Abstract
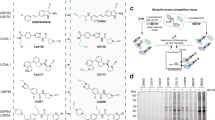
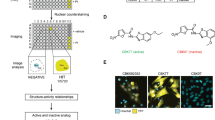
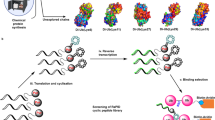
Protein ubiquitination and deubiquitination are central to the control of a large number of cellular pathways and signaling networks in eukaryotes. Although the essential roles of ubiquitination have been established in the eukaryotic DNA damage response, the deubiquitination process remains poorly defined. Chemical probes that perturb the activity of deubiquitinases (DUBs) are needed to characterize the cellular function of deubiquitination. Here we report ML323 (2), a highly potent inhibitor of the USP1–UAF1 deubiquitinase complex with excellent selectivity against human DUBs, deSUMOylase, deneddylase and unrelated proteases. Using ML323, we interrogated deubiquitination in the cellular response to UV- and cisplatin-induced DNA damage and revealed new insights into the requirement of deubiquitination in the DNA translesion synthesis and Fanconi anemia pathways. Moreover, ML323 potentiates cisplatin cytotoxicity in non–small cell lung cancer and osteosarcoma cells. Our findings point to USP1–UAF1 as a key regulator of the DNA damage response and a target for overcoming resistance to the platinum-based anticancer drugs.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Chen, Z.J. & Sun, L.J. Nonproteolytic functions of ubiquitin in cell signaling. Mol. Cell 33, 275–286 (2009).
Amerik, A.Y. & Hochstrasser, M. Mechanism and function of deubiquitinating enzymes. Biochim. Biophys. Acta 1695, 189–207 (2004).
Nijman, S.M. et al. A genomic and functional inventory of deubiquitinating enzymes. Cell 123, 773–786 (2005).
Fraile, J.M., Quesada, V., Rodriguez, D., Freije, J.M. & Lopez-Otin, C. Deubiquitinases in cancer: new functions and therapeutic options. Oncogene 31, 2373–2388 (2012).
Wang, W. Emergence of a DNA-damage response network consisting of Fanconi anaemia and BRCA proteins. Nat. Rev. Genet. 8, 735–748 (2007).
Kim, H. & D'Andrea, A.D. Regulation of DNA cross-link repair by the Fanconi anemia/BRCA pathway. Genes Dev. 26, 1393–1408 (2012).
Crossan, G.P. & Patel, K.J. The Fanconi anaemia pathway orchestrates incisions at sites of crosslinked DNA. J. Pathol. 226, 326–337 (2012).
Hoege, C., Pfander, B., Moldovan, G.L., Pyrowolakis, G. & Jentsch, S. RAD6-dependent DNA repair is linked to modification of PCNA by ubiquitin and SUMO. Nature 419, 135–141 (2002).
Stelter, P. & Ulrich, H.D. Control of spontaneous and damage-induced mutagenesis by SUMO and ubiquitin conjugation. Nature 425, 188–191 (2003).
Chen, J., Bozza, W. & Zhuang, Z. Ubiquitination of PCNA and its essential role in eukaryotic translesion synthesis. Cell Biochem. Biophys. 60, 47–60 (2011).
Chang, D.J. & Cimprich, K.A. DNA damage tolerance: when it's OK to make mistakes. Nat. Chem. Biol. 5, 82–90 (2009).
Haracska, L., Torres-Ramos, C.A., Johnson, R.E., Prakash, S. & Prakash, L. Opposing effects of ubiquitin conjugation and SUMO modification of PCNA on replicational bypass of DNA lesions in Saccharomyces cerevisiae. Mol. Cell. Biol. 24, 4267–4274 (2004).
Huang, T.T. et al. Regulation of monoubiquitinated PCNA by DUB autocleavage. Nat. Cell Biol. 8, 339–347 (2006).
Zhuang, Z. et al. Regulation of polymerase exchange between Polη and Polδ by monoubiquitination of PCNA and the movement of DNA polymerase holoenzyme. Proc. Natl. Acad. Sci. USA 105, 5361–5366 (2008).
