Abstract
Background:
KIBRA—initially identified as a neuronal associated protein is now shown to be functionally associated with other tissue types as well. KIBRA interacts with dyenin light chain 1 and this interaction is essential for oestrogen receptor transactivation in breast cancer cells. KIBRA as a substrate of Cdk1, Aurora kinase and ERK plays an important role in regulating cell cycle, cell proliferation and migration. Despite these evidences, the exact role of KIBRA in cancer progression is not known.
Methods:
We studied the expression of KIBRA in breast tissues and breast cancer cell lines by western blotting, immunohistochemisry (IHC) and RT-PCR. Stable over expression and knockdown clones were generated to study the transforming properties of KIBRA by conventional assays. Xenograft studies were performed in nude mice to study the in vivo tumourigenic efficacy of KIBRA. qPCR array was performed to understand the molecular mechanism behind oncogenic activity of KIBRA.
Results:
Our results showed that KIBRA is upregulated in breast cancer cells and in malignant human breast tumours by both western blotting and IHC. Interestingly, we found that KIBRA expression level goes up with increase in breast cancer progression in well-established MCF10A model system. Further, results from stable overexpression clones of KIBRA in fibroblasts (Rat-1) and epithelial breast cancer cells (ZR75) and lentiviral short hairpin RNA-mediated knockdown (KD) clones of KIBRA in ZR75 showed increase in transforming properties with KIBRA overexpression and vice-versa. Results also showed that fibroblasts stably overexpressing KIBRA showed increased tumourigenic potential in nude mice. By adopting a quantitative PCR array-based approach, we identified RASSF1A, a tumour suppressor, as a transcriptional target of KIBRA.
Conclusions:
This is the first study to demonstrate the in vivo tumourigenic property of KIBRA in a nude mouse model and also unravel the underlying molecular mechanism of KIBRA-mediated transformation via repression of RASSF1A.
Similar content being viewed by others
Main
Breast cancer is the most common cancer in women and is still a leading cause of cancer death worldwide. KIBRA (also called WWC1) is a multi-domain phospho-protein that is known to regulate cell polarity, cell migration and cell cycle through its interaction with different signalling molecules (Kremerskothen et al, 2003; Schneider et al, 2010; Yoshihama et al, 2012a). Further, KIBRA as a component of the Hippo signal pathway controls organ growth and cell proliferation (Yu et al, 2010). Recent data demonstrated that KIBRA is a target of kinases and that KIBRA phosphorylation is crucial for its cellular function (Büther et al, 2004; Xiao et al, 2011). KIBRA is reported to be frequently methylated in B-cell acute lymphocytic leukaemia (Hill et al, 2011). It has been shown that high KIBRA expression correlates with lymphatic invasion and poor prognosis in low atypical protein kinase C-expressing gastric cancers (Yoshihama et al, 2012b). KIBRA was reported to be upregulated in prostate tumours and also promote cell proliferation and migration in prostate cancer cells (Stauffer et al, 2016).
First evidence about the function of KIBRA in breast cancer development arose from earlier findings that the protein regulates the oestrogen receptor activity and binds to the dynein light chain 1 (DLC1) molecule (Rayala et al, 2006). Later, it was shown that KIBRA interacts with discoidin domain receptor 1 and modulates collagen-induced MAPK signalling in normal breast cells (Hilton et al, 2008). KIBRA expression is strongly upregulated during pregnancy, falls during lactation and is again high during involution of the gland at weaning, all of this could influence mammary gland development. These findings in combination with recently published observations that KIBRA as a substrate of Cdk1, Aurora kinase and ERK plays a critical role in cell cycle regulation, cell proliferation and migration (Xiao et al, 2011; Ming et al, 2012; Yang et al, 2014). It was recently reported that KIBRA modulates DNA damage response in breast cancer cells in a phosphorylation dependent manner (Mavuluri et al, 2016). All these studies point toward a crucial role for KIBRA in breast cancer development. However, despite the remarkable growth of information about the role of KIBRA in human breast cancer cells, it remains unknown whether KIBRA has any tumourigenic potential and the molecular mechanism by which KIBRA promotes oncogenic signalling in cancer cells. This prompted us to explore the oncogenic role of KIBRA in breast cancer. Here, we demonstrate that KIBRA expression is upregulated in breast cancer cell lines and breast tumours and is altered during breast tumour progression. Our in vitro and in vivo data from cell line and animal model systems respectively suggest that KIBRA plays a critical role in driving and enhancing the tumourigenic properties of breast cancer cells.
Materials and methods
Cell lines and tissues
The human breast cancer cell lines T47D, MDA MB 453, MDA MB 468, HBL 100, MCF7 and ZR75 were purchased from National Centre for Cell Science, India. MCF10A and MCF10AT were purchased from Karmanos Cancer Centre, USA. MCF10DCIS was purchased from Asterand Bioscience, USA. Rat-1 fibroblast cell line was a gift from Dr Robert A. Weinberg (Whitehead Institute for Biomedical Research, USA). MDA MB 231, SKBR3 and Hs587t were gift from Dr Asha Nair from Rajiv Gandhi Centre for Biotechnology, India. BT474 was a gift from Cancer Institute, Adayar, India. Matched breast tumour and adjacent normal tissues were obtained from Sri Ramachandra Medical College, India after getting ethical clearance. The disease status was confirmed by the pathologist.
