Abstract
Developments in genomic techniques have provided insight into the remarkable genetic complexity of malignant tumours. There is increasing evidence that solid tumours may comprise of subpopulations of cells with distinct genomic alterations within the same tumour, a phenomenon termed intra-tumour heterogeneity. Intra-tumour heterogeneity is likely to have implications for cancer therapeutics and biomarker discovery, particularly in the era of targeted treatment, and evidence for a relationship between intra-tumoural heterogeneity and clinical outcome is emerging. Our understanding of the processes that exacerbate intra-tumoural heterogeneity, both iatrogenic and tumour specific, is likely to increase with the development and more widespread implementation of advanced sequencing technologies, and adaptation of clinical trial design to include comprehensive tissue collection protocols. The current evidence for intra-tumour heterogeneity and its relevance to cancer therapeutics will be presented in this mini-review.
Similar content being viewed by others
Main
Tumour heterogeneity refers to the existence of subpopulations of cells, with distinct genotypes and phenotypes that may harbour divergent biological behaviours, within a primary tumour and its metastases, or between tumours of the same histopathological subtype (intra- and inter-tumour, respectively). With the advent of deep sequencing techniques, the extent and prevalence of intra- and inter-tumour heterogeneity is increasingly acknowledged. There are features of intra-tumour heterogeneity that form part of routine pathologic assessment, but its determination does not yet form part of the clinical decision-making process. This mini-review aims to summarise the evidence supporting the extent, causes and consequences of intra-tumour heterogeneity, and will suggest how this knowledge may be integrated into future clinical practice and research efforts to optimise patient care and clinical outcomes. Inter-tumour heterogeneity is well described in breast cancer and other solid tumours, but is beyond the scope of this article (for a comprehensive review, see Russnes et al, 2011).
The spectrum of cancer heterogeneity
Clinico-pathological heterogeneity and its molecular basis
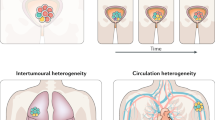
Morphological variation between different regions of a tumour has long been familiar to histopathologists. For this reason, it is routine for pathologists to examine multiple sections from the same tumour, but to report only the highest grade. Nuclear pleomorphism is another example of intra-tumour heterogeneity, which is accounted for in breast cancer grading. It is also readily apparent to those clinicians treating cancer that there is marked variation in tumour behaviour between patients with the same tumour type, and between different tumour sites in the same patient; the latter is usually manifested as differential or mixed responses to therapy (Figure 1). Intuitively, common clinico-pathological observations such as these could be attributable to intra-tumour heterogeneity, but studies are only now beginning to formally evaluate this relationship. It is also likely that other factors, such as pharmacodynamics, contribute to the non-uniformity of drug response.
Serial computed tomography (CT) scans from a patient treated with the mammalian target of rapamycin (mTOR) inhibitor everolimus for metastatic renal cell carcinoma demonstrate a reduction in the size of a metastasis at the medial right lung base (A) but an increase in the size of a metastatic lesion in the left retroperitoneum (B). A differential response such as this may reflect the presence of tumour heterogeneity, or altered pharmacodynamics at separate disease sites.
Initial proof that multiple subclones of differing genetic status existed within the same tumour was provided by G-banding karyotyping and fluorescent in situ hybridisation (FISH) studies, demonstrating discrete patterns of chromosomal rearrangements and copy number alterations of representative genomic loci (reviewed in Navin and Hicks, 2010). Completion of the human genome project was a prelude to understanding the true genetic complexity underlying cancer, providing the opportunity to systemically sequence somatic coding and non-coding aberrations on a large scale.
Although other forms of intra-tumoural heterogeneity exist, such as phenotypic heterogeneity, this review will focus on the impact of genetic intra-tumour heterogeneity on treatment stratification and therapeutic outcome.
Clonal evolution as a model of tumour progression and heterogeneity
A clonal evolutionary model of cancer development was first proposed by Nowell (1976) and elaborates upon Darwinian models of natural selection – that is, genetically unstable cells accumulate genetic alterations, and that selective pressures favour the growth and survival of variant subpopulations with a biological fitness advantage. This model, in partnership with cancer genomic instability mechanisms exacerbating structural and numerical chromosomal instability (McGranahan et al, 2012), might contribute to the presence of molecular heterogeneity within tumours, which increases the pool of genetic variants to be tested by selection and thus increase the chance that a subclone will have a growth and/or survival advantage. The concept of ‘driver’ and ‘passenger’ mutations is an important component of this model: driver somatic mutations are those that increase the fitness of the cell, allowing this cell lineage to populate the tumour, whereas passenger mutations are neutral or deleterious mutations that persist because they are genetically linked to driver mutations (for a detailed review, see Sprouffske et al, 2012). While driver mutations are central to our current understanding of the genetic basis of human cancer, it is our view that both types of mutation have direct implications for cancer therapeutics, because the heterogeneous events may determine resistance outgrowth and thus patient demise, discussed later in this review.
