during differentiation. Genome Res. 20, 320–331 (2010).
Lister, R. et al. Human DNA methylomes at base resolution show widespread epigenomic differences. Nature 462, 315–322 (2009).
Stresemann, C., Brueckner, B., Musch, T., Stopper, H. & Lyko, F. Functional diversity of DNA methyltransferase inhibitors in human cancer cell lines. Cancer Res. 66, 2794–2800 (2006).
Ma, D.K., Guo, J.U., Ming, G.L. & Song, H. DNA excision repair proteins and Gadd45 as molecular players for active DNA demethylation. Cell Cycle 8, 1526–1531 (2009).
Ford, E.C. et al. Localized CT-guided irradiation inhibits neurogenesis in specific regions of the adult mouse brain. Radiat. Res. 175, 774–783 (2011).
Illingworth, R.S. et al. Orphan CpG islands identify numerous conserved promoters in the mammalian genome. PLoS Genet. 6, e1001134 (2010).
Meissner, A. et al. Genome-scale DNA methylation maps of pluripotent and differentiated cells. Nature 454, 766–770 (2008).
Muotri, A.R. et al. Somatic mosaicism in neuronal precursor cells mediated by L1 retrotransposition. Nature 435, 903–910 (2005).
Kim, T.K. et al. Widespread transcription at neuronal activity-regulated enhancers. Nature 465, 182–187 (2010).
Weber, M. et al. Distribution, silencing potential and evolutionary impact of promoter DNA methylation in the human genome. Nat. Genet. 39, 457–466 (2007).
Klug, M. et al. Active DNA demethylation in human postmitotic cells correlates with activating histone modifications, but not transcription levels. Genome Biol. 11, R63 (2010).
Pierfelice, T., Alberi, L. & Gaiano, N. Notch in the vertebrate nervous system: an old dog with new tricks. Neuron 69, 840–855 (2011).
Irizarry, R.A. et al. The human colon cancer methylome shows similar hypo- and hypermethylation at conserved tissue-specific CpG island shores. Nat. Genet. 41, 178–186 (2009).
Herman, J.G. & Baylin, S.B. Gene silencing in cancer in association with promoter hypermethylation. N. Engl. J. Med. 349, 2042–2054 (2003).
Thompson, R.F. et al. Tissue-specific dysregulation of DNA methylation in aging. Aging Cell 9, 506–518 (2010).
Guo, J.U., Su, Y., Zhong, C., Ming, G.L. & Song, H. Hydroxylation of 5-methylcytosine by TET1 promotes active DNA demethylation in the adult brain. Cell 145, 423–434 (2011).
Simonsson, S. & Gurdon, J. DNA demethylation is necessary for the epigenetic reprogramming of somatic cell nuclei. Nat. Cell Biol. 6, 984–990 (2004).
Eaves, H.L. & Gao, Y. MOM: maximum oligonucleotide mapping. Bioinformatics 25, 969–970 (2009).
Acknowledgements
We thank G. Church, S. Baylin, D. Ginty and K. Christian for comments and suggestions; G. Sun for help with FACS; and W.Y. Kim (Johns Hopkins University) for breeding Gadd45b knockout mice. This work was supported by US National Institutes of Health (NIH; AG024984, NS047344), McKnight Scholar Award and NARSAD (Brain and Behavior Research Fund) to H.S.; by NIH (HD069184, NS048271), Johns Hopkins Brain Science Institute, Dr. Miriam and Sheldon G. Adelson Medical Research Foundation, and NARSAD grants to G.-l.M.; and Lieber Institute start-up funds to Y.G. J.U.G. was a FARMS (Foundation for Advanced Research in Medical Sciences) fellow. M.H.J. was supported by a US National Institute of Mental Health K99 award (MH090115). M.A.B. was partially supported by a fellowship from Maryland Stem Cell Research Fund (MSCRF); M.P.B. was supported by grants from the NIH to G. Church.
Author information
Authors and Affiliations
Contributions
J.U.G., D.K.M., G.M., Y.G. and H.S. designed the project. J.U.G. led and was involved in all aspect of the project. H.M. performed sequence mapping and methylation calculation. M.P.B. constructed libraries and performed initial methylation analysis. M.-H.J. assisted in BrdU analysis, irradiation and drug infusion procedures. M.A.B. performed FACS purification. J.A.B. wrote the gene mapping program. H.L.E. adapted MOM for the current project. B.X. and H.M. performed Illumina sequencing. E.F. contributed to irradiation studies. K.Z. contributed to data processing. J.U.G., G.M., Y.G. and H.S. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–22 and Supplementary Table 1 (PDF 3117 kb)
Supplementary Table 2
Primer sets used for bisulfite sequencing analysis of selected CpGs (XLS 32 kb)
Supplementary Table 3
Primer sets used for HpaII-qPCR analysis of selected CpGs (XLS 33 kb)
Supplementary Table 4
MSCC results for all MSCC30+ sites (XLS 10842 kb)
Supplementary Table 5
List of repetitive sequences examined for CpG methylation changes (XLS 63 kb)
Supplementary Table 6
Mouse exon array expression profiles of dentate granule cells at E0 and E4 (XLS 3410 kb)
Supplementary Table 7
Expression profiles of genes associated with activity-modified CpGs (XLS 680 kb)
Supplementary Table 8
Primer sets and results of q-PCR analysis of promoter CpG changes-associated genes (XLS 34 kb)
Supplementary Table 9
Functional pathways that contain activity-modified CpGs (XLS 8485 kb)
Rights and permissions
About this article
Cite this article
Guo, J., Ma, D., Mo, H. et al. Neuronal activity modifies the DNA methylation landscape in the adult brain. Nat Neurosci 14, 1345–1351 (2011). https://doi.org/10.1038/nn.2900
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nn.2900
This article is cited by
-
Brain histone beta-hydroxybutyrylation couples metabolism with gene expression
Cellular and Molecular Life Sciences (2023)
-
Altered hypothalamic DNA methylation and stress-induced hyperactivity following early life stress
Epigenetics & Chromatin (2021)
-
Maternal cholesterol levels during gestation: boon or bane for the offspring?
Molecular and Cellular Biochemistry (2021)
-
Neuronal ensemble-specific DNA methylation strengthens engram stability
Nature Communications (2020)
-
Synaptic control of DNA methylation involves activity-dependent degradation of DNMT3A1 in the nucleus
Neuropsychopharmacology (2020)