Abstract
Klebsiella pneumonia infection rates have increased dramatically. Molecular typing and virulence analysis are powerful tools that can shed light on Klebsiella pneumonia infections. Whereas 77.7% (28/36) of clinical isolates indicated multidrug resistant (MDR) patterns, 50% (18/36) indicated carpabenem resistance. Gene prevalence for the AcrAB efflux pump (82.14%) was more than that of the mdtK efflux pump (32.14%) in the MDR isolates. FimH-1 and mrkD genes were prevalent in wound and blood isolates. FimH-1 gene was prevalent in sputum while mrkD gene was prevalent in urine. Serum resistance associated with outer membrane protein coding gene (traT) was found in all blood isolates. IucC, entB, and Irp-1 were detected in 32.14%, 78.5% and 10.7% of MDR isolates, respectively. We used two Polymerase Chain Reaction (PCR) analyses: Enterobacterial Repetitive Intergenic Consensus (ERIC) and Random Amplified Polymorphic DNA (RAPD). ERIC-PCR revealed 21 and RAPD-PCR revealed 18 distinct patterns of isolates with similarity ≥80%. ERIC genotyping significantly correlated with resistance patterns and virulence determinants. RAPD genotyping significantly correlated with resistance patterns but not with virulence determinants. Both RAPD and ERIC genotyping methods had no correlation with the capsule types. These findings can help up better predict MDR Klebsiella pneumoniae outbreaks associated with specific genotyping patterns.
Similar content being viewed by others
Introduction
K. pneumonia belongs to family Enterobacteriaceaea and is related to other genera, such as Enterobacter, Escherichia, and Salmonella1. K. pneumoniae is considered one of the most common Gram negative bacteria2. It is also an important pathogen in nosocomial infections in Egypt3,4. A number of factors contribute to virulence and pathogenicity in K. pneumoniae such as the capsular serotype, lipopolysaccharide, iron-scavenging systems and adhesions5. Iron acquisition systems are essential for the growth of pathogenic bacteria6. Moreover, the iron chelator siderophore allows bacteria to take up protein-bound iron from the host cells7.
The incidence of microbial infections has been increasing in the past few decades. This has led to the continuous and uncontrolled use of antimicrobial drugs for prevention and treatment in several parts of the world. This, in turn, led to the emergence of specific drug and multidrug resistance among various strains of microorganisms including K. pneumonia8. Gram-negative bacteria have developed several mechanisms of resistance to currently used antimicrobials. One of the successful mechanisms for transmitting multiple-druge resistance among bacterial pathogens is horizontal transfer9. The spread of MDR isolates in the clinic has been attributed to commonly shared plasmids across bacteria such as K. pneumoniae, K. oxytoca, Escherichia coli, Enterobacter sp., and Salmonella sp10,11. The efflux pump systems are among the most important causes of MDR12. Efflux pump systems in K. pneumoniae include AcrAB and mdtK systems, These belong to the Resistance Nodulation Division (RND) and Multi Antimicrobial Extrusion (MATE) family efflux pumps, respectively. The AcrAB-TolC pump is composed of an outer-membrane channel (TolC), a secondary transporter located in the inner membrane (AcrB), and a periplasmic component (AcrA)13. This pump is responsible for resistance to quinolones, tetracyclines, and chloramphenicol in various MDR isolates14. The MATE pumps, such as the mdtK system, transport some of those antimicrobial agents15. Porins such as OmpK35 and OmpK36 are crucial for the penetration of antibiotics into the cells and for susceptibility to cephalosporins and carbapenems16.
Carbapenems have been used for the treatment of infections caused by Enterobacteriaceae17. The percentage of Carbapenem-resistant Enterobactericeae (CRE) has been on the rise18. One of the most prominent recent increases of MDR was observed with Klebsiella sp. In the period from 2001 through 201118. It is noteworthy that patients with infections due to carbapenemase-producing enterobacteriaceae, such as K. pneumoniae experience high mortality rates19,20,21. Normally, these MDR infections are hard-to-treat with limited available choices of antibiotics such as tigecycline, colistin, fosfomycin, and aminoglycosides22,23.
Molecular typing and virulence analysis of clinical isolates are powerful tools that can shed light on multidrug resistant (MDR) Klebsiella pneumonia infections. We also used two Polymerase Chain Reaction (PCR) genotyping analyses: Enterobacterial Repetitive Intergenic Consensus (ERIC) and Random Amplified Polymorphic DNA (RAPD) to assess correlations of each with resistance patterns, virulence determinants, or capsule types of K. pneumoniae isolates.
Results
Primers
Primers used for amplification are listed in Table 1. More detail is provided under materials and methods.
Clinical isolates
Thirty six of K. pneumoniae clinical isolates were collected as described under materials and methods. Isolates were recovered from specimens of urine (n = 16), wound (n = 4), cerebrospinal fluid CSF (n = 1), blood (n = 7), sputum (n = 8) on MacConkey’s agar. Colonies showing lactose fermenting ability were further identified both microscopically and biochemically.