Clonal evolution as a basis for tumour progression has been clearly demonstrated in recent studies of haematological cancers, and in brain, breast, and pancreatic tumours (Sidransky et al, 1992; Shah et al, 2009; Campbell et al, 2010; Yachida et al, 2010; Ding et al, 2012; Schuh et al, 2012; Walter et al, 2012). Sequencing analysis of paired primary tumour and relapse genomes from eight patients with acute myeloid leukaemia (AML) revealed a founding clone was present in the primary tumour, which had evolved and/or expanded in the relapsed tumour (Ding et al, 2012). In patients with transformed myelodysplastic syndromes, matched samples from the antecedent myelodysplastic stage and the secondary AML stage were also clonal; transformation to AML was associated with acquisition of many new mutations in a founding clone to form a daughter subclone (Walter et al, 2012). Another model for tumour progression, the cancer stem cell (CSC) hypothesis, may provide a complementary and not necessarily mutually exclusive explanation for intra-tumour heterogeneity. For a detailed discussion on the relationship between the CSC hypothesis and intra-tumour heterogeneity, readers are directed to reviews by Navin and Hicks (2010) and Sprouffske et al (2012).
Tumour heterogeneity: a dynamic state
Spatial and temporal heterogeneity may permit the tumour as a whole to adapt to a fluctuating tumour microenvironment. There are intriguing clinical examples of variation in temporal patterns of tumour genomic architectures within multiple myeloma patients, and in some patients a distinct pattern of complex clonal competition was observed (Keats et al, 2012). In one cytogenetically high-risk patient, serial samples were available for the entire disease course, enabling longitudional copy number analysis. This study demonstrated two major clonal progenitors and subsequent divergence. At different time points, the suppression and reappearance of clones appeared to correlate with timing of drug therapy but the plasma cell leukaemic clone 2.2, which determined patient death, was hardly detectable at the outset, raising clear problems for predictive or prognostic biomarker strategies at diagnosis. A similar finding was recently reported in patients with chronic lymphocytic leukaemia (Schuh et al, 2012). Our further interpretation of the experimental data is that two subclones can exist in a ‘dynamic equilibrium’, constantly competing for clonal dominance in the context of systemic therapy. A third study using whole genome sequencing in a patient whose disease had transformed from multiple myeloma to plasma cell leukaemia indicated the presence of subclones at diagnosis, which waxed and waned in dominance as the cancer progressed (Egan et al, 2012).
Clonal cooperativity has also been reported; Inda et al (2010) showed in vitro and in vivo that in glioblastoma, the mixture of epidermal growth factor receptor (EGFR)-mutant and EGFR-wild type glioma cells enhanced tumour growth. This occurred through a paracrine mechanism, in which EGFR-mutant cells express cytokines such as IL-6 and leukaemia inhibitor factor that activate EGFR-wild type, driving their proliferation.
In summary, it is argued that heterogeneous tumours should be viewed as complex ecosystems or societies, in which even a minor tumour subpopulation may influence growth of the entire tumour and thereby actively maintain tumour heterogeneity (Heppner, 1984; Marusyk and Polyak, 2010; Bonavia et al, 2011). In this model, subclones occupy various niches within the tumour microenvironment and the survival advantage of the tumour ‘society’ exceeds those of the individual subpopulation; relationships between subclones may be competitive, commensal, or mutualistic for this purpose.
Current evidence for intra-tumour heterogeneity
Next-generation sequencing (NGS) technologies are adding new evidence for genetic diversity both within and between common tumours. NGS methodologies comprehensively and systematically determine both nucleotide sequence and copy number of genomic loci, and have the advantage of being able to simultaneously sequence heterogeneous mixtures of genomes in a given sample, which might include tumour, stromal, and immune cells.
Both NGS and comparative genomic hybridisation studies in breast cancer suggest marked genetic intra-tumour heterogeneity (Torres et al, 2007; Shah et al, 2009; Navin et al, 2010). In particular, Navin et al (2010) demonstrated the existence of subclones that were derived from a common clonal progenitor, which varied between samples taken from different regions of the same primary tumour as well as subclones with distinct patterns of DNA copy number variation within the same sector of tumour. In these polygenomic tumours, subpopulation of cells could be anatomically separate or intermixed. There was no correlation between different tumour grades or immunohistochemical staining patterns and genomic heterogeneity.
Intratumour heterogeneity in glioblastoma heterogeneity in glioblastoma is also apparent (Snuderl et al, 2011; Nickel et al, 2012; Szerlip et al, 2012). One study identified intermingled population of cells with mutually exclusive amplifications of different receptor tyrosine kinases (RTKs) such as PDGFRA, MET, and EGFR, with subpopulation sharing similar alterations in CDKN2A and TP53 genes, consistent with a single common ancestral precursor (Snuderl et al, 2011). A second study confirmed intra-tumour heterogeneity of amplification of RTKs in glioblastoma and demonstrated that such heterogeneity results in functionally distinct and reduced sensitivity to targeted therapeutics (Szerlip et al, 2012). Spatial and temporal heterogeneity was characterised by targeted NGS in seven primary and recurrent tumour samples from one patient with glioblastoma; in this study, the variant allelic frequency of somatic mutations in EGFR, PI3KCA, PTEN, and TP53 vs wild type varied between focal regions of the same tumour, and between the time points of diagnosis, first recurrence and second recurrence (Nickel et al, 2012).
Spatial genomic heterogeneity has been recently documented in renal cell carcinoma (RCC) raising the potential for tumour sampling bias to confound biomarker interpretation (Gerlinger et al, 2012). Exome sequencing of multiple tumour samples from primary and metastatic lesions in two patients with clear cell RCC revealed extensive intra-tumour heterogeneity, which was demonstrated in genetic and transcriptomic analyses. Distinct loss of function somatic events in multiple tumour suppressor genes occurred in spatially separated regions of the same tumour, demonstrating convergent evolution and potentially predictable routes to tumour progression. Approximately 30–35% of mutations were shared between regions taken from multiple sites of the primary and metastases. Supporting evidence for intratumour heterogeneity in clear cell RCC, single cell exome sequencing of a clear cell RCC confirmed a complex genomic landscape, with a small number of genes mutated in a large proportion of cells and a greater number of genes mutated at low frequency (Xu et al, 2012).