Transfection and stable cell line
Transfection was done using FuGENE HD (Promega, Madison, WI, USA) transfection reagent according to manufacturer’s instructions. For generating stable cell lines ZR75 and Rat-1 cells were selected with blasticidin (MP Biomedicals, Santa Ana, CA, USA) after transfection at 10 μg ml−1 and puromycin (MP Biomedicals) at 2 μg ml−1. Plasmids pcDNA-6B and Flag-tagged-KIBRA were gift from Dr J. Kremerskothen, University of Muenster, Germany. pLKO.1 puro (Addgene plasmid # 8453), pCMV-dR8.2 dvpr (Addgene plasmid # 8455), pCMV-VSV-G (Addgene plasmid # 8454) were gift from Bob Weinberg (Stewart et al, 2003). pLKO.1puro-shKIBRA-A (Addgene plasmid # 40888) and pLKO.1puro-shKIBRA-B (Addgene plasmid # 40889) were gift from Paul Reynolds (Moleirinho et al, 2013). RASSF1A was cloned into pBabepuro from pClneoFH human RASSF1A, a gift from YukataHata (Addgene #37016) (Ikeda et al, 2009). BrdU cell proliferation assay kit was purchased from BD Biosciences (San Jose, CA, USA).
Immunohistochemistry
For immunohistochemistry (IHC), tissue microarrays (TMA) were purchased from US-Biomax (Rockville, MD, USA). Each core was 5 μm in thickness. IHC was done using rabbit anti-KIBRA antibody purchased from Santa Cruz Biotechnology (Dallas, TX, USA). The methodology is described in Rajhans et al (2007). In short, TMAs were deparafinised in xylene and ethanol followed by rehydration in water. The TMAs were boiled in 10 mM citrate buffer for 10 min followed by blocking with BSA (3%) for 1 h. the TMAs were then incubated with anti-KIBRA antibody overnight. The TMAs were stained using Biogenex IHC detection system according to manufacturer’s protocol. Stained arrays were scored by pathologist as Q score, where Q=P (% cells stained)* I (score based on intensity of staining). P was described as: 1=< 0–25%; 2=26–50%; 3=51–75%; 4>75%. I was described as: 1+ (low intensity), 2+ (moderate intensity) and 3+ (high intensity). As control, BSA was used as negative control instead of primary antibody.
Cells and tissue extract, western blotting and real-time PCR
Protein lysates were prepared in Radioimmunoassay precipitation buffer (RIPA) with protease inhibitor cocktail (Roche life science). RNA was isolated using TRIzol reagent (Life Technologies, Invitrogen, Carlsbad, CA, USA) according to manufacturer’s instructions. Real time PCR analysis was done for KIBRA using TaqMan probes according to manufacturer’s instructions. Actin was used as reference gene. Expression levels of KIBRA were normalised to MCF10A as reference.
Cell growth assay, wound healing assay, soft agar, clonogenic assay and confocal imaging
Cell growth assay was done as described earlier (Rayala et al, 2006). For wound healing assay, cells were grown to confluence. Scratch was made using a 20 μl tip. Cells were washed with PBS and images were captured using Carls Zeiss microscope. Soft agar assay was done, as described earlier (Jagadeeshan et al, 2014). To perform phalloidin staining, cells were cultured on coverslips. To perform staining, cells were fixed with 1% paraformaldehyde followed by treatment with 0.1% triton X-100. After treatment cells were incubated with Alexafluor 488 conjugated phalloidin (Life Technologies, Invitrogen). For cell cycle analysis, cells were fixed with ethanol and stained with propidium iodide before analysis. For BrdU cell proliferation assay protocol was followed as instructed by the manufacturer. For clonogenic assay, 500 cells were seeded and stained after 2–3 weeks with crystal violet.
Immunoprecipitation assay
For immunoprecipitaion assay, 1 mg of total protein was incubated with anti-Flag M2 affinity gel (Sigma-Aldrich, St Louis, MO, USA) overnight at 4 °C. The precipitates were washed and probed for KIBRA. Rabbit anti KIBRA antibody was purchased from Cell Signalling Technology (Danvers, MA, USA).
Animal studies
A total of 20 female nude mice (nu/nu) were purchased from Reliance Life Sciences, India after ethical clearance. Mice were divided into two groups. 5 × 106 Rat-1 cells expressing either vector or KIBRA were injected in both the mammary fat pads. Tumour volume was measured using the formula (π/6) × (l × b2), where l=length of tumour and b=breadth of tumour.
Statistical analysis
Data are represented as mean±s.e.m. Prism6 (GraphPad, USA) was used for statistical analysis.
Results
Clinical significance of kibra
KIBRA expression is upregulated in breast cancer cells and tumours
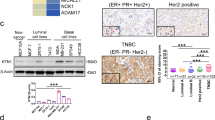
KIBRA was initially identified as a dendrin-interacting protein expressed predominantly in kidney and brain. Several emerging studies have shown that KIBRA is an important protein in cell and is expressed in various tissue types. To examine this, we initially checked the Gene enrichment profile of KIBRA in primary tissue data set from publicly available database and found that KIBRA is widely expressed in various tissues (Benita et al, 2010). Interestingly, breast tissue ranked second in gene enrichment profiling for KIBRA in various tissues. This indicates KIBRA plays an important role in breast tissue growth and development (Supplementary Figure S1). Further, we analysed the expression of KIBRA levels in a panel of 10 breast cancer cell lines and in the normal cell line MCF10A by western blot and qPCR. Results showed elevated expression of KIBRA in most of the malignant cell lines compared with the non-tumourigenic control (Figure 1A and B). Further, to examine whether the deregulation of KIBRA expression occurs during breast cancer progression, we used lysates from well characterised MCF10A model system cells to analyse KIBRA expression by western blot. The MCF10A model system contains a continuum of cell lines that enable the analysis of gene expression during the progression of breast cancer (Soule et al, 1990; Dawson et al, 1996; So et al, 2012). Western blot results revealed steady increase in the expression of KIBRA corroborating with the reported tumourigenic and metastatic potential of MCF10A series (Figure 1C). Further, in order to study the role of KIBRA in breast cancer, we analysed the expression of KIBRA protein in 17 paired samples of human breast tumours and adjacent normal breast tissue by Western blotting, and found that KIBRA levels are significantly high in 11 out of 17 tumours as compared with that in normal tissue (Figure 1D). It was interesting to note that KIBRA levels were relatively not detectable in adjacent normal tissues. These results suggest that KIBRA expression is upregulated in breast cancer cell lines and tumours.