Antimicrobial susceptibility pattern and detection of genes coding for MDR efflux pumps and outer membrane porins
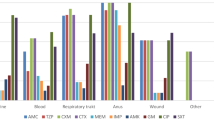
As determined by disc diffusion antimicrobial susceptibility testing method, a percentage 77.7% (28/36) of isolates showed multidrug resistance (MDR) patterns, but all these MDR isolates were sensitive to colisitin (10 μg). All MDR isolates were resistant to beta lactam antibiotics and 64.28%, 82.15%, and 85.7% showed resistance to carbapenem, quinolone, and aminoglycosides, respectively. Tetracycline and chloramphenicol were effective against 61.1% of carbapenem-resistant isolates. The tested isolates were distributed into 24 antimicrobial resistance patterns (Table 2). Most patterns showed resistance to cephalosporin and beta lactam/beta lactamase inhibitors. The most predominant pattern was A6 and A8.
Gene prevalence for the AcrAB efflux pump system (82.14%) was more than that of the mdtK efflux pump (32.14%) in the MDR isolates. Incomplete AcrAB efflux pump system was detected in the remaining five isolates. The genes coding for porin protein (ompK35) and (ompK36) were not detected in six and four MDR isolates, respectively. Genes coding for the porins (ompK 35 and ompK36) were detected in all isolates recovered from wound and CSF specimens. The presence of these two porin-coding genes was variable in blood, sputum, and urine samples.
Detection of virulence genes
The prevalence and distribution of virulence factors are shown in Table 3. The fimH-1 and mrkD genes, encoding type 1 and type 3 fimbrial adhesins, were present in all wound and blood isolates. The fimH-1 gene was prevalent in sputum isolates whereas mrkD gene was prevalent in all urine samples. Serum resistance associated with the outer membrane protein coding gene (traT) was detected in all blood isolates.
The iron siderophores, aerobactin synthase gene (IucC), enterobactin biosynthesis gene (entB) and Yersinibactin biosynthesis gene (Irp-1) were detected in 32.14%, 85.7% and 28.5% of MDR isolates, respectively.
The prevalence of capsule K genotypes in the 28 MDR isolates revealed that K1 (n = 8), K2 (n = 2) and the remaining isolates were non-typable as K1 or K2 genotypes. Isolates showing K1 genotypes were obtained from urine, blood, and sputum specimens, while K2 isolates was recovered from urine and wound samples. There were 16 virulence profiles according to detected virulence genes. It is noteworthy that virulence genetic profiles indicated that virulence determinants were variable among K. pneumoniae strains that possess the same capsule genotype (Table 3).
Genotyping of Klebsiella pneumoniae isolates by RAPD and ERIC analyses
According to the dendrograms, Enterobacterial Repetitive Intergenic Consensus (ERIC) and Random Amplified Polymorphic DNA (RAPD) analyses revealed 21 and 18 distinct patterns of K. pneumoniae isolates with similarity >80%, respectively (Figs 1 and 2). The 21 ERIC genotypes were designated E1 to E21 while the RAPD genotypes were designated R1 to R18 and each of their variant subtypes were indicated by a letter suffix. Dendrogram analysis of ERIC genotyping showed three clusters (A–C): clusters A, B, and C contained 12/28, 9/28, and 7/28 of the MDR isolates, respectively. The isolates (18 and 21) and (28, 35, 51, and 56) showed high similarity which may suggest that those isolates constitute a clonal lineage (Fig. 1). On the other hand, the RAPD genotyping revealed different pattern with 6 clusters (A–F). The isolates (15 and 58), (20 and 21), (9 and 18), and (36 and 56) showed high similarity (Fig. 2). Based on Simpson’s index of diversity, the discriminatory potential of different typing techniques used with K. pneumoniae isolates varied from 0.519 to 0.984 (Table 4). The high Simpson’s index of diversity for the antibiotyping, virulence, RAPD, and ERIC typing indicates greater diversity. Kendall’s tau-b correlation coefficient was calculated between RAPD and ERIC genotyping methods versus resistance patterns, virulence determinants, and capsule types of K. pneumoniae. Based on the statistical correlation tests (Table 5), the ERIC genotyping significantly correlates with resistance patterns (p < 0.01) and virulence determinants (p < 0.05). On the other hand, the RAPD genotyping significantly correlates with resistance patterns (p < 0.05) but not with virulence determinants (p > 0.05). Both RAPD and ERIC genotyping methods have no correlation with the capsule types (p > 0.05).
Discussion
K. pneumoniae is the causative agent of several different healthcare-associated infections, such as bloodstream infections, wound infections, pneumonia, and meningitis. The extensive use of antimicrobials led to high incidence of resistance in K. pneumoniae24. In our study, K. pneumoniae isolates showing multidrug resistance comprised 71.1% of total samples. Rates as high as 66.7% of MDR K. pneumoniae isolates were also detected in other studies25. The high rates of antimicrobial resistance detected in our study can be attributed to the lack of strict policies that govern the use of antibiotics in Egypt3.