Importantly, some of the earlier studies based on Sanger sequencing or low depth NGS may underestimate the degree of genomic heterogeneity due to limitations of sequencing depth that preclude the identification of rarer tumour subpopulations, and the study by Xu et al suggests that single cell approaches may be required to assay rare subclones. Furthermore, aberrations mediated through post-translational and epigenetic modifications as well as stochastic and unpredictable behavioural diversity of subclones with similar genotypic identities (Kreso et al, 2012) are likely to complicate the picture of intratumour heterogeneity further.
The genomic relationship between primary and metastatic tumours
Emerging data from deep sequencing analyses suggest that understanding clonal heterogeneity and the evolution of tumour subclonal architecture from the primary to the metastatic tumour sites and during therapy may provide important insight into the metastatic process and the emergence of drug resistance during systemic therapy.
There is now compelling evidence to support a branched evolutionary pattern of tumour growth across many haematological (Anderson et al, 2011) and solid tumours (Shah et al, 2009; Campbell et al, 2010; Yachida et al, 2010; Gerlinger et al, 2012; Wu et al, 2012). Comparison of the somatic mutational status of primary and metastatic tumour sites indicates that these may vary substantially, and phylogenetic reconstruction of tumour evolution in some of these studies suggests that a minor subpopulation. This was elegantly demonstrated in a recent ‘bicompartmental’ mouse model of medulloblastoma; complex genetic events were shared between metastatic tumour sites and a restricted population of their matched primaries, highly supportive of a common progenitor cell initiating metastatic outgrowth (Wu et al, 2012). The observation that some genetic aberrations were found solely in either primary or metastatic tumours lead to the proposal that further genetic divergence and evolution occurred independently at both sites after tumour dissemination. Similarly, the identification of somatic mutations in clear cell RCC that are shared by the primary tumour or metastatic sites only or are specific to one tumour region (private mutations) provides further evidence for ongoing independent subclonal evolution within distinct and spatially separated tumour regions (Gerlinger et al, 2012).
Implications for targeted therapeutics
The issue of cancer heterogeneity, including the relationships between subpopulation within and between tumour lesions, may have profound implications for drug therapy in cancer. Targeted therapy, which attempts to exploit a tumour’s dependence on a critical proliferation or survival pathway, has significantly improved patient outcomes in a range of solid tumour types, but in the majority of advanced disease cases, it is also apparent that targeted therapeutics do not help all molecularly selected patients and even when clinical benefit is observed, it is often of limited duration (Gore and Larkin, 2011; Diaz et al, 2012). Tumour heterogeneity may partly explain these clinical phenomena.
Considering an ‘actionable mutation’ within a model of clonal dominance, occurring in early cancer cell progenitors and present in the trunk of the tumour, may provide a more tractable approach for therapeutic targeting and for the identification of robust predictive biomarkers which are less susceptible to tumour sampling bias (Yap et al, 2012); such actionable ‘trunk’ mutations would be ubiquitous, clonally dominant driver events present in all tumour cells (Figure 2). However, if a tumour contains multiple branched events, depicting intra-tumour heterogeneity, then even the targeting of a driver event may not significantly influence treatment outcome due to a low-frequency subpopulation harbouring a resistance event in the tumour branches, leading to subclonal selection and the acquisition of drug resistance, as observed for the low-frequency gatekeeper mutation in EGFR in non-small cell lung cancer (NSCLC) (Su et al, 2012). Furthermore, we hypothesise that the role of somatic mutations as driver or passenger events is dynamic, subject to environmental and treatment selection pressures where passengers may become drivers and vice versa (Yap et al, 2012). Conceivably, clinical strategies may have to adapt to the possibility that branched, low frequency somatic events determine patient outcome and therapeutic failure. Indeed, by extension, heterogeneous somatic events may render tumour dependence of early truncal drivers redundant.
Trunk-branch model of tumour heterogeneity using single patient examples of renal cell carcinoma (Gerlinger et al, 2012). Ubiquitous driver events are common to all tumour sites and represent the trunk, while the heterogeneous passenger mutations, found at one site of disease but not another, are found in the branches of this model. It seems likely that ‘actionable mutations’, that is, those that could be therapeutically targeted and which could serve as useful predictive biomarkers, may prove more tractable when confined to the trunk. However, it is also plausible that heterogeneous minority subpopulation in the branches contribute to treatment resistance and failure, and targeting heterogeneous events in the branches may prove beneficial if paracrine signalling occurs from these subclones stimulating growth of dominant clones. It is also worth noting that in some glioblastoma and breast tumours, genetic events may be present in both the trunk and the branches (Shah et al, 2009).
