Abstract
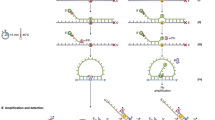
We describe a rapid target enrichment method for next-generation sequencing, termed anchored multiplex PCR (AMP), that is compatible with low nucleic acid input from formalin-fixed paraffin-embedded (FFPE) specimens. AMP is effective in detecting gene rearrangements (without prior knowledge of the fusion partners), single nucleotide variants, insertions, deletions and copy number changes. Validation of a gene rearrangement panel using 319 FFPE samples showed 100% sensitivity (95% confidence limit: 96.5–100%) and 100% specificity (95% confidence limit: 99.3–100%) compared with reference assays. On the basis of our experience with performing AMP on 986 clinical FFPE samples, we show its potential as both a robust clinical assay and a powerful discovery tool, which we used to identify new therapeutically important gene fusions: ARHGEF2-NTRK1 and CHTOP-NTRK1 in glioblastoma, MSN-ROS1, TRIM4-BRAF, VAMP2-NRG1, TPM3-NTRK1 and RUFY2-RET in lung cancer, FGFR2-CREB5 in cholangiocarcinoma and PPL-NTRK1 in thyroid carcinoma. AMP is a scalable and efficient next-generation sequencing target enrichment method for research and clinical applications.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Mamanova, L. et al. Target-enrichment strategies for next-generation sequencing. Nat. Methods 7, 111–118 (2010).
Meyerson, M., Gabriel, S. & Getz, G. Advances in understanding cancer genomes through second-generation sequencing. Nat. Rev. Genet. 11, 685–696 (2010).
Kwak, E.L. et al. Anaplastic lymphoma kinase inhibition in non-small-cell lung cancer. N. Engl. J. Med. 363, 1693–1703 (2010).
Bergethon, K. et al. ROS1 rearrangements define a unique molecular class of lung cancers. J. Clin. Oncol. 30, 863–870 (2012).
Drilon, A. et al. Response to Cabozantinib in patients with RET fusion-positive lung adenocarcinomas. Cancer Discov. 3, 630–635 (2013).
Heuckmann, J.M. et al. Differential protein stability and ALK inhibitor sensitivity of EML4-ALK fusion variants. Clin. Cancer Res. 18, 4682–4690 (2012).
Crystal, A.S. & Shaw, A.T. Variants on a theme: a biomarker of crizotinib response in ALK-positive non-small cell lung cancer? Clin. Cancer Res. 18, 4479–4481 (2012).
Takeuchi, K. et al. RET, ROS1 and ALK fusions in lung cancer. Nat. Med. 18, 378–381 (2012).
Frohman, M.A., Dush, M.K. & Martin, G.R. Rapid production of full-length cDNAs from rare transcripts: amplification using a single gene-specific oligonucleotide primer. Proc. Natl. Acad. Sci. USA 85, 8998–9002 (1988).
Zheng, Z. et al. Titration-free 454 sequencing using Y adapters. Nat. Protoc. 6, 1367–1376 (2011).
Zheng, Z. et al. Titration-free massively parallel pyrosequencing using trace amounts of starting material. Nucleic Acids Res. 38, e137 (2010).
Neiman, M. et al. Library preparation and multiplex capture for massive parallel sequencing applications made efficient and easy. PLoS ONE 7, e48616 (2012).
Wang, T., Yu, D. & Lamb, M.L. Trk kinase inhibitors as new treatments for cancer and pain. Expert Opin. Ther. Pat. 19, 305–319 (2009).
Dias-Santagata, D. et al. Rapid targeted mutational analysis of human tumours: a clinical platform to guide personalized cancer medicine. EMBO Mol. Med. 2, 146–158 (2010).
Chmielecki, J. et al. Targeted next-generation sequencing of DNA regions proximal to a conserved GXGXXG signaling motif enables systematic discovery of tyrosine kinase fusions in cancer. Nucleic Acids Res. 38, 6985–6996 (2010).
Aird, D. et al. Analyzing and minimizing PCR amplification bias in Illumina sequencing libraries. Genome Biol. 12, R18 (2011).
Oyola, S.O. et al. Optimizing Illumina next-generation sequencing library preparation for extremely AT-biased genomes. BMC Genomics 13, 1 (2012).