Lee, K.Y. et al. Human ELG1 regulates the level of ubiquitinated proliferating cell nuclear antigen (PCNA) through its interactions with PCNA and USP1. J. Biol. Chem. 285, 10362–10369 (2010).
Kim, J.M. et al. Inactivation of murine Usp1 results in genomic instability and a Fanconi anemia phenotype. Dev. Cell 16, 314–320 (2009).
Guervilly, J.H., Renaud, E., Takata, M. & Rosselli, F. USP1 deubiquitinase maintains phosphorylated CHK1 by limiting its DDB1-dependent degradation. Hum. Mol. Genet. 20, 2171–2181 (2011).
Bell, D.W. et al. Predisposition to cancer caused by genetic and functional defects of mammalian Atad5. PLoS Genet. 7, e1002245 (2011).
Sowa, M.E., Bennett, E.J., Gygi, S.P. & Harper, J.W. Defining the human deubiquitinating enzyme interaction landscape. Cell 138, 389–403 (2009).
Oestergaard, V.H. et al. Deubiquitination of FANCD2 is required for DNA crosslink repair. Mol. Cell 28, 798–809 (2007).
Chen, J. et al. Selective and cell-active inhibitors of the USP1/UAF1 deubiquitinase complex reverse cisplatin resistance in non-small cell lung cancer cells. Chem. Biol. 18, 1390–1400 (2011).
Post, R.M., Jimerson, D.C., Bunney, W.E. Jr. & Goodwin, F.K. Dopamine and mania: behavioral and biochemical effects of the dopamine receptor blocker pimozide. Psychopharmacology (Berl.) 67, 297–305 (1980).
Brown, P.J. et al. Identification of a subtype selective human PPARα agonist through parallel-array synthesis. Bioorg. Med. Chem. Lett. 11, 1225–1227 (2001).
Prage, E.B. et al. Location of inhibitor binding sites in the human inducible prostaglandin E synthase, MPGES1. Biochemistry 50, 7684–7693 (2011).
Xiao, H. et al. Insights into the mechanism of microtubule stabilization by Taxol. Proc. Natl. Acad. Sci. USA 103, 10166–10173 (2006).
Khrapunovich-Baine, M. et al. Distinct pose of discodermolide in taxol binding pocket drives a complementary mode of microtubule stabilization. Biochemistry 48, 11664–11677 (2009).
Villamil, M.A. et al. Serine phosphorylation is critical for the activation of ubiquitin-specific protease 1 and its interaction with WD40-repeat protein UAF1. Biochemistry 51, 9112–9123 (2012).
Borodovsky, A. et al. Chemistry-based functional proteomics reveals novel members of the deubiquitinating enzyme family. Chem. Biol. 9, 1149–1159 (2002).
Altun, M. et al. Activity-based chemical proteomics accelerates inhibitor development for deubiquitylating enzymes. Chem. Biol. 18, 1401–1412 (2011).
Köberle, B., Tomicic, M.T., Usanova, S. & Kaina, B. Cisplatin resistance: preclinical findings and clinical implications. Biochim. Biophys. Acta 1806, 172–182 (2010).
Chou, T.C. Drug combination studies and their synergy quantification using the Chou-Talalay method. Cancer Res. 70, 440–446 (2010).
Albertella, M.R., Green, C.M., Lehmann, A.R. & O'Connor, M.J. A role for polymerase η in the cellular tolerance to cisplatin-induced damage. Cancer Res. 65, 9799–9806 (2005).
Johnson, R.E., Prakash, S. & Prakash, L. Efficient bypass of a thymine-thymine dimer by yeast DNA polymerase, Polη. Science 283, 1001–1004 (1999).
You, C. et al. A quantitative assay for assessing the effects of DNA lesions on transcription. Nat. Chem. Biol. 8, 817–822 (2012).
Kim, H., Yang, K., Dejsuphong, D. & D'Andrea, A.D. Regulation of Rev1 by the Fanconi anemia core complex. Nat. Struct. Mol. Biol. 19, 164–170 (2012).