Heterogeneity of secondary somatic mutations and drug resistance
Drug resistance is arguably the most critical problem faced by oncologists, and as a result almost all patients with metastatic solid tumours (with some notable exceptions such as seminoma) die of their disease. There are many examples of drug resistance conferred by the emergence of subclones harbouring specific somatic gene mutations. Imatinib-resistant mutations in the BCR-ABL fusion gene have been identified in patients with chronic myeloid leukaemia; some of these mutations have been shown to precede systemic treatment, and additionally, to co-exist with subclones carrying different imatinib-resistant mutations in treatment naïve patients (Shah et al, 2002). Intra-tumour heterogeneity of drug resistance mechanisms occurs in gastrointestinal stromal tumours (GIST) treated with imatinib or sunitinib; 9 of 11 patients with oncogenic KIT mutations developed secondary drug-resistant mutations, 6 of whom had two or more different mutations in separate metastases, and 3 of whom had 2 secondary KIT mutations in the same metastasis (Liegl et al, 2008). The bewildering complexity witnessed by multiple distinct mutations occurring in separate or identical metastases begins to illuminate the vast somatic mutational reservoir present in these tumours.
A similar theme exists in NSCLCs with a mutation in the EGFR – secondary mutations confer insensitivity to the EGFR tyrosine kinase inhibitor (EGFR-TKI) gefitinib, and have been identified in patients with clinical resistance to gefitinib, but also in untreated patients (Inukai et al, 2006). Su et al (2012) analysed NSCLC samples for one such mutation in EGFR, the T790M mutation, from treatment-naïve patients, and from patients before and during treatment with EGFR-TKIs. The presence of low-frequency T790M mutations before treatment predicted for shorter progression-free survival. In anaplastic lymphoma kinase (ALK)-rearranged lung cancers, ∼25% of patients with acquired resistance to the ALK inhibitor crizotinib exhibited a secondary mutation in the ALK tyrosine kinase domain, but these differed substantially between patients, and in a subset of patients, more than one resistance mechanism appeared to occur simultaneously (Katayama et al, 2012).
A unique example of intra-tumour heterogeneity and transplacental transfer of melanoma, intratumour heterogeneity and its contribution to therapeutic failure was recently described in a tragic case of transplacental transfer of melanoma (Sekulic et al, 2012). Genomic analysis of melanoma tumours from the mother (pre-vemurafenib treatment and at relapse) and the baby (before vemurafenib treatment) revealed two distinct but related clones in the mother, only one of which was present in the infant’s tumours. The shared clone appeared to be sensitive to vemurafenib treatment, while the unique clone progressed and lead to relapse in the mother.
Taken together, these findings suggest that biomarker efforts may have to rise to the challenge of identifying low frequency events, that may conceivably be spatially separated, in tumours before therapy to predict outcome. Such observations suggest that suitable combinatorial approaches specific to each individual patient’s tumour subclonal genetic heterogeneity composition might have to be considered to limit the acquisition of drug resistance; a single drug may not be adequate to treat a genetically heterogeneous tumour, since a pre-treatment cancer cell population harbouring resistance mutations, even if present at a low frequency, can contribute to therapeutic failure and poor outcome.
It is also plausible that there is a therapeutic window of opportunity in which to treat a driver mutation, before clonal expansion and divergence occurs (Gerlinger and Swanton, 2010). In a study from Ding et al (2012), evidence that the loss of founding clones and gain of mutations in the genomes of eight patients with relapsed AML was influenced by interval cytotoxic chemotherapy raised the possibility that cytotoxic treatment may contribute to relapse by inducing subclonal evolution.
It is important to note that cancer drug resistance occurs for reasons other than tumour heterogeneity, with the tumour microenvironment playing a key role. For example, innate resistance to BRAF inhibition in BRAF-mutant tumour cell lines was shown to be mediated by secretion of hepatocyte growth factor and its receptor, MET, by stromal cells, and reversal of resistance was achieved by combining BRAF and MET inhibitors (Straussman et al, 2012). Drug resistance can also be driven by non-mutational and epigenetic mechanisms that may be reversible (Lackner et al, 2012).
The impact of tumour heterogeneity on biomarker validation for targeted therapy
Clinically useful and economically viable targeted therapy relies upon biomarkers that are predictive of response to treatment. Tumour heterogeneity and tumour sampling bias may have had a negative impact on the discovery and validation of predictive biomarkers. For example, in metastatic RCC, despite identification of biologically relevant signalling pathways, there are limited genomics tools that facilitate prediction of response or resistance to targeted therapy. It should be noted that this may also reflect a lack of systematic tissue collection and translational research in the drug development trials for this disease.
Base-line genomic instability and intra-tumour heterogeneity may itself be a biomarker of poor clinical outcome (Birkbak et al, 2011; McGranahan et al, 2012). In Barrett’s oesophagus for example, clonal diversity appears to predict for increased risk of progression to invasive adenocarcinoma (Maley et al, 2006) and many studies have revealed the association of CIN with poor clinical outcome (reviewed in McGranahan et al, 2012). There is accumulating evidence to support intra-tumour heterogeneity, to which CIN is likely a major contributor, as a mechanism of drug resistance (Gerlinger and Swanton, 2010); CIN ovarian cancers may be more resistant to taxane chemotherapy (Swanton et al, 2009) and CIN colorectal cancer cell lines appear to be intrinsically multidrug resistant (Lee et al, 2011). Emerging NGS data may support these concepts; whole genome or exome sequencing of pre- and post-treatment biopsy samples of breast cancer patients treated with neoadjuvant aromatase inhibitors (AIs) in the ACOSOG Z1031 trial suggested that tumours which were more molecularly heterogeneous at the outset had a worse outcome; the background mutation rate among AI-resistant tumours was nearly double that of AI-sensitive tumours (Ellis et al, 2012).