Antibiotic efflux pumps represent one of the most important antimicrobial resistance mechanisms used by K. pneumoniae clinical isolates26,27. The increased efflux of the antimicrobial agent leads to the reduction of its intracellular concentration, which can enhance bacterial survival28. The AcrAB efflux pump was more common than mdtK. The presence of the multidrug efflux pump system (AcrAB-TolC) was significantly correlated with the MDR pattern. On the other hand, five MDR isolates was missing either the AcrAB efflux pump or the TolC outer membrane protein or both.
Gram negative bacterial outer membranes are poorly permeable to both hydrophobic and hydrophilic molecules. Thus, most antimicrobial agents other than β-lactam must cross the membrane in order to reach their intracellular drug targets and so require the presence of porin to bypass the asymmetric bilayer of phospholipid and lipopolysaccharide membrane29. Consequently, it has been reported that loss of porins ompK 35 and ompK 36 led to an increase in carbapenem, ciprofloxacin, and chloramphenicol resistance30. Surprisingly, in our research, porin loss was not significantly correlated to the MDR pattern (P > 0.05). This could be attributed to the presence of point mutations, disruption in the protein coding sequence, or promoter region mutations31.
In the current study, about fifty percent of the total isolates showed resistance to both imipenem and ertapenem. In this context, there has been a significant increase in carbapenem resistance among K. pneumoniae isolates in Egypt during the last few years (from 13.9% to 44.4%)3,32,33. K. pneumoniae isolates possess several mechanisms to evade the activity of carbapenems. These include AmpC production or ESBL production together with porin loss, carbapenemase production, and production of acquired Metallobetalactamase (MBL)34.
Contrary to the New Delhi metallo-β-lactamase, which is a broad spectrum carbapenemase with ability to inactivate β-lactams except aztreonam35, all carabapenem resistant isolates in our study were also resistant to aztreonem. This may be due to the development of a new antimicrobial resistance pattern in Egyptian hospitals. In all tested carbapenem-resistant isolates (n = 18), there was no simultaneous porin loss with AmpC or ESBL production. In another study36, Szabó et al. showed that OmpD and OmpF in an ertapenem-resistant E. coli strain were less permeable than those of a susceptible control strain. This suggested that the possession of these two porins could lead to higher resistance due to an associated pump system36. This is relevant to our study given the fact that OmpF genes in E. coli are homologues to OmpK35 genes in K. pneumoniae29.
Carbapenemase-producing enterobacteriaceae, such as K. pneumonia, can cause deadly infections20,21. Colistin was used to treat Gram-negative infections but was abandoned because of its toxicity. Recently, it has been revived again as a treatment for life-threatening infections caused by some resistant Gram-negative bacteria, such as Pseudomonas aeruginosa and Acinetobacter baumannii37. Interestingly, all the tested K. pneumoniae isolates in this study were sensitive to colistin.
Both OmpK35 and OmpK36 play a role in K. pneumoniae virulence and infection. Deletion of OmpK36 or OmpK35/OmpK36 can lead to the reduction in virulence of highly virulent strains and can increase their susceptibility to neutrophil phagocytosis38,39. In our investigation, both Ompk 35 and OmpK36 porin- coding genes were simultaneously detected in all K. pneumoniae isolates recovered from wound and CSF samples. Their presence was variable though in sputum, blood, and urine samples. A direct correlation between efflux pumps and virulence of pathogenic bacteria was reported by Padilla et al40. Several genes essential for intracellular invasion and survival were downregulated in mutant strains lacking acrAB-tolC efflux pumps41.
Type 1 fimbriae are the most common adhesive organelles in enterobacteriaceae and can lead to urinary tract infections42. Type 3 fimbrial adhesin can mediate the binding of K. pneumoniae to endothelial cells and to epithelial cells of the respiratory and urinary tracts. MrkD protein is a crucial factor in binding bacteria to the collagen molecules of the mammalian cells43. Many K. pneumoniae clinical isolated normally express both type 1 and type 3 fimbrial adhesins44. In the current study, the two genes coding for these adhesive structures (Type 3 fimbrial adhesin and MrkD) were detected in all wound, blood, and CSF isolates and in about 80% of sputum and urine isolates. The plasmidic traT gene encodes an outer membrane protein involved in bacterial conjugation and blocks the complement-mediated cascade, and act as an invasin45. We detected the traT gene in twenty two K. pneumoniae isolates (78.5%). The prevalence of traT gene in our isolates was relatively high as it was frequently associated with the K1 capsule serotype.
Most enterobacteriaceae strains contain genes encoding iron uptake systems, such as enterochelin or aerobactin46. These siderophores have dual roles as they can also inhibit T cell proliferation in addition to their role in enhancing iron uptake47. The iron siderophores aerobactin synthase gene (IucC), enterobactin biosynthesis gene (entB), and yersinibactin biosynthesis gene (Irp-1) were detected in 32.14%, 78.5%, and 10.7% of MDR K. pneumoniae isolates, respectively. Highly pathogenic Yersinia strains have high-pathogenicity island (HPI) that contain the gene Irp-1. This HPI is also prevalent in Klebsiella and other enterbocateria48, such as E. coli, K. oxytoca, K. pneumonia, Citrobacter species, and Enterobacter species46.