Chevet, E., Lemaitre, G. & Katinka, M.D. Low concentrations of tetramethylammonium chloride increase yield and specificity of PCR. Nucleic Acids Res. 23, 3343–3344 (1995).
Tewhey, R. et al. Microdroplet-based PCR enrichment for large-scale targeted sequencing. Nat. Biotechnol. 27, 1025–1031 (2009).
Turner, E.H. et al. Massively parallel exon capture and library-free resequencing across 16 genomes. Nat. Methods 6, 315–316 (2009).
Beltran, H. et al. Targeted next-generation sequencing of advanced prostate cancer identifies potential therapeutic targets and disease heterogeneity. Eur. Urol. 63, 920–926 (2013).
Clark, M.J. et al. Performance comparison of exome DNA sequencing technologies. Nat. Biotechnol. 29, 908–914 (2011).
Yousem, S.A. et al. Pulmonary Langerhans cell histiocytosis: profiling of multifocal tumors using next-generation sequencing identifies concordant occurrence of BRAF V600E mutations. Chest 143, 1679–1684 (2013).
Do, H. et al. Reducing sequence artifacts in amplicon-based massively parallel sequencing of formalin-fixed paraffin-embedded DNA by enzymatic depletion of uracil-containing templates. Clin. Chem. 59, 1376–1383 (2013).
Johansson, H. et al. Targeted resequencing of candidate genes using selector probes. Nucleic Acids Res. 39, e8 (2011).
Varley, K.E. & Mitra, R.D. Nested Patch PCR enables highly multiplexed mutation discovery in candidate genes. Genome Res. 18, 1844–1850 (2008).
Donner, A. & Rotondi, M.A. Sample size requirements for interval estimation of the kappa statistic for interobserver agreement studies with a binary outcome and multiple raters. Int. J. Biostat. 6, 31 (2010).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Kent, W.J. BLAT–the BLAST-like alignment tool. Genome Res. 12, 656–664 (2002).
Koboldt, D.C. et al. VarScan 2: somatic mutation and copy number alteration discovery in cancer by exome sequencing. Genome Res. 22, 568–576 (2012).
Untergasser, A. et al. Primer3—new capabilities and interfaces. Nucleic Acids Res. 40, e115 (2012).
Acknowledgements
This study was supported by a US National Institutes of Health grant (R21CA161590) to A.J.I. and a postdoctoral fellowship from the Swedish Research Council (350-2012-368) to Z.Z.
Author information
Authors and Affiliations
Contributions
Z.Z., M.L., B.Z., Y.C., D.P., K.D.L., J.C., H.E.R., H.S.S., J.C., W.P. & J.A.E. conducted the experiments; Z.Z. and L.P.L. conducted the data analyses; Z.Z. and L.P.L. wrote the manuscript; A.J.I. supervised the project.
Corresponding authors
Ethics declarations
Competing interests
A provisional patent has been filed by A.J.I., L.P.L. and Z.Z. for the anchored multiplex PCR technology with the US Patent Office. A.J.I., L.P.L. and Z.Z. are equity holders in Enzymatics, Inc., a licensee of the technology.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 and Supplementary Tables 1–7 (PDF 10310 kb)
Rights and permissions
About this article
Cite this article
Zheng, Z., Liebers, M., Zhelyazkova, B. et al. Anchored multiplex PCR for targeted next-generation sequencing. Nat Med 20, 1479–1484 (2014). https://doi.org/10.1038/nm.3729
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.3729
This article is cited by
-
TRIM4 Expression Related to Malignant Progression and Cisplatin Resistance in Osteosarcoma
Applied Biochemistry and Biotechnology (2024)
-
satmut_utils: a simulation and variant calling package for multiplexed assays of variant effect
Genome Biology (2023)
-
Utilizing cell-free DNA to predict risk of developing brain metastases in patients with metastatic breast cancer
npj Breast Cancer (2023)
-
Personalized Cancer Monitoring Assay for the Detection of ctDNA in Patients with Solid Tumors
Molecular Diagnosis & Therapy (2023)
-
Metastatic pituitary tumors: an institutional case series
Pituitary (2023)