It seems apparent that predicting drug response with clinically qualified biomarkers, is going to be challenging in practice due to the polygenic nature of drug resistance and the contributions of intra-tumour heterogeneity to this process, where low frequency, regionally separated subclones may influence therapeutic outcome. Conceivably, baseline estimates of intra-tumour heterogeneity may prove a more tractable approach to assess the propensity of a tumour to adapt to drug exposure.
Using intra-tumour heterogeneity to therapeutic advantage
The prospect of treating multiple genetic subpopulation within a tumour is discouraging, but it is possible that intra-tumour heterogeneity could be exploited for therapeutic advantage. Gatenby et al argue that an adaptive and dynamic model of cancer therapy is more likely to provide durable tumour control, and propose an experimental ‘evolutionary double blind therapy’ in which two therapies are given sequentially, each driving tumour cells to specific adaptations which render them vulnerable to the second therapy (Cunningham et al, 2011). This strategy has yet to be applied to clinical cancer therapy, but provides a good example of how improved knowledge of intra-tumour heterogeneity may be highly relevant to advances in the field of medical oncology. There is recent evidence of improved prognosis in breast cancer patients with tumours defined as having extreme CIN compared with those with intermediate CIN, suggesting that there may be an ideal level of genomic instability for tumorigenesis and that an excessive level may be deleterious to cancer cell viability, that may be influenced by cytotoxic therapy (Birkbak et al, 2011; Roylance et al, 2011). It is intriguing to speculate that such a paradoxical relationship between this pattern of genomic instability, contributing to intra-tumour heterogeneity, and clinical outcome might be therapeutically exploitable. Finally, identifying drivers or suppressors of genome instability in solid tumours, whose activation or inactivation is required to initiate intra-tumour heterogeneity and diversity, may provide a tractable route to ultimately attempt to limit tumour evolutionary processes.
Future directions
A systematic approach to the study of molecular heterogeneity in cancer is required, which is somewhat hindered by current trial design and funding mechanisms. Single biopsy specimens of primary tumours performed for diagnostic purposes, which are frequently archival, may not fully represent a genetically diverse malignancy with multiple metastatic sites, and sequencing techniques may not be sufficiently sensitive to detect low frequency events in tumour subclones. Often clinical practice is guided by the molecular analysis of primary tumours and we assume that all primary tumour characteristics are carried over to metastases later in the disease course. While this would hold true for trunk driver events, potentially important molecular changes will be missed if repeat biopsies are not performed during the evolution of the disease. Anticancer treatments may cause selection pressures and influence the complex mutational landscape within a tumour, analogous to the pruning of some tumour branches and the selection of other heterogeneous branched events through therapy.
Future clinical trials and biomarker studies might consider longitudinal analyses of tumour evolution through the disease course. We argue that much will be learned through the acquisition of multiple tumour samples from different regions of the primary and metastatic tumour sites and at sequential time points in the disease course, including before, during therapy and at disease progression, to study the changing nature of tumours over time. The requirement for multiple tumour biopsies is ethically and clinically challenging and new approaches, such as plasma circulating free tumour DNA sequencing technologies are primed to analyse tumour evolution over time (Dennis and Chiu 2011).
Conclusion
Cancer heterogeneity has potentially far-reaching consequences for cancer therapeutics, but translational research examining this issue is in its early stages. This is likely to change rapidly, with the ongoing development of advanced genomic technologies and increased access to these, and adaptation of clinical trial design to include comprehensive longitudinal tissue collection protocols. There is accumulating evidence for substantial genetic diversity both within and between many common solid tumours, but less is known about how such diversity is generated or its impact upon clinical outcomes such as response or resistance to anticancer therapies and the natural history of the disease. Conceivably, tumour heterogeneity may impede the identification of predictive biomarkers, and the quest for personalised, or even curative treatment, and is an area of cancer research worthy of intensive and collaborative effort.