The capsular serotypes K1 and K2 are associated with the predominant virulent strains of K. pneumoniae49. Feizabadi et al. has shown that K1 and K2 serotypes represented 11.2% and 14.6%, respectively, of the total K. pneumoniae isolates50. In our study, K1 and K2 serotypes represented 28.5% and 7.14% of the MDR K. pneumoniae isolates.
Molecular typing is a potent tool for the study of nosocomial infections51. RAPD is a widely used genotyping tool for K. pneumoniae strains with ESBLs production (Gori et al., 1996). Out of a total of 28 MDR K. pneumoniae isolates in the current investigation, ERIC-PCR revealed 21 and RAPD-PCR revealed 18 distinct patterns of K. pneumoniae isolates. This may be attributed to the genetic variation in pathogenic these K. pneumoniae strains. Our data confirms the observations of Lai et al.52 that pathogenic K. pneumoniae is highly heterogeneous, due to differences in nucleotide sequences. The large number of serotypes in this species could also explain this genetic diversity highlighted by the RAPD-PCR genotypic analysis53.
Correlations between RAPD-PCR genotyping and antibiotic resistance patterns of K. pneumonia were observed by Ashayeri-Panah et al.54 and Espinar et al.55. In the current study, both RAPD-PCR and ERIC-PCR genotypic analyses revealed correlations with resistance patterns of K. pneumonia. The highest correlation coefficients were observed with ERIC genotyping, indicating that the latter may be more valuable in prediction of resistance patterns of K. pneumoniae as compared to the RAPD-PCR genotyping method. Moreover, ERIC, but not RAPD-PCR, revealed statistically significant correlations with virulence determinants of K. pneumoniae. Finally, both RAPD-PCR and ERIC-PCR showed no statistically significant correlation with the detected capsule types of K. pneumoniae isolates. Results included in this study can help up better predict MDR Klebsiella pneumoniae outbreaks associated with specific genotyping patterns in the future.
Materials and Methods
Bacterial strains
Thirty six K. pneumonia clinical isolates were recovered from patients at Kasr El Aini Hospitals, Cairo, Egypt. Approvals from the institutional review board of the hospitals and the Research Ethics committee of the October University for Modern Sciences and Arts, Giza, Egypt were obtained prior to conducting the study. All methods were performed in accordance with the required guidelines and regulations.
For experiments involving human samples, informed consent was obtained from all subjects. Strains were isolated from sputum, urine, blood, wound, and cerebrospinal fluid (CSF) specimens. Specimens were collected in the period from August 2015 through December 2015. Isolates were identified by conventional and biochemical tests as described previously56 and then were stored at −20 °C in brain heart infusion broth with 15% v/v glycerol.
Antimicrobial susceptibility
Antibiotic susceptibility testing of Klebsiella sp., was performed according to the Kirby-Bauer disk diffusion method57. The antimicrobial sensitivity assays to nineteen antibacterial drugs were done using commercially available antibiotic discs (OXOID, UK) including Ampicillin (AMP, 10 μg), Amoxicillin/Clavulanic acid (AMC, 20/10 μg), Piperacillin/Tazobactam (TZP, 110/10 μg), Cefoxitin (FOX, 30 μg), Ceftazidime (CAZ, 30 μg), Cefuroxime (CXM, 30 μg), Aztreonam (ATM, 30 μg), Ertapenem (ETP, 10 μg), Impinem (IMP, 10 μg), Gentamicin (CN, 10 μg), Tobramycin (TOB, 10 μg), Amikacin (AK, 30 μg), Tetracycline (TE, 30 μg), Doxycycline (DO, 30 μg), Ciprofloxacin (CIP, 5 μg), Nalidixic acid (NA, 30 g), Co-trimoxazole (SXT, 30 μg), Colistin (CT, 10 μg) and Chloramphenicol (C, 30 μg). For all tested antimicrobials except colistin (10 μg), the plates were then incubated at 37 °C for 24 hours, the diameters of the inhibition zones were measured in millimeter and interpretation of results was done according to CLSI standards57. For Colistin, breakpoints were used for interpretation58. Multidrug resistant (MDR) isolates were selected according to their non-susceptibility to at least one agent in three or more antimicrobial categories59.
Detection of multidrug resistance and virulence determinants using PCR
DNA extraction
Genomic DNA was extracted from overnight culture using ZYMO Quick-gDNA™ MiniPrep (ZYMO Research, CA, USA). Concentration of the DNA extract and purity was determined by measuring absorbance at wavelengths 260 and 280 nm. The integrity of genomic DNA was tested by resolving DNA extracts on a 0.8% w/v agarose gel by electrophoresis. These crude DNA extracts were frozen at −20 °C.