Renaud, E. & Rosselli, F. FANC pathway promotes UV-induced stalled replication forks recovery by acting both upstream and downstream Polη and Rev1. PLoS ONE 8, e53693 (2013).
Li, X. & Heyer, W.D. Homologous recombination in DNA repair and DNA damage tolerance. Cell Res. 18, 99–113 (2008).
Niedzwiedz, W. et al. The Fanconi anaemia gene FANCC promotes homologous recombination and error-prone DNA repair. Mol. Cell 15, 607–620 (2004).
Xia, B. et al. Control of BRCA2 cellular and clinical functions by a nuclear partner, PALB2. Mol. Cell 22, 719–729 (2006).
Sonoda, E. et al. Multiple roles of Rev3, the catalytic subunit of polζ in maintaining genome stability in vertebrates. EMBO J. 22, 3188–3197 (2003).
Takata, M., Ishiai, M. & Kitao, H. The Fanconi anemia pathway: Insights from somatic cell genetics using DT40 cell line. Mutat. Res. 668, 92–102 (2009).
Lebwohl, D. & Canetta, R. Clinical development of platinum complexes in cancer therapy: an historical perspective and an update. Eur. J. Cancer 34, 1522–1534 (1998).
Kelland, L. The resurgence of platinum-based cancer chemotherapy. Nat. Rev. Cancer 7, 573–584 (2007).
Brabec, V. & Kasparkova, J. Molecular aspects of resistance to antitumor platinum drugs. Drug Resist. Updat. 5, 147–161 (2002).
Park, E. et al. FANCD2 activates transcription of TAp63 and suppresses tumorigenesis. Mol. Cell 50, 908–918 (2013).
Williams, S.A. et al. USP1 deubiquitinates ID proteins to preserve a mesenchymal stem cell program in osteosarcoma. Cell 146, 918–930 (2011).
Xu, G., Paige, J.S. & Jaffrey, S.R. Global analysis of lysine ubiquitination by ubiquitin remnant immunoaffinity profiling. Nat. Biotechnol. 28, 868–873 (2010).
Burkhart, R.A. et al. Mitoxantrone targets human ubiquitin-specific peptidase 11 (USP11) and is a potent inhibitor of pancreatic cancer cell survival. Mol. Cancer Res. 11, 901–911 (2013).
Hibbert, R.G. & Sixma, T.K. Intrinsic flexibility of ubiquitin on proliferating cell nuclear antigen (PCNA) in translesion synthesis. J. Biol. Chem. 287, 39216–39223 (2012).
Inglese, J. et al. Quantitative high-throughput screening: a titration-based approach that efficiently identifies biological activities in large chemical libraries. Proc. Natl. Acad. Sci. USA 103, 11473–11478 (2006).
You, C. et al. Translesion synthesis of 8,5′-cyclopurine-2′-deoxynucleosides by DNA polymerases η, ι, and ζ. J. Biol. Chem. 288, 28548–28556 (2013).
Baker, D.J. et al. Nucleotide excision repair eliminates unique DNA-protein cross-links from mammalian cells. J. Biol. Chem. 282, 22592–22604 (2007).
Yuan, B. et al. The roles of DNA polymerases κ and ι in the error-free bypass of N2-carboxyalkyl-2′-deoxyguanosine lesions in mammalian cells. J. Biol. Chem. 286, 17503–17511 (2011).
Mitra, D. et al. An ultraviolet-radiation–independent pathway to melanoma carcinogenesis in the red hair/fair skin background. Nature 491, 449–453 (2012).
Burns, J.A., Dreij, K., Cartularo, L. & Scicchitano, D.A. O6-methylguanine induces altered proteins at the level of transcription in human cells. Nucleic Acids Res. 38, 8178–8187 (2010).
Sanchez, J.A., Marek, D. & Wangh, L.J. The efficiency and timing of plasmid DNA replication in Xenopus eggs: correlations to the extent of prior chromatin assembly. J. Cell Sci. 103, 907–918 (1992).
