Abstract
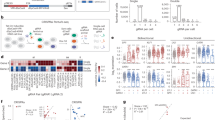
The genetic variation causal for predisposition to type 1 diabetes (T1D) remains unidentified for the majority of known T1D risk loci. MicroRNAs function as post-transcriptional gene regulators by targeting microRNA-binding sites in the 3′ untranslated regions (UTR) of mRNA. Genetic variation within the 3′-UTR of T1D-associated genes may contribute to T1D development by altering microRNA-mediated gene regulation. In silico analysis of variable sites predicted altered microRNA binding in established T1D loci. Functional implications were assessed for variable sites in the 3′-UTR of T1D candidate risk genes CTLA4 and IL10, both involved in immune regulation. We confirmed that in these genes 3′-UTR variation either disrupted or introduced a microRNA-binding site, affecting the repressive capacity of miR-302a* and miR-523, respectively. Our study points to the potential of 3′-UTR variation to affect T1D pathogenesis by altering post-transcriptional gene regulation by microRNAs.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 6 digital issues and online access to articles
$119.00 per year
only $19.83 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Coppieters KT, Dotta F, Amirian N, Campbell PD, Kay TW, Atkinson MA et al. Demonstration of islet-autoreactive CD8 T cells in insulitic lesions from recent onset and long-term type 1 diabetes patients. J Exp Med 2012; 209: 51–60.
Burton PR, Clayton DG, Cardon LR, Craddock N, Deloukas P, Duncanson A et al. Association scan of 14,500 nonsynonymous SNPs in four diseases identifies autoimmunity variants. Nat Genet 2007; 39: 1329–1337.
Barrett JC, Clayton DG, Concannon P, Akolkar B, Cooper JD, Erlich HA et al. Genome-wide association study and meta-analysis find that over 40 loci affect risk of type 1 diabetes. Nat Genet 2009; 41: 703–707.
Maier LM, Hafler DA . Autoimmunity risk alleles in costimulation pathways. Immunol Rev 2009; 229: 322–336.
Cooper JD, Smyth DJ, Smiles AM, Plagnol V, Walker NM, Allen JE et al. Meta-analysis of genome-wide association study data identifies additional type 1 diabetes risk loci. Nat Genet 2008; 40: 1399–1401.
Manolio TA, Collins FS, Cox NJ, Goldstein DB, Hindorff LA, Hunter DJ et al. Finding the missing heritability of complex diseases. Nature 2009; 461: 747–753.
Clayton DG . Prediction and interaction in complex disease genetics: experience in type 1 diabetes. PLoS Genet 2009; 5: e1000540.
DeGobbi M, Viprakasit V, Hughes JR, Fisher C, Buckle VJ, Ayyub H et al. A regulatory SNP causes a human genetic disease by creating a new transcriptional promoter. Science 2006; 312: 1215–1217.
Nicoloso MS, Sun H, Spizzo R, Kim H, Wickramasinghe P, Shimizu M et al. Single-nucleotide polymorphisms inside microRNA target sites influence tumor susceptibility. Cancer Res 2010; 70: 2789–2798.
Abelson JF, Kwan KY, O′Roak BJ, Baek DY, Stillman AA, Morgan TM et al. Sequence Variants in SLITRK1 Are Associated with Tourette’s Syndrome. Science 2005; 310: 317–320.
Kim J, Bartel DP . Allelic imbalance sequencing reveals that single-nucleotide polymorphisms frequently alter microRNA-directed repression. Nat Biotechnol 2009; 27: 472–477.
Bao L, Zhou M, Wu L, Lu L, Goldowitz D, Williams RW et al. PolymiRTS Database: linking polymorphisms in microRNA target sites with complex traits. Nucleic Acids Res 2007; 35: D51–D54.
Long D, Lee R, Williams P, Chan CY, Ambros V, Ding Y . Potent effect of target structure on microRNA function. Nat Struct Mol Biol 2007; 14: 287–294.
Kertesz M, Iovino N, Unnerstall U, Gaul U, Segal E . The role of site accessibility in microRNA target recognition. Nat Genet 2007; 39: 1278–1284.
Oleksyk TK, Thio CL, Truelove AL, Goedert JJ, Donfield SM, Kirk GD et al. Single nucleotide polymorphisms and haplotypes in the IL10 region associated with HCV clearance. Genes Immun 2005; 6: 347–357.
Orban T, Bundy B, Becker DJ, DiMeglio LA, Gitelman SE, Goland R et al. Co-stimulation modulation with abatacept in patients with recent-onset type 1 diabetes: a randomised, double-blind, placebo-controlled trial. Lancet 2011; 378: 412–419.
Acknowledgements
This work was funded by grants from the Dutch Diabetes Research Foundation, The Netherlands, The Juvenile Diabetes Research Foundation, The Leiden University Medical Center and the 7th Framework Program of the European Commission.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on Genes and Immunity website
Supplementary information
Rights and permissions
About this article
Cite this article
de Jong, V., Zaldumbide, A., van der Slik, A. et al. Post-transcriptional control of candidate risk genes for type 1 diabetes by rare genetic variants. Genes Immun 14, 58–61 (2013). https://doi.org/10.1038/gene.2012.38
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gene.2012.38
Keywords
This article is cited by
-
Regulatory mechanisms of immune checkpoints PD-L1 and CTLA-4 in cancer
Journal of Experimental & Clinical Cancer Research (2021)
-
Type 1 diabetes mellitus as a disease of the β-cell (do not blame the immune system?)
Nature Reviews Endocrinology (2021)
-
Association of MTMR3 rs12537 at miR-181a binding site with rheumatoid arthritis and systemic lupus erythematosus risk in Egyptian patients
Scientific Reports (2019)
-
SNP rs3202538 in 3′UTR region of ErbB3 regulated by miR-204 and miR-211 promote gastric cancer development in Chinese population
Cancer Cell International (2017)
-
Autoimmunity against a defective ribosomal insulin gene product in type 1 diabetes
Nature Medicine (2017)