Primers used for amplification are listed in Table 1 and were prepared by Invitrogen® (Thermo Fisher scientific Inc., MA, USA). Primers were designed using the complete genome sequence of K. pneumoniae MGH 78578 (accession no. CP000647) and the internet based software Basic Local Alignment Tool (BLAST) in NCBI and Multiple sequence alignment using the CLUSTAL Omega in EMBL-EBI.
PCR detection of multidrug resistance genes
Isolates that showed multidrug resistance phenotypes were tested for genes coding for the multidrug efflux pump system AcrAB-TolC and MdtK, in addition to porin coding genes (OmpK35 and OmpK36). The amplifications of these genes were performed in cycles with initial denaturing at 94 °C for 5 min followed by 35 cycles, each cycle consisting of 30 seconds at 94 °C for denaturation, 30 seconds for primer annealing (Table 1), and 1.5 min at 72 °C for elongation. After these cycles, the final elongation step was carried out at 72 °C for 10 min45.
PCR detection of virulence-associated genes
PCR was used to amplify the virulence-associated genes. These genes include those encoding for regulators of mucoid phenotype A (rmpA), type 1 and type 3 adhesins (fimH-1, mrkD), aerobactin (iron siderophore) synthase (IucC), bacteriocin biosynthesis [enterobactin (entB), and yersiniabactin (irP-1)], and serum resistance-associated outer membrane lipoprotein (traT). Measurements of the prevalence of capsule serotypes K1 and K2 were also included.
The PCR conditions were similar to those used for detection of multidrug resistance genes with annealing temperatures included in Table 1.
Molecular typing of Klebsiella pneumoniae isolates using Random Amplified Polymorphic DNA (RAPD) and Enterobacterial Repetitive Intergenic Consensus (ERIC) methods
Typing by randomly amplified polymorphic DNA (RAPD) analysis was performed according to the protocol published by Deschaght et al.60 using the primer RAPD4 (5′-AAGACGCCGT-3′). Briefly, two microliters of the DNA template were added to 12.5 μL multiplex mastermix (MyTaq™HM Mix, Bioline®, MA, USA), 1 μL primer (10 pmol), and 9.5 μL H2O. PCR cycles of initial incubation at 94 °C for 15 min followed by cycling for 40 times at 94 °C for 1 min, 37 °C for 1 min, and a final elongation at 72 °C for 2 min was performed.
ERIC typing was carried out using the primer ERIC2 (5′-AAGTAAGTGACTGGGGTGAGCG-3′) using a similar PCR program to that of the RAPD method except for an extension time of 8 min59.
RAPD and ERIC fragments were visualized by 1.5% w/v agarose gel electrophoresis and results were analyzed using GelCompar II software (Version 6.6.11, Applied Maths, Kortrijk, Belgium). The patterns were normalized with bands of the marker and bands that were consistently present in all patterns. Computer-assisted analyses implemented in this study were performed according to the manufacturer’s instructions.
Comparing different typing methods and calculation of discriminatory index
Simpsons index of diversity [discriminatory index (D)], based on the probability that two unrelated isolate samples from the test population are located in different typing groups, was calculated according to the following equation:

where N is the total number of isolates in the sample population, s is the total number of types described, and nj is the number of strains belonging to the jth type. Simpsons index of diversity ranges from 0.0 to 1.0, where 1.0 indicates that a typing method is able to distinguish each member of a population from all other members of that population and, conversely, 0.0 indicates that all members of a strain population are of an identical type61.
Statistical analysis
All statistical analyses were performed using SPSS, version 18.0 (SPSS Inc., NY, USA). Chi-square tests were used to compare categorical measures between groups (Fisher’s exact test where appropriate). Statistical correlation tests, including Kendall’s tau-b nonparametric correlation coefficients, were determined at the two-tailed significance level for correlation of genotyping methods with virulence determinants, antimicrobial resistance, and capsule types. Data output of correlation analyses with p values less than 0.05 were considered statistically significant.
Additional Information
How to cite this article: Wasfi, R. et al. Molecular typing and virulence analysis of multidrug resistant Klebsiella pneumoniae clinical isolates recovered from Egyptian hospitals. Sci. Rep. 6, 38929; doi: 10.1038/srep38929 (2016).
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Brisse, S. & Verhoef, J. Phylogenetic diversity of Klebsiella pneumoniae and Klebsiella oxytoca clinical isolates revealed by randomly amplified polymorphic DNA, gyrA and parC genes sequencing and automated ribotyping. Int J Syst Evol Microbiol 51, 915–924, doi: 10.1099/00207713-51-3-915 (2001).
Lin, W. H. et al. Clinical and microbiological characteristics of Klebsiella pneumoniae isolates causing community-acquired urinary tract infections. Infection 38, 459–464, doi: 10.1007/s15010-010-0049-5 (2010).
Daef, E. A. & Elsherbiny, N. M. Clinical and Microbiological Profile of Nosocomial Infections in Adult Intensive Care Units at Assiut University Hospitals, Egypt. Journal of American Science 8, 1239–1250 (2012).