KIBRA expression in breast cancer cell lines and tissues. (A) KIBRA expression levels were examined in breast cancer cell lines by western blotting. MCF10A was used as normal cell line for control. (B) Relative mRNA levels of KIBRA in breast cancer cell lines by real time PCR (RE: relative expression). Data are presented as mean±s.e.m. Real time PCR data were normalised to MCF10A. (C) KIBRA expression was examined in MCF10AT and MCF10DCIS in order of their tumorogenicity by western blotting. (D) KIBRA expression in matched breast tumour and adjacent normal tissues.
KIBRA expression is deregulated in breast cancer tissues
On the basis of the result that KIBRA expression levels correlated with increase in breast cancer progression in MCF10A system, we next analysed the expression of KIBRA by IHC on human breast tissue array containing 272 cores. Of this, 173 were malignant, 41 benign and 58 normal cases. Mean Q-score was used for data analysis with scores 0 for no/low staining, 1–4 for medium staining and >=6 for intense staining. KIBRA expression was found to be nuclear in majority of cases. Few samples have both the cytoplasmic, as well as nuclear positivity. The expression is predominantly high in malignant breast cancer cases (Figure 2A). On the basis of mean Q-score, malignant cases scored an average of 4.53, benign scored an average of 2.41 and normal scored an average of 1.52. The box plot (Figure 2B) denotes the quartile Q scores in the given three categories and the findings are clearly indicative of the fact that the Q scores are more in the malignant tumour samples (at least by a quarter of the samples), as compared to either normal or benign tumour tissue samples. In depth analysis revealed that a significant increase in KIBRA expression was observed in malignant type compared to normal (P value=0.0005) and benign compared to normal (P value=0.03) tissue (Table 1). In total, 86.1% cores of malignant tumour cases stained positively for KIBRA expression compared to 63.4% in benign and 43.1% in normal cases. Further analysis showed that 41% malignant cases showed an intense staining compared to 19.5 and 5.2% in benign and normal cases with a Q-score of=>6 (Table 2). While 56.9% normal cases show no/low staining, it is 36.6 and 13.9% in benign and malignant cases, respectively (Table 2). Our IHC data suggest a significant increase in KIBRA expression levels in malignant breast cancer. Altogether, these results suggest that KIBRA expression is altered during breast tumour progression and upregulated in malignant breast tumours.
KIBRA expression in human breast tumour and normal tissue sections. (A) Representative images of KIBRA expression by immunohistochemistry in tissue microarray representing normal breast, benign and malignant tumours at 200 × magnification. (B) Box plot showing the distribution of Q-score in normal, benign and malignant categories.
Transforming and tumourigenic properties of kibra
KIBRA enhances the transforming properties of breast cancer cells
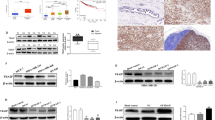
Results from the above experiments led us to investigate the oncogenic potential of KIBRA. To explore this, we generated stable clones of KIBRA overexpression in ZR75 breast cancer cell line. Stable over expression of KIBRA was confirmed by immunoprecipitation of Flag-tagged KIBRA (Figure 3A). Since KIBRA through its interaction with various signalling proteins is involved with adhesion regulation, motile and invasive phenotype, (Traer et al, 2007; Rosse et al, 2009; Yoshihama et al, 2011, 2012a), we next analysed whether KIBRA overexpression induced any changes in cell morphologic alterations indicative of a more motile phenotype. Confocal microscopic analysis revealed that modulating KIBRA levels induced long cytoplasmic extensions, increase in actin stress fibres, filopodia formation, and abnormal outgrowths in KIBRA overexpressing ZR75 cells, as compared with the vector control cells (Figure 3B). Further to provide functional proof-of-principle evidence for the observed changes in actin cytoskeleton due to KIBRA overexpression, we used an established wound-healing assay to assess the migration of ZR75 clones. Consistent with the confocal results, in vitro wound healing assay and boyden chamber migration assay results showed that KIBRA overexpression leads to improved wound healing and migration rates as compared with the control vector ZR75 cell line (Figure 3C and D). Further, to strengthen the role of KIBRA in enhancing the transforming properties of breast cancer cells, we performed conventional cellular transforming assays with the KIBRA overexpressing ZR75 cells and found that KIBRA overexpression significantly enhanced the cell proliferation capability, increased the ability to form more colonies in anchorage independent soft agar and clonogenic assays (Figure 4A–C). Since KIBRA was shown to regulate mitosis (Xiao et al, 2011; Ming et al, 2012), we tested if KIBRA overexpression affects the cell cycle profile. Results showed a significant increase in the S phase population (Figure 4D). This increase in S phase population was further confirmed by BrdU cell proliferation assay which showed an increase in proliferation of KIBRA overexpressing clones upon incubation with BrdU (Figure 4E).
KIBRA promotes migration in ZR75 breast cancer cells. (A) Characterisation of ZR75 cell line expressing vector or KIBRA by immunoprecipitation of Flag tag and probed with KIBRA antibody. (B) Representative images of confocal microscopy for actin staining by phalloidinAlexa flour in ZR75 cells expressing vector or KIBRA. Arrow head represents the filopodia structures. (C) Representative images of wound healing assay at 0 h and 24 h. (*P value=0.027; unpaired t test with Welch’s correction). (D) Representative images of migration assay with ZR75 cells overexpressing vector or KIBRA. (**P value=0.0078, unpaired t test with Welch’s correction).