Taylor, E.R. & Morgan, I.M. A novel technique with enhanced detection and quantitation of HPV-16 E1- and E2-mediated DNA replication. Virology 315, 103–109 (2003).
Delaney, J.C. & Essigmann, J.M. Mutagenesis, genotoxicity, and repair of 1-methyladenine, 3-alkylcytosines, 1-methylguanine, and 3-methylthymine in alkB Escherichia coli. Proc. Natl. Acad. Sci. USA 101, 14051–14056 (2004).
Delaney, J.C. & Essigmann, J.M. Assays for determining lesion bypass efficiency and mutagenicity of site-specific DNA lesions in vivo. Methods Enzymol. 408, 1–15 (2006).
Acknowledgements
We thank M. Cordeiro-Stone (University of North Carolina at Chapel Hill) for xeroderma pigmentosum variant (XPV) human fibroblasts GM02359-hTERT (XP115LO) and Polη-complemented GM02359-hTERT (XPV + Polη) and M. Jasin (Memorial Sloan-Kettering Cancer Center) for DR-GFP U2OS cells. We also thank C. Arrowsmith (University of Toronto and Ontario Cancer Institute) for the USP21 plasmid; A. Tencer for assistance with protein purification; S. Michael and R. Jones for automation support; P. Shinn and D. van Leer for assistance with compound management; and W. Leister, H. Baker, C. Leclair and E. Fernandez for analytical chemistry and compound purification support. This work was supported by US National Institutes of Health (NIH) grant R03DA030552 and in part by NIH grant R01GM097468 to Z.Z. C.A.O. was supported by NIH training grant T32GM008550. Work in the laboratory of Y.W. was supported by NIH grant R01DK082779. T.S.D., A.S.R., D.K.L., A.S., A.J. and D.J.M. were supported by the intramural research program of the National Center for Advancing Translational Sciences and the Molecular Libraries Initiative of the National Institutes of Health Roadmap for Medical Research (U54MH084681).
Author information
Authors and Affiliations
Contributions
Q.L., T.S.D. and J.C. designed, performed and analyzed in vitro biochemical assays for DUBs and other proteases. T.S.D. and A.S. designed and performed the HTS assay. Q.L. and P.Z. designed, performed and analyzed cell culture experiments. A.S.R., D.K.L. and D.J.M. performed chemical synthesis and oversaw the chemistry efforts. M.A.V. performed protein purification and native gel analysis. C.Y. and B.Y. performed the cellular DNA replication assay under the guidance of Y.W. Q.Z. and H.X. performed and analyzed HDX experiments. C.A.O. generated substrates for the DUB assay. H.S. and A.J. analyzed HTS data and structure-activity relationship results. Z.Z. and D.J.M. designed the research plan and wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
A patent application on the compounds described in the paper as DUB inhibitors has been filed.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Figures 1–20, Supplementary Tables 1–3 and Supplementary Note. (PDF 23259 kb)
Rights and permissions
About this article
Cite this article
Liang, Q., Dexheimer, T., Zhang, P. et al. A selective USP1–UAF1 inhibitor links deubiquitination to DNA damage responses. Nat Chem Biol 10, 298–304 (2014). https://doi.org/10.1038/nchembio.1455
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1455
This article is cited by
-
Spotlights on ubiquitin-specific protease 12 (USP12) in diseases: from multifaceted roles to pathophysiological mechanisms
Journal of Translational Medicine (2023)
-
The promotive role of USP1 inhibition in coordinating osteogenic differentiation and fracture healing during nonunion
Journal of Orthopaedic Surgery and Research (2023)
-
Deubiquitinase USP1 enhances CCAAT/enhancer-binding protein beta (C/EBPβ) stability and accelerates adipogenesis and lipid accumulation
Cell Death & Disease (2023)
-
Discovery of an OTUD3 inhibitor for the treatment of non-small cell lung cancer
Cell Death & Disease (2023)
-
Contribution of microRNA-30d to the prevention of the thyroid cancer occurrence and progression: mechanism and implications
Apoptosis (2023)