Abdel-Wahab, F., Ghoneim, M., Khashaba, M., El-Gilany, A. H. & Abdel-Hady, D. Nosocomial infection surveillance in an Egyptian neonatal intensive care unit. J Hosp Infect 83, 196–199, doi: 10.1016/j.jhin.2012.10.017 (2013).
Fuursted, K. et al. Virulence of a Klebsiella pneumoniae strain carrying the New Delhi metallo-beta-lactamase-1 (NDM-1). Microbes Infect 14, 155–158, doi: 10.1016/j.micinf.2011.08.015 (2012).
Lawlor, M. S., O’Connor, C. & Miller, V. L. Yersiniabactin Is a Virulence Factor for Klebsiella pneumoniae during Pulmonary Infection. Infection and Immunity 75, 1463–1472, doi: 10.1128/IAI.00372-06 (2007).
Wu, C.-C. et al. IscR Regulation of Capsular Polysaccharide Biosynthesis and Iron-Acquisition Systems in Klebsiella pneumoniae CG43. PLoS One 9, e107812, doi: 10.1371/journal.pone.0107812 (2014).
Tanwar, J., Das, S., Fatima, Z. & Hameed, S. Multidrug Resistance: An Emerging Crisis. Interdisciplinary Perspectives on Infectious Diseases 2014, 7, doi: 10.1155/2014/541340 (2014).
Munoz-Price, L. S. & Quinn, J. P. The Spread of Klebsiella pneumoniae Carbapenemases: A Tale of Strains, Plasmids, and Transposons. Clinical Infectious Diseases 49, 1739–1741, doi: 10.1086/648078 (2009).
Falagas, M. E. & Karageorgopoulos, D. E. Extended-spectrum beta-lactamase-producing organisms. J Hosp Infect 73, 345–354, doi: 10.1016/j.jhin.2009.02.021 (2009).
Miro, E. et al. Spread of plasmids containing the bla(VIM-1) and bla(CTX-M) genes and the qnr determinant in Enterobacter cloacae, Klebsiella pneumoniae and Klebsiella oxytoca isolates. J Antimicrob Chemother 65, 661–665, doi: 10.1093/jac/dkp504 (2010).
Meletis, G., Exindari, M., Vavatsi, N., Sofianou, D. & Diza, E. Mechanisms responsible for the emergence of carbapenem resistance in Pseudomonas aeruginosa. Hippokratia 16, 303–307 (2012).
Du, D. et al. Structure of the AcrAB-TolC multidrug efflux pump. Nature 509, 512–515, doi: 10.1038/nature13205 (2014).
Okusu, H., Ma, D. & Nikaido, H. AcrAB efflux pump plays a major role in the antibiotic resistance phenotype of Escherichia coli multiple-antibiotic-resistance (Mar) mutants. Journal of Bacteriology 178, 306–308 (1996).
Sun, J., Deng, Z. & Yan, A. Bacterial multidrug efflux pumps: Mechanisms, physiology and pharmacological exploitations. Biochemical and Biophysical Research Communications 453, 254–267, doi: 10.1016/j.bbrc.2014.05.090 (2014).
Shi, W. et al. Carbapenem and cefoxitin resistance of Klebsiella pneumoniae strains associated with porin OmpK36 loss and DHA-1 beta-lactamase production. Braz J Microbiol 44, 435–442, doi: 10.1590/s1517-83822013000200015 (2013).
Vardakas, K. Z., Tansarli, G. S., Rafailidis, P. I. & Falagas, M. E. Carbapenems versus alternative antibiotics for the treatment of bacteraemia due to Enterobacteriaceae producing extended-spectrum β-lactamases: a systematic review and meta-analysis. Journal of Antimicrobial Chemotherapy 67, 2793–2803, doi: 10.1093/jac/dks301 (2012).
CDC. Vital Signs: Carbapenem-Resistant Enterobacteriaceae. 165–170 (Center for Disease Control and prevention (CDC), 2013).
Ben-David, D. et al. Outcome of carbapenem resistant Klebsiella pneumoniae bloodstream infections. Clinical Microbiology and Infection 18, 54–60, doi: 10.1111/j.1469-0691.2011.03478.x (2012).
Patel, G. et al. Outcomes of Carbapenem-Resistant Klebsiella pneumoniae Infection and the Impact of Antimicrobial and Adjunctive Therapies. Infection Control and Hospital Epidemiology 29, 1099–1106, doi: 10.1086/592412 (2008).
Chang, H.-J. et al. Risk factors and outcomes of carbapenem-nonsusceptible Escherichia coli bacteremia: A matched case–control study. Journal of Microbiology, Immunology and Infection 44, 125–130, doi: 10.1016/j.jmii.2010.06.001 (2011).
Falagas, M. E. & Kopterides, P. Old antibiotics for infections in critically ill patients. Current Opinion in Critical Care 13, 592–597, doi: 10.1097/MCC.0b013e32827851d7 (2007).