KIBRA promotes cell transformation properties in ZR75 cell line. (A) Cell proliferation rate was measured every 24 h for 6 days (**P value<0.005; ****P value<0.00005; two-way ANOVA with Sidak’s multiple comparisions test). (B and C) Representative images of soft agar and clonogenic assay with bar graph representing the number of colonies. (****P value<0.0001 and **P value=0.0067, unpaired t test with Welch’s correction). (D and E) Bar graphs showing cell cycle analysis and BrdU cell proliferation assay results. (**P value=0.0029, *P value=0.0342; unpaired t test with Welch’s correction).
Collectively, all these results suggest that KIBRA plays a critical role in enhancing tumourigenic properties of breast cancer cells.
KIBRA induces transforming property to Rat-1 fibroblast cell line
To further confirm the tumourigenic ability of KIBRA, we used Rat-1 fibroblasts. Rat-1 cell line is a well-established model system to study various oncogenic properties (Reynolds et al, 1987; LaMontagne et al, 1998; Young et al, 1998). We established stable clones of KIBRA overexpression in Rat-1 fibroblast cell line. Stable overexpression of KIBRA was confirmed by immunoprecipitation of Flag-tagged KIBRA (Figure 5A). Consistent with our results with ZR75 clones, KIBRA overexpressing Rat-1 clones also showed significant increase in cell proliferation (Figure 5B), enhanced wound healing (Figure 5C) and increased ability to form more colonies in clonogenic cell survival assay as compared to control vector Rat-1 clone (Figure 5D). These results suggest that KIBRA induces tumourigenic potential to fibroblast cells.
KIBRA promotes cell transformation properties in Rat-1 fibroblast cell line and tumour formation in nude mice. (A) Characterisation of Rat-1 fibroblast cell line expressing vector or KIBRA by immunoprecipitation with flag tag. (B) Cell growth was measured every 24 h for 6 days (*P<0.05; ****P<0.0001; two-way ANOVA with Sidak’s multiple comparision test). (C and D) Representative images of wound healing and clonogenic assay with bar graph. (*P value=0.0092 and *P value=0.0015, unpaired t test with Welch’s correction). Data represent the mean±s.e.m of minimum three independent experiments. (E) Representative images showing the tumour formation in nude mice with bar graph showing the tumour volume. (**P value=0.004, unpaired t test with Welch’s correction).
Tumourigenic potential of KIBRA in nude mouse model
Further to investigate the KIBRA’s ability to induce tumourigenic potential in vivo, we used a nude mouse xenograft model. Rat-1 cells stably expressing vector (n=8) or KIBRA (n=8) were injected s.c. into mice and tumourigenic potential was monitored. Tumours were visible after two weeks in mice injected with KIBRA overexpressing cells. All 8/8 mice with KIBRA overexpression developed tumours. Relatively small tumour was visible in two mice injected with vector control cells (Figure 5E). Tumour measurements showed that there is a significant increase in tumour growth in KIBRA overexpression group as compared to vector control group (Supplementary Table 1). Collectively, these in vivo results were consistent with our in vitro results—indicating the importance of KIBRA in driving the tumourigenic property.
KIBRA knockdown inhibits the cell survival and proliferation of breast cancer cells
Our above results demonstrated that KIBRA overexpression enhances cell proliferation, migration, cell survival and anchorage independence of breast cancer ZR75 and Rat-1 fibroblast cells. Next, we asked, if KIBRA knockdown has any effect on the tumourigenic potential of breast cancer cells. For this, we developed stable lentiviral short hairpin RNA-mediated knockdown (shKIBRA) clones of KIBRA in ZR75 cells (Figure 6A and Supplementary Figure S3) and studied the changes in transforming properties of these cells. In the colony formation assay, we observed significantly fewer number of colonies formed in KIBRA downregulated clones as compared with the control clone (Figure 6B). We also observed a decrease in proliferation of KIBRA shRNA clone upon incubation with BrdU (Figure 6C).
KIBRA knockdown inhibits cell transforming properties in ZR75 cells and induces RASSF1 expression. (A) Western blot showing the KIBRA knockdown. (B) Representative image and bar graph of clonogenic assay with vector and shKIBRAA clones. (****P value< 0.0001, unpaired t test with Welch’s correction) (C) Bar graph for BrdU cell proliferation assay results performed in vector or shKIBRAA clones. (D) Heat map for the qPCR array performed with KIBRA overexpression clones in ZR75 breast cancer cell line. (E) Bar graph (qRT-PCR) and western blots showing RASSF1A expression in ZR75 cells expressing vector or KIBRA. (RE: relative expression; **P value=0.0016, unpaired t test with Welch’s correction) (F) Western blot for RASFF1A expression in vector or shRNA knockdown ZR75 cells. (G) Representative image and bar graph for the clonogenic assay showing the effect of ectopic expression of RASSF1A in vector or KIBRA expressing ZR75 cells. (***P value<0.0005, one way ANOVA with Tukey’s multiple comparison test). (H) Proposed mechanism of action of KIBRA by regulating RASSF1A expression.