References
Anderson K, Lutz C, van Delft FW, Bateman CM, Guo Y, Colman SM, Kempski H, Moorman AV, Titley I, Swansbury J, Kearney L, Enver T, Greaves M (2011) Genetic variegation of clonal architecture and propagating cells in leukaemia. Nature 469: 356–361
Birkbak NJ, Eklund AC, Li Q, McClelland SE, Endesfelder D, Tan P, Tan IB, Richardson AL, Szallasi Z, Swanton C (2011) Paradoxical relationship between chromosomal instability and survival outcome in cancer. Cancer Res 71: 3447–3452
Bonavia R, Inda MM, Cavenee WK, Furnari FB (2011) Heterogeneity maintenance in glioblastoma: a social network. Cancer Res 71: 4055–4060
Campbell PJ, Yachida S, Mudie LJ, Stephens PJ, Pleasance ED, Stebbings LA, Morsberger LA, Latimer C, McLaren S, Lin ML, McBride DJ, Varela I, Nik-Zainal SA, Leroy C, Jia M, Menzies A, Butler AP, Teague JW, Griffin CA, Burton J, Swerdlow H, Quail MA, Stratton MR, Iacobuzio-Donahue C, Futreal PA (2010) The patterns and dynamics of genomic instability in metastatic pancreatic cancer. Nature 467: 1109–1113
Cunningham JJ, Gatenby RA, Brown JS (2011) Evolutionary dynamics in cancer therapy. Mol Pharm 8: 2094–2100
Dennis Lo Y, Chiu RW (2011) Plasma nucleic acid analysis by massively parallel sequencing: pathological insights and diagnostic implications. J Pathol 225: 318–323
Diaz LA, Williams RT, Wu J, Kinde I, Hecht JR, Berlin J, Allen B, Bozic I, Reiter JG, Nowak MA, Kinzler KW, Oliner KS, Vogelstein B (2012) The molecular evolution of acquired resistance to targeted EGFR blockade in colorectal cancers. Nature 486: 537–540
Ding L, Ley TJ, Larson DE, Miller CA, Koboldt DC, Welch JS, Ritchey JK, Young MA, Lamprecht T, McLellan MD, McMichael JF, Wallis JW, Lu C, Shen D, Harris CC, Dooling DJ, Fulton RS, Fulton LL, Chen K, Schmidt H, Kalicki-Veizer J, Magrini VJ, Cook L, McGrath SD, Vickery TL, Wendl MC, Heath S, Watson MA, Link DC, Tomasson MH, Shannon WD, Payton JE, Kulkarni S, Westervelt P, Walter MJ, Graubert TA, Mardis ER, Wilson RK, DiPersio JF (2012) Clonal evolution in relapsed acute myeloid leukaemia revealed by whole-genome sequencing. Nature 481: 506–510
Egan JB, Shi CX, Tembe W, Christoforides A, Kurdoglu A, Sinari S, Middha S, Asmann Y, Schmidt J, Braggio E, Keats JJ, Fonseca R, Bergsagel PL, Craig DW, Carpten JD, Stewart AK (2012) Whole genome sequencing of multiple myeloma from diagnosis to plasma cell leukemia reveals genomic initiating events, evolution and clonal tides. Blood 120 (5): 1060–1066
Ellis MJ, Ding L, Shen D, Luo J, Suman VJ, Wallis JW, Van Tine BA, Hoog J, Goiffon RJ, Goldstein TC, Ng S, Lin L, Crowder R, Snider J, Ballman K, Weber J, Chen K, Koboldt DC, Kandoth C, Schierding WS, McMichael JF, Miller CA, Lu C, Harris CC, McLellan MD, Wendl MC, DeSchryver K, Allred DC, Esserman L, Unzeitig G, Margenthaler J, Babiera GV, Marcom PK, Guenther JM, Leitch M, Hunt K, Olson J, Tao Y, Maher CA, Fulton LL, Fulton RS, Harrison M, Oberkfell B, Du F, Demeter R, Vickery TL, Elhammali A, Piwnica-Worms H, McDonald S, Watson M, Dooling DJ, Ota D, Chang LW, Bose R, Ley TJ, Piwnica-Worms D, Stuart JM, Wilson RK, Mardis ER (2012) Whole-genome analysis informs breast cancer response to aromatase inhibition. Nature 486: 353–360
Gerlinger M, Rowan A, Horswell S, Larkin J, Endesfelder D, Gronroos E, Matthews PM, Stewart N, Mcdonald A, Butler N, Jones A, Raine D, Santos K, Varela C, Nohadani I, Eklund M, Spencer A, Dene B, Clark G, Pickering L, Stamp G, Gore M, Szallasi Z, Downward J, Futreal PA, Swanton C (2012) Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N Engl J Med 366: 883–893
Gerlinger M, Swanton C (2010) How Darwinian models inform therapeutic failure initiated by clonal heterogeneity in cancer medicine. Br J Cancer 103: 1139–1143
Gore ME, Larkin JM (2011) Challenges and opportunities for converting renal cell carcinoma into a chronic disease with targeted therapies. Br J Cancer 104: 399–406
Heppner GH (1984) Tumor heterogeneity. Cancer Res 44: 2259–2265
Inda MM, Bonavia R, Mukasa A, Narita Y, Sah DW, Vandenberg S, Brennan C, Johns TG, Bachoo R, Hadwiger P, Tan P, Depinho RA, Cavenee W, Furnari F (2010) Tumor heterogeneity is an active process maintained by a mutant EGFR-induced cytokine circuit in glioblastoma. Genes Dev 24: 1731–1745
Inukai M, Toyooka S, Ito S, Asano H, Ichihara S, Soh J, Suehisa H, Ouchida M, Aoe K, Aoe M, Kiura K, Shimizu N, Date H (2006) Presence of epidermal growth factor receptor gene T790M mutation as a minor clone in non-small cell lung cancer. Cancer Res 66: 7854–7858