Falagas, M. E., Kasiakou, S. K. & Saravolatz, L. D. Colistin: The Revival of Polymyxins for the Management of Multidrug-Resistant Gram-Negative Bacterial Infections. Clinical Infectious Diseases 40, 1333–1341, doi: 10.1086/429323 (2005).
Cao, X. et al. Molecular characterization of clinical multidrug-resistant Klebsiella pneumoniae isolates. Annals of Clinical Microbiology and Antimicrobials 13, 16–16, doi: 10.1186/1476-0711-13-16 (2014).
Paneru, T. P. Surveillance of Klebsiella pneumoniae and antibiotic resistance a retrospective and comparative study through a period in Nepal. Danish journal of medical and biology sciences, 29–36 (2015).
Peleg, A. Y. & Hooper, D. C. Hospital-acquired infections due to gram-negative bacteria. N Engl J Med 362, 1804–1813, doi: 10.1056/NEJMra0904124 (2010).
Zavascki, A. P., Carvalhaes, C. G., Picao, R. C. & Gales, A. C. Multidrug-resistant Pseudomonas aeruginosa and Acinetobacter baumannii: resistance mechanisms and implications for therapy. Expert Rev Anti Infect Ther 8, 71–93, doi: 10.1586/eri.09.108 (2010).
Piddock, L. J. Clinically relevant chromosomally encoded multidrug resistance efflux pumps in bacteria. Clin Microbiol Rev 19, 382–402, doi: 10.1128/cmr.19.2.382-402.2006 (2006).
Nikaido, H. Molecular basis of bacterial outer membrane permeability revisited. Microbiol Mol Biol Rev 67, 593–656 (2003).
Kaczmarek, F. M., Dib-Hajj, F., Shang, W. & Gootz, T. D. High-Level Carbapenem Resistance in a Klebsiella pneumoniae Clinical Isolate Is Due to the Combination of bla(ACT-1) β-Lactamase Production, Porin OmpK35/36 Insertional Inactivation, and Down-Regulation of the Phosphate Transport Porin PhoE. Antimicrobial Agents and Chemotherapy 50, 3396–3406, doi: 10.1128/AAC.00285-06 (2006).
Doumith, M., Ellington, M. J., Livermore, D. M. & Woodford, N. Molecular mechanisms disrupting porin expression in ertapenem-resistant Klebsiella and Enterobacter spp. clinical isolates from the UK. J Antimicrob Chemother 63, 659–667, doi: 10.1093/jac/dkp029 (2009).
Ashour, H. M. & El-Sharif, A. Species distribution and antimicrobial susceptibility of gram-negative aerobic bacteria in hospitalized cancer patients. J Transl Med 7, 14, doi: 10.1186/1479-5876-7-14 (2009).
Metwally, L., Gomaa, N., Attallah, M. & Kamel, N. High prevalence of Klebsiella pneumoniae carbapenemase-mediated resistance in K. pneumoniae isolates from Egypt. East Mediterr Health J 19, 947–952 (2013).
Pfeifer, Y., Cullik, A. & Witte, W. Resistance to cephalosporins and carbapenems in Gram-negative bacterial pathogens. International Journal of Medical Microbiology 300, 371–379, doi: 10.1016/j.ijmm.2010.04.005 (2010).
Shakil, S. et al. New Delhi metallo-beta-lactamase (NDM-1): an update. J Chemother 23, 263–265, doi: 10.1179/joc.2011.23.5.263 (2011).
Szabó, D. et al. Outer Membrane Protein Changes and Efflux Pump Expression Together May Confer Resistance to Ertapenem in Enterobacter cloacae. Antimicrobial Agents and Chemotherapy 50, 2833–2835, doi: 10.1128/AAC.01591-05 (2006).
Karabinis, A. et al. Colistin for Klebsiella pneumoniae—Associated Sepsis. Clinical Infectious Diseases 38, e7–e9, doi: 10.1086/380461 (2004).
Chen, J. H. et al. Contribution of outer membrane protein K36 to antimicrobial resistance and virulence in Klebsiella pneumoniae. J Antimicrob Chemother 65, 986–990, doi: 10.1093/jac/dkq056 (2010).
Tsai, Y.-K. et al. Klebsiella pneumoniae Outer Membrane Porins OmpK35 and OmpK36 Play Roles in both Antimicrobial Resistance and Virulence. Antimicrobial Agents and Chemotherapy 55, 1485–1493, doi: 10.1128/AAC.01275-10 (2011).
Padilla, E. et al. Klebsiella pneumoniae AcrAB efflux pump contributes to antimicrobial resistance and virulence. Antimicrob Agents Chemother 54, 177–183, doi: 10.1128/aac.00715-09 (2010).
Webber, M. A., Randall, L. P., Cooles, S., Woodward, M. J. & Piddock, L. J. V. Triclosan resistance in Salmonella enterica serovar Typhimurium. Journal of Antimicrobial Chemotherapy 62, 83–91, doi: 10.1093/jac/dkn137 (2008).