Molecular mechanism of kibra induced tumourigenesis
RASSF1A – A novel molecular target of KIBRA
The evidence from the above results suggested that KIBRA promotes oncogenic properties in cells. To identify the downstream molecular targets that might be regulated through KIBRA signalling and also associated with the tumourigenesis, we performed qPCR expression profiling of 84 genes associated with tumourigenesis utilising KIBRA overexpressing ZR75 clones (Figure 6D). A complete list of genes analysed through the gene array and the genes that are modulated by KIBRA is given as Supplementary Table 2. The shortlisted genes that are modulated by KIBRA are given as Supplementary Table 3. As determined by this method, we identified RASSF1A as one of the potential target that was drastically downregulated in the KIBRA overexpressing clone as compared with vector control. We further validated RASSF1A in overexpression model of ZR75 using individual qPCR assays and immunoblotting for RASSF1A. Results showed that RASSF1A mRNA and protein levels were significantly decreased in KIBRA overexpressing ZR75 cell line as compared with the empty vector transfect (Figure 6E). Similarly, there is a significant increase in the RASSF1A protein levels in the stable shRNA clone of KIBRA in ZR75 cells (Figure 6F). Further, to strengthen the mechanistic studies implicating the role of RASSF1A in KIBRA functions, we overexpressed RASSF1A in ZR75 cells overexpressing either vector control or KIBRA and performed clonogenic survival assay. Results showed that there is a significant reduction in tumourigenic property of KIBRA upon overexpressing RASSF1A (Figure 6G and Supplementary Figure S4).
Discussion
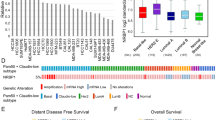
With the advent of herceptin and other targeted therapeutics for cancer, research into the discovery of novel biomarkers for advanced breast cancer is in full swing. Although the progression from early to metastatic stage breast cancer is a critical determinant of survival, little is known about the molecules that contribute to the progression and metastasis of breast tumours. In this study, we identified that KIBRA plays a role in breast cancer progression and its expression is deregulated in breast tumours. Using a panel of breast cancer cells, fibroblasts, human breast tissue samples, and mouse xenograft models, we have provided evidence that modulating KIBRA expression contributes to and enhances oncogenic property to breast cancer cells. In addition, progressively increased expression of KIBRA in MCF10A model system with distinct invasive and metastatic potential revealed the functional importance of KIBRA during breast cancer progression. In support of this, we determined the KIBRA expression in human breast tumour samples, benign breast diseases and normal breast epithelium and found that KIBRA expression and localisation are deregulated in breast tumours. The expression of KIBRA was predominantly high in malignant breast tumour tissue samples, and was nuclear in majority of samples; only a few samples showed both nuclear and cytoplasmic localisation. In support of our results, it was interesting to note that preliminary data (10 cores of breast carcinoma) from IHC staining from Protein Atlas database also revealed an enhanced expression (9 out of 10 showed medium to high) of KIBRA in breast cancer tissues and especially ductal breast carcinoma showed high levels of KIBRA (Supplementary Figure S2; Uhlén et al, 2005; Berglund et al, 2008; Uhlén et al, 2015). It was shown that in low atypical protein kinase C expressing gastric cancers higher KIBRA expression level correlates with increased lymphatic invasion and poor prognosis (Yoshihama et al, 2012b). KIBRA expression was also reported to be high in prostate tumours (Stauffer et al, 2016). Previous reports have shown that nuclear localisation of KIBRA has functional significance in deregulating cell division and cell cycle (Ming et al, 2012; Zhang et al, 2012). It is known that cancer cells show excessive proliferation rates as a manifestation of defective cell cycle and cell division. Collectively, this could mean that there exists a strong correlation between higher KIBRA expressions with nuclear localisation and aggressiveness of breast cancer.
We observed that KIBRA modulated dynamic changes in cytoplasm were translated into filamentous actin cytoskeletal rearrangements, adoption of motile cell phenotypes, and increased ability to migrate. This is supported by the report that KIBRA has been shown to be present at leading edge and acts as a scaffold protein for the efficient migration of cells (Rosse et al, 2009).
Further, we observed an increase in proliferation, anchorage independence and cell survival on KIBRA overexpression in cells. This trend is reversed on knockdown of KIBRA. These findings suggest that KIBRA plays an essential role in attributing to the transforming and tumourigenic potential of breast cancer cells. This is in agreement with an earlier study where KIBRA knockdown has been demonstrated to reduce the proliferation in MCF7 and MDA MB 231 breast cancer cell lines (Rayala et al, 2006; Yang et al, 2014). This is further supported by the evidence that KIBRA overexpression enhances proliferation and migration in prostate cancer cells (Stauffer et al, 2016). This was supported by the finding that KIBRA expression is decreased in a mouse model of failed mammary gland proliferation indicating importance of KIBRA in cell proliferation and mammary gland development (Hilton et al, 2008). It is well established that ERK activation promotes the cell proliferation. It was shown that overexpression of KIBRA increases ERK activation in a collagen dependent manner. KIBRA was reported to be a substrate for ERK-RSK cascade and thus activated KIBRA modulates cell proliferation, migration and invasion (Yang et al, 2014).
KIBRA has been reported to be necessary for chromosomal alignment and centrosome formation during mitosis and its knockdown lead to chromosomal misalignment and spindle defects (Xiao et al, 2011; Ming et al, 2012; Zhang et al, 2012). KIBRA is also reported to be a phospho-protein and its phosphorylation by CDK1 has been shown to regulate the cell cycle (Ming et al, 2012). Our results also indicate that KIBRA can induce cell proliferation by enhancing the S phase. This was also supported by an earlier report where KIBRA has been shown to induce Cyclin D1 and Bcl2 levels in an oestrogen dependent manner (Rayala et al, 2006). Further, tumour xenograft studies from our KIBRA overexpressing Rat-1 fibroblasts provided evidence for the transforming role of KIBRA. This is the first study to demonstrate the in vivo tumourigenic property of KIBRA in a nude mouse model.
