Abstract
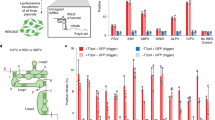
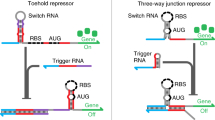
Recent studies have demonstrated the important enzymatic, structural and regulatory roles of RNA in the cell. Here we present a post-transcriptional regulation system in Escherichia coli that uses RNA to both silence and activate gene expression. We inserted a complementary cis sequence directly upstream of the ribosome binding site in a target gene. Upon transcription, this cis-repressive sequence causes a stem-loop structure to form at the 5′–untranslated region of the mRNA. The stem-loop structure interferes with ribosome binding, silencing gene expression. A small noncoding RNA that is expressed in trans targets the cis-repressed RNA with high specificity, causing an alteration in the stem-loop structure that activates expression. Such engineered riboregulators may lend insight into mechanistic actions of endogenous RNA-based processes and could serve as scalable components of biological networks, able to function with any promoter or gene to directly control gene expression.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Gesteland, R.F., Cech, T.R. & Atkins, J.F. The RNA World, edn. 2 (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, 1999).
Joyce, G.F. The antiquity of RNA-based evolution. Nature 418, 214–221 (2002).
Eddy, S.R. Non-coding RNA genes and the modern RNA world. Nat. Rev. Genet. 2, 919–929 (2001).
Kruger, K. et al. Self-splicing RNA: autoexcision and autocyclization of the ribosomal RNA intervening sequence of Tetrahymena. Cell 31, 147–157 (1982).
Guerrier-Takada, C., Gardiner, K., Marsh, T., Pace, N. & Altman, S. The TNA moiety of ribonuclease P is the catalytic subunit of the enzyme. Cell 35, 849–857 (1983).
Doudna, J.A. & Cech, T.R. The chemical repertoire of natural ribozymes. Nature 418, 222–228 (2002).
Steward, P., Molin, S. & Nordstrom, K. RNAs involved in copy-number control and incompatibility of plasmid R1. Proc. Natl. Acad. Sci. USA 78, 6008–6012 (1981).
Wagner, E.G.H. & Simons, R.W. Antisense RNA control in bacteria, phages, and plasmids. Annu. Rev. Microbiol. 48, 713–742 (1994).
Lee, R.C., Feinbaum, R.L. & Ambros, V. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 75, 843–854 (1993).
Gottesman, S. Stealth regulation: biological circuits with small RNA switches. Genes Dev. 16, 2829–2842 (2002).
Gelfand, M.S., Mironov, A.A., Jomantas, J., Kozlov, Y.I. & Perumov, D.A. A conserved RNA structure element involved in the regulation of bacterial riboflavin synthesis genes. Trends Genet. 15, 439–442 (1999).
Winkler, W., Nahvi, A. & Breaker, R.R. Thiamine derivatives bind messenger RNAs directly to regulate bacterial gene expression. Nature 419, 952–656 (2002).
Johansson, J. et al. An RNA thermosensor controls expression of virulence genes in Listeria monocytogenes. Cell 110, 551–561 (2002).
Mironov, A. et al. Sensing small molecules by nascent RNA: a mechanism to control transcription in bacteria. Cell 111, 747–756 (2002).
Morita, M.T. et al. Translational induction of heat shock transcription factor sigma32: evidence for a built-in RNA thermosensor. Genes Dev. 13, 655–665 (2002).
Mandal, M., Boese, B., Barrick, J.E., Winkler, W.C. & Breaker, R.R. Riboswitches control fundamental biochemical pathways in Bacillus subtilis and other bacteria. Cell 113, 577–586 (2003).
Winkler, W., Nahvi, A., Roth, A., Collins, J.A. & Breaker, R.R. Control of gene expression by a natural metabolite-responsive ribozyme. Nature 428, 281–286 (2004).
Lease, R.A. & Belfort, M. A trans-acting RNA as a control switch in Escherichia coli: DsrA modulates function by forming alternative structures. Proc. Natl. Acad. Sci. USA 97, 9919–9924 (2000).
Majdalani, N., Hernandez, D. & Gottesman, S. Regulation and mode of action of the second small RNA activator of RpoS translation, RprA. Mol. Microbiol. 46, 813–826 (2002).