Katayama R, Shaw AT, Khan TM, Mino-Kenudson M, Solomon BJ, Halmos B, Jessop NA, Wain JC, Yeo AT, Benes C, Drew L, Saeh JC, Crosby K, Sequist LV, Iafrate AJ, Engelman JA (2012) Mechanisms of acquired crizotinib resistance in ALK-rearranged lung cancers. Sci Transl Med 4: 120ra17
Keats JJ, Chesi M, Egan JB, Garbitt VM, Palmer SE, Braggio E, Van Wier S, Blackburn PR, Baker AS, Dispenzieri A, Kumar S, Rajkumar SV, Carpten JD, Barrett M, Fonseca R, Stewart AK, Bergsagel PL (2012) Clonal competition with alternating dominance in multiple myeloma. Blood 120 (5): 1067–1076
Kreso A, O'Brien CA, Van Galen P, Gan O, Notta F, Brown AMK, Ng K, Ma J, Wienholds E, Dunant C, Pollett A, Gallinger S, McPherson J, Mullighan CG, Shibata D, Dick JE . Variable clonal repopulation dynamics influence chemotherapy response in colorectal cancer. Science e-pub ahead of print, 13 December 2012
Lackner MR, Wilson TR, Settleman J (2012) Mechanisms of acquired resistance to targeted cancer therapies. Future Oncol 8: 999–1014
Lee AJ, Endesfelder D, Rowan AJ, Walther A, Birkbak NJ, Futreal PA, Downward J, Szallasi Z, Tomlinson IP, Howell M, Kschischo M, Swanton C (2011) Chromosomal instability confers intrinsic multidrug resistance. Cancer Res 71: 1858–1870
Liegl B, Kepten I, Le C, Zhu M, Demetri GD, Heinrich MC, Fletcher CDM, Corless CL, Fletcher JA (2008) Heterogeneity of kinase inhibitor resistance mechanisms in GIST. J Pathol 216: 64–74
Maley CC, Galipeau PC, Finley JC, Wongsurawat VJ, Li X, Sanchez CA, Paulson TG, Blount PL, Risques RA, Rabinovitch PS, Reid BJ (2006) Genetic clonal diversity predicts progression to esophageal adenocarcinoma. Nat Genet 38: 468–473
Marusyk A, Polyak K (2010) Tumor heterogeneity: causes and consequences. Biochim Biophys Acta 1805: 105–117
McGranahan N, Burrell RA, Endesfelder D, Novelli MR, Swanton C (2012) Cancer chromosomal instability: therapeutic and diagnostic challenges. EMBO Rep 13: 528–538
Navin N, Krasnitz A, Rodgers L, Cook K, Meth J, Kendall J, Riggs M, Eberling Y, Troge J, Grubor V, Levy D, Lundin P, Maner S, Zetterberg A, Hicks J, Wigler M (2010) Inferring tumor progression from genomic heterogeneity. Genome Res 20: 68–80
Navin NE, Hicks J (2010) Tracing the tumor lineage. Mol Oncol 4: 267–283
Nickel GC, Barnholtz-Sloan J, Gould MP, McMahon S, Cohen A, Adams MD, Guda K, Cohen M, Sloan AE, LaFramboise T (2012) Characterizing mutational heterogeneity in a glioblastoma patient with double recurrence. PLoS One 7: e35262
Nowell PC (1976) The clonal evolution of tumor cell populations. Science 194: 23–28
Roylance R, Endesfelder D, Gorman P, Burrell RA, Sander J, Tomlinson I, Hanby AM, Speirs V, Richardson AL, Birkbak NJ, Eklund AC, Downward J, Kschischo M, Szallasi Z, Swanton C (2011) Relationship of extreme chromosomal instability with long-term survival in a retrospective analysis of primary breast cancer. Cancer Epidemiol Biomarkers Prev 20: 2183–2194
Russnes HG, Navin N, Hicks J, Borresen-Dale AL (2011) Insight into the heterogeneity of breast cancer through next-generation sequencing. J Clin Invest 121: 3810–3818
Schuh A, Becq J, Humphray S, Alexa A, Burns A, Clifford R, Feller SM, Grocock R, Henderson S, Khrebtukova I, Kingsbury Z, Luo S, McBride D, Murray L, Menju T, Timbs A, Ross M, Taylor J, Bentley D (2012) Monitoring chronic lymphocytic leukemia progression by whole genome sequencing reveals heterogeneous clonal evolution patterns. Blood 120 (20): 4191–4196
Sekulic A, Hingorani P, Lenkiewicz E, Holley T, Barrett M, Zizmann V, Trent J (2012) Clonal evolution underlying transplacental transfer and vemurafenib resistance in melanoma. Pigment Cell Melanoma Res 25: 886
Shah NP, Nicoll JM, Nagar B, Gorre ME, Paquette RL, Kuriyan J, Sawyers CL (2002) Multiple BCR-ABL kinase domain mutations confer polyclonal resistance to the tyrosine kinase inhibitor imatinib (STI571) in chronic phase and blast crisis chronic myeloid leukemia. Cancer Cell 2: 117–125
Shah SP, Morin RD, Khattra J, Prentice L, Pugh T, Burleigh A, Delaney A, Gelmon K, Guliany R, Senz J, Steidl C, Holt RA, Jones S, Sun M, Leung G, Moore R, Severson T, Taylor GA, Teschendorff AE, Tse K, Turashvili G, Varhol R, Warren RL, Watson P, Zhao Y, Caldas C, Huntsman D, Hirst M, Marra MA, Aparicio S (2009) Mutational evolution in a lobular breast tumour profiled at single nucleotide resolution. Nature 461: 809–813
Sidransky D, Mikkelsen T, Schwechheimer K, Rosenblum ML, Cavanee W, Vogelstein B (1992) Clonal expansion of p53 mutant cells is associated with brain tumour progression. Nature 355: 846–847
Snuderl M, Fazlollahi L, Le LP, Nitta M, Zhelyazkova BH, Davidson CJ, Akhavanfard S, Cahill DP, Aldape KD, Betensky RA, Louis DN, Iafrate AJ (2011) Mosaic amplification of multiple receptor tyrosine kinase genes in glioblastoma. Cancer Cell 20: 810–817