It was shown that KIBRA is a positive regulator of Hippo pathway activity in mammalian cells suggesting that KIBRA functions as a tumour suppressive protein as it does in Drosophila (Yu et al, 2010; Genevet et al, 2010). Consistent with these observations, a recent study showed that KIBRA knockdown enhanced migration and invasion of immortalised breast epithelial cells (Moleirinho et al, 2013). The gene expression data analysis also showed that KIBRA expression is decreased in Claudin-low primary breast cancers. However, our current study shows that KIBRA overexpression significantly increases cell proliferation, migration and anchorage independent growth of ZR75 cells (Figures 3 and 4) and KIBRA is highly expressed in malignant breast tumours (Figures 1 and 2). This contrasts to the study in immortalised non-cancerous cells (Moleirinho et al, 2013) and our IHC data for KIBRA does not exclusively take any particular subtype into consideration. Interestingly, several studies have also demonstrated that KIBRA positively regulates cell migration and proliferation (Rosse et al, 2009; Yang et al, 2014; Stauffer et al, 2016). Thus, the tumour suppressive function of KIBRA could be tissue/cell type specific and KIBRA could function as a positive regulator of cell proliferation, cell motility and invasion depending on the context. Detailed parallel studies with different tissue/cell types are required to get a clear picture on resolving this dual functionality. Our study is consistent with these later studies, supporting a positive role of KIBRA in proliferation and migration in breast cancer cells.
Our efforts to unravel the underlying molecular mechanism of KIBRA-mediated transformation by using qPCR array identified 5 genes (BAD, CDH1, ESR2, RASSF1A and TP53) which were downregulated by KIBRA overexpression. Only RASSF1A was found to be consistently and significantly downregulated. This suggests that KIBRA acts by regulating RASSF1A expression. RASSF1A is a well-known tumour suppressor gene in several cancers. It is known to be inactivated by hypermethylation of its promoter in majority of cancers. It has been shown that RASSF1A promoter is hypermethylated in invasive breast cancer and DCIS (Agathanggelou et al, 2001; Honorio et al, 2003; Donninger et al, 2007). RASSF1A is shown to reduce the clonogenic survival of cancer cells (Whang et al, 2005, 2009; Thaler et al, 2012). Our results indicated that KIBRA inhibits RASSF1A expression and thereby promotes tumourigenesis. Given the fact that KIBRA is localised to the nucleus (this study) (Rayala et al, 2006; Mavuluri et al, 2016) and interacts with histones, (Rayala et al, 2006), it is quite possible that KIBRA somehow might be acting as transcriptional repressor for RASSF1A via modulating its promoter methylation. This ultimately leads to inactivation of tumour suppressor RASSF1A and this could be the possible mechanism by which KIBRA contributes to tumourigenesis (Figure 6H).
Change history
08 August 2017
This paper was modified 12 months after initial publication to switch to Creative Commons licence terms, as noted at publication
References
Agathanggelou A, Honorio S, MacartneyDP, Martinez D, Dallol A, Rader J, Fullwood P, Chauhan A, Walker R, Shaw JA, Hosoe S, Lerman MI, Minna JD, Maher ER, Latif F (2001) Methylation associated inactivation of RASSF1A from region 3p21.3 inlung, breast and ovarian tumours. Oncogene 20: 1509–1518.
Benita Y, Cao Z, Giallourakis C, Li C, Gardet A, Xavier RJ (2010) Gene enrichment profiles reveal T-cell development, differentiation, and lineage-specific transcription factors including ZBTB25 as a novel NF-AT repressor. Blood 115 (26): 5376–5384.
Berglund L, Björling E, Oksvold P, Fagerberg L, Asplund A, Szigyarto CA, Persson A, Ottosson J, Wernérus H, Nilsson P, Lundber E, Sivertsson A, Navani S, Wester K, Kampf C, Hober S, Pontén F, Uhlén M (2008) A genecentric Human Protein Atlas for expression profiles based on antibodies. Mol Cell Proteomics 7 (10): 2019–2027.
Büther K, Plaas C, Barnekow A, Kremerskothen J (2004) KIBRA is a novel substrate for protein kinase Cζ. Biochem Biophys Res Commun 317 (3): 703–707.
Dawson PJ, Wolman SR, Tait L, Heppner GH, Miller FR (1996) MCF10AT: a model for the evolution of cancer from proliferative breast disease. Am J Pathol 148 (1): 313–319.
Donninger H, Michele DV, Clark GJ (2007) The RASSF1A tumor suppressor. J Cell Sci 120: 3163–3172.
Genevet A, Wehr MC, Brain R, Thompson BJ, Tapon N (2010) Kibra is a regulator of the Salvador/Warts/Hippo signaling network. Dev Cell 18 (2): 300–308.
Hill VK, Dunwell T, Catchpoole D, Krex D, Brini AT, Griffiths M, Craddock C, Maher ER, Latif F (2011) Frequent epigenetic inactivation of KIBRA, an upstream member of the Salvador/Warts/Hippo (SWH) tumor suppressor network, is associated with specific genetic event in B-cell acute lymphocytic leukemia. Epigenetics 6 (3): 326–332.
Hilton HN, Stanford PM, Harris J, Oakes SR, Kaplan W, Daly RJ, Ormandy CJ (2008) KIBRA interacts with discoidin domain receptor 1 to modulate collagen-induced signalling. Biochim Biophys Acta 1783 (3): 383–393.
Honorio S, Agathaggagenlou A, Schuermann M, Pankow W, Viacava P, Maher ER, Latif F (2003) Detection of RASSF1A aberrant promoter hypermethylation in sputum from chronic smokers and ductal carcinoma in situ from breast cancer patients. Oncogene 22: 147–150.