Opalinska, J.B. & Gewirtz, A.M. Nucleic–acid therapeutics: basic principles and recent applications. Nat. Rev. Drug Discov. 1, 503–514 (2002).
Good, L. Translation repression by antisense sequences. Cell. Mol. LifeSci. 60, 854–861 (2003).
Ji, Y. et al. Identification of critical Staphylococcal gene using conditional phenotypes generated by antisense RNA. Science 293, 2266–2269 (2001).
Dykxhoorn, D., Novina, C.D. & Sharp, P.A. Killing the messenger: Short RNAs that silence gene expression. Nat. Rev. Mol. Cell Biol. 4, 457–467 (2003).
Wagner, E.G.H. & Flardh, K. Antisense RNAs everywhere? Trends Genet. 18, 223–226 (2002).
Engdahl, H.M., Lindell, M. & Wagner, E.G.H. Introduction of an RNA stability element at the 5′–end of an antisense RNA cassette increases the inhibition of target RNA translation. Antisense Nucleic A. 11, 29–40 (2001).
Morfeldt, E., Taylor, D., von Gabain, A. & Arvidson, S. Activation of alpha-toxin translation in Staphylococcus aureus by the trans-encoded antisense RNA, RNAIII. EMBO J. 14, 4569–4577 (1995).
Franch, T., Petersen, M., Wagner, E.G.H., Jacobsen, J.P. & Gerdes, K. Antisense RNA regulation in prokaryotes: rapid RNA/RNA interaction facilitated by a general U-turn loop structure. J. Mol. Biol. 294, 1115–1125 (1999).
Zuker, M. Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res. 31, 3406–3415 (2003).
Lutz, R. & Bujard, H. Independent and tight regulation of transcriptional units in Escherichia coli via the LacR/O, the TetR/O and AraCI1-I2 regulatory elements. Nucleic Acids Res. 25, 1203–1210 (1997).
Cormack, B.P., Valdivia, R.C. & Falkow, S. FACS-optimized mutants of the green fluorescent protein (GFP). Gene 173, 33–38 (1996).
Hjalt, T.A.H. & Wagner, E.G.H. Bulged-out nucleotides protect an antisense RNA from RNase III cleavage. Nucleic Acids Res. 23, 571–579 (1995).
Court, D. RNA processing and degradation by RNase III. Control of Messenger RNA Stability, (eds. Belasco, J. & Brawerman, G.) 71–116. (Academic Press, New York, 1993).
Rivas, E., Klein, R.J., Jones, T.A. & Eddy, S.R. Computational identification of noncoding RNAs in E. coli by comparative genomics. Curr. Biol. 11, 1369–1373 (2001).
Altuvia, S. & Wagner, E.G.H. Switching on and off with RNA. Proc. Natl. Acad. Sci. USA 97, 9824–9826 (2000).
Hasty, J., McMillen, D. & Collins, J.J. Engineered gene circuits. Nature 420, 224–230 (2002).
Gardner, T.S., di Bernardo, D., Lorenz, D. & Collins, J.J. Inferring genetic networks and identifying compound mode of action via expression profiling. Science 301, 102–105 (2003).
Bartel, D.P. & Szostak, J.W. Isolation of new ribozymes from a large pool of random sequences. Science 261, 1411–1418 (1993).
Wilson, D.S. & Szostak, J.W. In vitro selection of functional nucleic acids. Annu. Rev. Biochem. 68, 611–647 (1999).
Ellington, A.D. & Szostak, J.W. In vitro selection of RNA molecules that bind specific ligands. Nature 346, 818–822 (1990).
Tuerk, C. & Gold, L. Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA polymerase. Science 249, 505–510 (1990).
Joyce, G.F. Amplification, mutation and selection of catalytic RNA. Gene 82, 83–87 (1989).
Robertson, M.P. & Ellington, A.D. In vitro selection of an allosteric ribozyme that transduces analytes to amplicons. Nat. Biotechnol. 17, 62–66 (1999).
Faulhammer, D., Cukras, A.R., Lipton, R.J. & Landweber, L.F. Molecular computation: RNA solutions to chess problems. Proc. Natl. Acad. Sci. USA 97, 1385–1389 (2000).