Sprouffske K, Merlo LM, Gerrish PJ, Maley CC, Sniegowski PD (2012) Cancer in light of experimental evolution. Curr Biol 22: R762–R771
Straussman R, Morikawa T, Shee K, Barzily-Rokni M, Qian ZR, Du J, Davis A, Mongare MM, Gould J, Frederick DT, Cooper ZA, Chapman PB, Solit DB, Ribas A, Lo RS, Flaherty KT, Ogino S, Wargo JA, Golub TR (2012) Tumour micro-environment elicits innate resistance to RAF inhibitors through HGF secretion. Nature 487 (7408): 500–504
Su KY, Chen HY, Li KC, Kuo ML, Yang JC, Chan WK, Ho BC, Chang GC, Shih JY, Yu SL, Yang PC (2012) Pretreatment epidermal growth factor receptor (EGFR) T790M mutation predicts shorter EGFR tyrosine kinase inhibitor response duration in patients with non-small-cell lung cancer. J Clin Oncol 30 (4): 433–440
Swanton C, Nicke B, Schuett M, Eklund AC, Ng C, Li Q, Hardcastle T, Lee A, Roy R, East P, Kschischo M, Endesfelder D, Wylie P, Kim SN, Chen JG, Howell M, Ried T, Habermann JK, Auer G, Brenton JD, Szallasi Z, Downward J (2009) Chromosomal instability determines taxane response. Proc Natl Acad Sci USA 106: 8671–8676
Szerlip NJ, Pedraza A, Chakravarty D, Azim M, McGuire J, Fang Y, Ozawa T, Holland EC, Huse JT, Jhanwar S, Leversha MA, Mikkelsen T, Brennan CW (2012) Intratumoral heterogeneity of receptor tyrosine kinases EGFR and PDGFRA amplification in glioblastoma defines subpopulations with distinct growth factor response. Proc Natl Acad Sci USA 109: 3041–3046
Torres L, Ribeiro FR, Pandis N, Andersen JA, Heim S, Teixeira MR (2007) Intratumor genomic heterogeneity in breast cancer with clonal divergence between primary carcinomas and lymph node metastases. Breast Cancer Res Treat 102: 143–155
Walter MJ, Shen D, Ding L, Shao J, Koboldt DC, Chen K, Larson DE, McLellan MD, Dooling D, Abbott R, Fulton R, Magrini V, Schmidt H, Kalicki-Veizer J, O'Laughlin M, Fan X, Grillot M, Witowski S, Heath S, Frater JL, Eades W, Tomasson M, Westervelt P, DiPersio JF, Link DC, Mardis ER, Ley TJ, Wilson RK, Graubert TA (2012) Clonal architecture of secondary acute myeloid leukemia. N Engl J Med 366: 1090–1098
Wu X, Northcott PA, Dubuc A, Dupuy AJ, Shih DJH, Witt H, Croul S, Bouffet E, Fults DW, Eberhart CG, Garzia L, Van Meter T, Zagzag D, Jabado N, Schwartzentruber J, Majewski J, Scheetz TE, Pfister SM, Korshunov A, Li X-N, Scherer SW, Cho Y-J, Akagi K, Mac D, onald TJ, Koster J, McCabe MG, Sarver AL, Collins VP, Weiss WA, Largaespada DA, Collier LS, Taylor MD (2012) Clonal selection drives genetic divergence of metastatic medulloblastoma. Nature 482: 529–533
Xu X, Hou Y, Yin X, Bao L, Tang A, Song L, Li F, Tsang S, Wu K, Wu H, He W, Zeng L, Xing M, Wu R, Jiang H, Liu X, Cao D, Guo G, Hu X, Gui Y, Li Z, Xie W, Sun X, Shi M, Cai Z, Wang B, Zhong M, Li J, Lu Z, Gu N, Zhang X, Goodman L, Bolund L, Wang J, Yang H, Kristiansen K, Dean M, Li Y (2012) Single-cell exome sequencing reveals single-nucleotide mutation characteristics of a kidney tumor. Cell 148: 886–895
Yachida S, Jones S, Bozic I, Antal T, Leary R, Fu B, Kamiyama M, Hruban RH, Eshleman JR, Nowak MA, Velculescu VE, Kinzler KW, Vogelstein B, Iacobuzio-Donahue CA (2010) Distant metastasis occurs late during the genetic evolution of pancreatic cancer. Nature 467: 1114–1117
Yap TA, Gerlinger M, Futreal PA, Pusztai L, Swanton C (2012) Intratumor heterogeneity: seeing the wood for the trees. Sci Transl Med 4: 127ps10
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
This work is licensed under the Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Fisher, R., Pusztai, L. & Swanton, C. Cancer heterogeneity: implications for targeted therapeutics. Br J Cancer 108, 479–485 (2013). https://doi.org/10.1038/bjc.2012.581
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bjc.2012.581
Keywords
This article is cited by
-
Noninvasive prediction of metastasis in esophageal cancer using ensemble-based feature selection
International Journal of System Assurance Engineering and Management (2024)
-
NQO1 Mediates Lenvatinib Resistance by Regulating ROS-induced Apoptosis in Hepatocellular Carcinoma
Current Medical Science (2024)
-
A Cocktail of Natural Compounds Holds Promise for New Immunotherapeutic Potential in Head and Neck Cancer
Chinese Journal of Integrative Medicine (2024)
-
Supervised graph contrastive learning for cancer subtype identification through multi-omics data integration
Health Information Science and Systems (2024)
-
Patient-derived organoids for precision oncology: a platform to facilitate clinical decision making
BMC Cancer (2023)