Ikeda M, Kawata A, Nishikawa M, Tateishi Y, Yamaguchi M, Nakagawa K, Hirabayashi S, Bao Y, Hidaka S, Hirata Y, Hata Y (2009) Hippo pathway–dependent and –independent roles of RASSF6. Sci Signal 2 (90): 1–11.
Jagadeeshan S, Krishnamoorthy YR, Singhal M, Subramanian A, Mavuluri J, Lakshmi A, Roshini A, Baskar G, Ravi M, Joseph LD, Sadasivan K, Krishnan A, Nair AS, Venkatraman G, Rayala SK (2014) Transcriptional regulation of fibronectin by p21-activated kinase-1 modulates pancreatic tumourigenesis. Oncogene 34 (4): 455–464.
Kremerskothen J, Plaas C, Büther K, Finger I, Veltel S, Matanis T, Liedtke T, Barnekow A (2003) Characterization of KIBRA, a novel WW domain-containing protein. Biochem Biophys Res Commun 300 (4): 862–867.
LaMontagne KR, Hannon G, Tonks NK (1998) Protein tyrosine phosphatase PTP1B suppresses p210 bcr-abl-induced transformation of rat-1 fibroblasts and promotes differentiation of K562 cells. Proc Natl Acad Sci 95 (24): 14094–14099.
Mavuluri J, Beesetti S, Surabhi R, Kremerskothen J, Venkatraman G, Rayala SK (2016) Phosphorylation-dependent regulation of the DNA damage response of adaptor protein KIBRA in cancer cells. Mol Cell Biol 36: 1354–1365.
Ming JI, Yang S, Chen Y, Xiao L, Zhang L, Dong J (2012) Phospho-regulation of KIBRA by CDK1 and CDC14 phosphatase controls cell-cycle progression. Biochem J 447 (1): 93–102.
Moleirinho S, Chang N, Sims AH, Tilston-Lünel AM, Angus L, Steele A, Boswell V, Barnett SC, Ormandy C, Faratian D, Gunn-Moore FJ, Reynolds PA (2013) KIBRA exhibits MST-independent functional regulation of the Hippo signaling pathway in mammals. Oncogene 32 (14): 1821–1830.
Rajhans R, Nair S, Holden AH, Kumar R, Tekmal RR, Vadlamudi RK (2007) Oncogenic potential of the nuclear receptor coregulator proline-, glutamic acid–, leucine-rich protein 1/modulator of the nongenomic actions of the estrogen receptor. Cancer Res 67 (11): 5505–5512.
Rayala SK, den Hollander P, Manavathi B, Talukder AH, Song C, Peng S, Barnekow A, Kremerskothen J, Kumar R (2006) Essential role of KIBRA in co-activator function of dynein light chain 1 in mammalian cells. J Biol Chem 281 (28): 19092–19099.
Reynolds VL, DiPietro M, Lebovitz RM, Lieberman MW (1987) Inherent tumourigenic and metastatic properties of rat-1 and rat-2 cells. Cancer Res 47 (23): 6384–6387.
Rosse C, Formstecher E, Boeckeler K, Zhao Y, Kremerskothen J, White MD, Camonis JH, Parker PJ (2009) An aPKC-exocyst complex controls paxillin phosphorylation and migration through localised JNK1 activation. PLoS Biol 7 (11): e1000235.
Schneider A, Huentelman MJ, Kremerskothen J, Duning K, Spoelgen R, Nikolich K (2010) KIBRA: a new gateway to learning and memory? Front Aging Neurosci 2: 1–9.
So JY, Lee HJ, Kramata P, Minden A, Suh N (2012) Differential expression of key signaling proteins in MCF10 cell lines, a human breast cancer progression model. Mol Cell Pharmacol 4 (1): 31–40.
Soule HD, Maloney TM, Wolman SR, Peterson WD, Brenz R, McGrath CM, Russo J, Pauley RJ, Jones RF, Brooks SC (1990) Isolation and characterization of a spontaneously immortalized human breast epithelial cell line, MCF-10. Cancer Res 50 (18): 6075–6086.
Stauffer S, Chen X, Zhang L, Chen Y, Dong J (2016) KIBRA promotes prostate cancer cell proliferation and motility. FEBS J 283: 1800–1811.
Stewart SA, Dykshoorn DM, Palliser D, Mizuno H, Yu EY, An DS, Sabatini DM, Chen IS, Hahn WC, Sharp PA, Weinberg RA, Novina CD (2003) Lentivirus-delivered stable gene silencing by RNAi in primary cells. RNA 9: 493–501.
Thaler S, Schmidt M, Schad A, Sleeman JP (2012) RASSF1A inhibits estrogen receptor alpha expression andestrogen-independent signalling: implications for breast cancer development. Oncogene 31: 4912–4922.
Traer CJ, Rutherford AC, Palmer KJ, Wassmer T, Oakley J, Attar N, Carlton JG, Kremerskothen J, Stephens DJ, Cullen PJ (2007) SNX4 coordinates endosomal sorting of TfnR with dynein-mediated transport into the endocytic recycling compartment. Nat Cell Biol 9 (12): 1370–1380.