Smith, H.O., Hutchison, C.A. III, Pfannkoch, C. & Venter, J.C. Generating a synthetic genome by whole genome assembly: φx174 bacteriophage from synthetic oligonucleotides. Proc. Natl. Acad. Sci. USA 100, 15440–15445 (2003).
Tabor, J.J. & Ellington, A.D. Playing to win at DNA computation. Nat. Biotechnol. 21, 1013–1015 (2003).
Stojanovic, M.N. & Stefanovic, D. A deoxyribozyme–based molecular automaton. Nat. Biotechnol. 21, 1069–1074 (2003).
Sambrook, J., Fritsch, E.F. & Maniatis, T. Molecular Cloning: A Laboratory Manual, edn. 2 (Cold Spring Harbor Laboratory Press, Plainview, New York, 1989).
Andersen, J.B. et al. New unstable variants of green fluorescent protein for studies of transient gene expression in bacteria. Appl. Environ. Microbiol. 64, 2240–2246 (1998).
Ding, C. & Cantor, C.R. A high-throughput gene expression analysis technique using competitive PCR and matrix-assisted laser desorption ionization time-of-flight MS. Proc. Natl. Acad. Sci. USA 100, 3059–3064 (2003).
Marky, L.A. & Breslauer, K.J. Calculating thermodynamic data for transitions of any molecularity from equilibrium melting curves. Biopolymers 26, 1601–1620 (1987).
Acknowledgements
We thank T. Yoshida for providing access to the UV spectrophotometer; E. Protozanova for discussions and advice with RNA melting experiments; I. Smolina for help and advice with reverse transcription experiments; W. Blake, J. Hasty, D.H. Lee, J. Graber and members of our lab for helpful discussions and advice in preparing the manuscript. This work was supported by the National Science Foundation and Defense Advanced Research Projects Agency.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare a pending patent application whose value may be affected by publication of this paper.
Supplementary information
Supplementary Table 1
List of plasmids in this study. aThe pBADHisA vector was obtained from Invitrogen. (PDF 32 kb)
Supplementary Table 2
Sequences of cis-repressed RNA constructs, loop containing the YUNR (TTGG) recognition motif, ribosome binding site (RBS), and trans-activating RNA constructs used in this work. (PDF 80 kb)
Supplementary Table 3
Real-competitive PCR assay design. List of primers used to amplify RTPCR products obtained from RNA cell preparations. A terminator mix contains three different ddNTPs and one dNTP. For example, CGT mix for 16S rRNA is ddCTP/ddGTP/ddTTP/dATP. (PDF 9 kb)
Supplementary Notes
Rational attempts to increase dynamic range of taR12-crR12 (PDF 100 kb)
Supplementary Fig. 1
Set of plasmids used in the artificial riboregulator systems. (PDF 49 kb)
Supplementary Fig. 2
Reverse transcription profiles of taRNA-crRNA complexes. (PDF 140 kb)
Supplementary Fig. 3
Determination of equilibrium dissociation constants for the taR7-crR12 pair. (PDF 34 kb)
Supplementary Fig. 4
RNA Melting curves for crR7, crR10, and crR12. Absorbance measurements at 260 nm (OD260) were determined between 10–95°C for each construct. (PDF 47 kb)
Rights and permissions
About this article
Cite this article
Isaacs, F., Dwyer, D., Ding, C. et al. Engineered riboregulators enable post-transcriptional control of gene expression. Nat Biotechnol 22, 841–847 (2004). https://doi.org/10.1038/nbt986
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt986
This article is cited by
-
Cas9-assisted biological containment of a genetically engineered human commensal bacterium and genetic elements
Nature Communications (2024)
-
Engineering the gut microbiome
Nature Reviews Bioengineering (2023)
-
Elimination of editing plasmid mediated by theophylline riboswitch in Zymomonas mobilis
Applied Microbiology and Biotechnology (2023)
-
Integral feedback in synthetic biology: negative-equilibrium catastrophe
Journal of Mathematical Chemistry (2023)
-
aRNAque: an evolutionary algorithm for inverse pseudoknotted RNA folding inspired by Lévy flights
BMC Bioinformatics (2022)