Uhlén M, Björling E, Agaton C, Szigyarto CA-K, Amini B, Andersen E, Andersson A-C, Angelidou P, Asplund A, Asplund C, Berglund L, Bergström K, Brumer H, Cerjan D, Ekström M, Elobeid A, Eriksson C, Fagerberg L, Falk R, Fall J, Forsberg M, Björklund MG, Gumbel K, Halimi A, Hallin I, Hamsten C, Hansson M, Hedhammar M, Hercules G, Kampf C, Larsson K, Lindskog M, Lodewyckx W, Lund J, Lundeberg J, Magnusson K, Malm E, Nilsson P, Odling J, Oksvold P, Olsson I, Oster E, Ottosson J, Paavilainen L, Persson A, Rimini R, Rockberg J, Runeson M, Sivertsson A, Sköllermo A, Steen J, Stenvall M, Sterky F, Strömberg S, Sundberg M, Tegel H, Tourle S, Wahlund E, Waldén A, Wan J, Wernérus H, Westberg J, Wester K, Wrethagen U, Xu LL, Hober S, Pontén F (2005) A human protein atlas for normal and cancer tissues based on antibody proteomics. Mol Cell Proteomics 4 (12): 1920–1932.
Uhlén M, Fagerberg L, Hallström BM, Lindskog C, Oksvold P, Mardinoglu A, Sivertsson Å, Kampf C, Sjöstedt E, Asplund A, Olsson I, Edlund K, Lundberg E, Navani S, Szigyarto CA, Odeberg J, Djureinovic D, Takanen JO, Hober S, Alm T, Edqvist PH, Berling H, Tegel H, Mulder J, Rockberg J, Nilsson P, Schwenk JM, Hamsten M, von Feilitzen K, Forsberg M, Persson L, Johansson F, Zwahlen M, von Heijne G, Nielsen J, Pontén F (2015) Tissue-based map of the human proteome. Science 347 (6220): 1260419.
Whang YM, Kim YH, Kim JS, Yoo YD (2005) RASSF1A suppresses the c-Jun-NH2-kinase pathway and inhibits cell cycle progression. Cancer Res 65 (9): 3682–3690.
Whang YM, Park KH, Jung HY, Jo UH, Kim YH (2009) Microtubule-damaging agents enhance RASSF1A-induced cell death in lung cancer cell lines. Cancer 115: 1253–1266.
Xiao L, Chen Y, Ji M, Volle DJ, Lewis RE, Tsai M-Y, Dong J (2011) KIBRA protein phosphorylation is regulated by mitotic kinase aurora and protein phosphatase 1. J Biol Chem 286 (42): 36304–36315.
Yang S, Ji M, Zhang L, Chen Y, Wennmann DO, Kremerskothen J, Dong J (2014) Phosphorylation of KIBRA by the extracellular signal-regulated kinase (ERK)-ribosomal S6 kinase (RSK) cascade modulates cell proliferation and migration. Cell Signal 26 (2): 343–351.
Yoshihama Y, Chida K, Ohno S (2012a) The KIBRA-aPKC connection: a potential regulator of membrane trafficking and cell polarity. Commun Integr Biol 5 (2): 146–151.
Yoshihama Y, Izumisawa Y, Akimoto K, Satoh Y, Mizushima T, Satoh K, Chida K, Takagawa R, Akiyama H, Ichikawa Y (2012b) High expression of KIBRA in low atypical protein kinase C-expressing gastric cancer correlates with lymphatic invasion and poor prognosis. Cancer Sci 104 (2): 259–265.
Yoshihama Y, Sasaki K, Horikoshi Y, Suzuki A, Ohtsuka T, Hakuno F, Takahashi S-I, Ohno S, Chida K (2011) KIBRA suppresses apical exocytosis through inhibition of aPKC kinase activity in epithelial cells. Curr Biol 21 (8): 705–711.
Young CS, Kitamura M, Hardy S, Kitajewski J (1998) Wnt-1 induces growth, cytosolic β-catenin, and Tcf/Lef transcriptional activation in Rat-1 fibroblasts. Mol Cell Biol 18 (5): 2474–2485.
Yu J, Zheng Y, Dong J, Klusza S, Deng W-M, Pan D (2010) Kibra functions as a tumor suppressor protein that regulates Hippo signaling in conjunction with Merlin and Expanded. Dev Cell 18 (2): 288–299.
Zhang L, Iyer J, Chowdhury A, Ji M, Xiao L, Yang S, Chen Y, Tsai M-Y, Dong J (2012) KIBRA regulates aurora kinase activity and is required for precise chromosome alignment during mitosis. J Biol Chem 287 (41): 34069–34077.
Acknowledgements
We thank Dr Vijayalakshmi, Cancer Institute, Adyar, Chennai, India for helping us in IHC staining. We thank Mr Rakesh Nankar and Mr Shabeer Ahmed for help with the statistical analysis and valuable suggestions. We thank Mr Shashi Bhushan for capturing the images and Ms. Sansrity Sinha for helping in preparing figure panels for the manuscript. Suggestions and help received from all the students of IIT Madras and Sri Ramachandra University is appreciated. We thank DBT and DST (Grant # BT/IN/01/GERMAN/RSK/2010 and DST/INT/SL/P-09), Government of India for financial support to SKR and Indian Institute of Technology Madras (IITM) for all other facilities.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
This work is published under the standard license to publish agreement. After 12 months the work will become freely available and the license terms will switch to a Creative Commons Attribution-NonCommercial-Share Alike 4.0 Unported License.
Supplementary Information accompanies this paper on British Journal of Cancer website
Rights and permissions
From twelve months after its original publication, this work is licensed under the Creative Commons Attribution-NonCommercial-Share Alike 4.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Anuj, Arivazhagan, L., Surabhi, R. et al. KIBRA attains oncogenic activity by repressing RASSF1A. Br J Cancer 117, 553–562 (2017). https://doi.org/10.1038/bjc.2017.192
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bjc.2017.192
Keywords
This article is cited by
-
KIBRA Team Up with Partners to Promote Breast Cancer Metastasis
Pathology & Oncology Research (2020)
-
KIBRA; a novel biomarker predicting recurrence free survival of breast cancer patients receiving adjuvant therapy
BMC Cancer (2018)