Struve, C., Bojer, M. & Krogfelt, K. A. Characterization of Klebsiella pneumoniae type 1 fimbriae by detection of phase variation during colonization and infection and impact on virulence. Infect Immun 76, 4055–4065, doi: 10.1128/iai.00494-08 (2008).
Langstraat, J., Bohse, M. & Clegg, S. Type 3 fimbrial shaft (MrkA) of Klebsiella pneumoniae, but not the fimbrial adhesin (MrkD), facilitates biofilm formation. Infect Immun 69, 5805–5812 (2001).
Sahly, H. et al. Extended-spectrum beta-lactamase production is associated with an increase in cell invasion and expression of fimbrial adhesins in Klebsiella pneumoniae. Antimicrob Agents Chemother 52, 3029–3034, doi: 10.1128/aac.00010-08 (2008).
El Fertas-Aissani, R., Messai, Y., Alouache, S. & Bakour, R. Virulence profiles and antibiotic susceptibility patterns of Klebsiella pneumoniae strains isolated from different clinical specimens. Pathol Biol (Paris) 61, 209–216, doi: 10.1016/j.patbio.2012.10.004 (2013).
Schubert, S., Cuenca, S., Fischer, D. & Heesemann, J. High-pathogenicity island of Yersinia pestis in enterobacteriaceae isolated from blood cultures and urine samples: prevalence and functional expression. J Infect Dis 182, 1268–1271, doi: 10.1086/315831 (2000).
Autenrieth, I., Hantke, K. & Heesemann, J. Immunosuppression of the host and delivery of iron to the pathogen: a possible dual role of siderophores in the pathogenesis of microbial infections? Med Microbiol Immunol 180, 135–141 (1991).
Carniel, E. The Yersinia high-pathogenicity island: an iron-uptake island. Microbes Infect 3, 561–569 (2001).
Turton, J. F., Baklan, H., Siu, L. K., Kaufmann, M. E. & Pitt, T. L. Evaluation of a multiplex PCR for detection of serotypes K1, K2 and K5 in Klebsiella sp. and comparison of isolates within these serotypes. FEMS Microbiol Lett 284, 247–252, doi: 10.1111/j.1574-6968.2008.01208.x (2008).
Feizabadi, M. M., Raji, N. & Delfani, S. Identification of Klebsiella pneumoniae K1 and K2 Capsular Types by PCR and Quellung Test. Jundishapur J Microbiol 6, e7585, doi: 10.5812/jjm.7585 (2013).
Boccia, S. et al. Genotypic analysis by 27A DNA fingerprinting of Candida albicans strains isolated during an outbreak in a neonatal intensive care unit. Infect Control Hosp Epidemiol 23, 281–284, doi: 10.1086/502052 (2002).
Lai, Y. C., Yang, S. L., Peng, H. L. & Chang, H. Y. Identification of genes present specifically in a virulent strain of Klebsiella pneumoniae. Infect Immun 68, 7149–7151 (2000).
de Souza Lopes, A. C., Falcão Rodrigues, J. & Antônio de Morais Júnior, M. Molecular typing of Klebsiella pneumoniae isolates from public hospitals in Recife, Brazil. Microbiological Research 160, 37–46, doi: 10.1016/j.micres.2004.09.007 (2005).
Ashayeri-Panah, M., Feizabadi, M. M. & Eftekhar, F. Correlation of Multi-drug Resistance, Integron and blaESBL Gene Carriage With Genetic Fingerprints of Extended-Spectrum β-Lactamase Producing Klebsiella pneumoniae. Jundishapur Journal of Microbiology 7, e8747, doi: 10.5812/jjm.8747 (2014).
Espinar, M. J. et al. Urinary Tract Infections in Kidney Transplant Patients Due to Escherichia coli and Klebsiella pneumoniae-Producing Extended-Spectrum ?-Lactamases: Risk Factors and Molecular Epidemiology. PLoS ONE 10, e0134737, doi: 10.1371/journal.pone.0134737 (2015).
Collee, J. In Mackie & Mccartney Practical Medical Microbiology (ed Collee, J. G., Fraser, A. G., Marimon, B. P., Simmons, A. ) (Elsevier, 2007).
CLSI. (Clinical and Laboratory Standards Institute, PA, USA, 2014).
EUCAST. 1–91 (European committee on antimicrobil susceptibility testing, 2016).
Cabral, A. B., Melo Rde, C., Maciel, M. A. & Lopes, A. C. Multidrug resistance genes, including bla(KPC) and bla(CTX)-M-2, among Klebsiella pneumoniae isolated in Recife, Brazil. Rev Soc Bras Med Trop 45, 572–578 (2012).
Deschaght, P. et al. Rapid genotyping of Achromobacter xylosoxidans, Acinetobacter baumannii, Klebsiella pneumoniae, Pseudomonas aeruginosa and Stenotrophomonas maltophilia isolates using melting curve analysis of RAPD-generated DNA fragments (McRAPD). Research in Microbiology 162, 386–392, doi: 10.1016/j.resmic.2011.02.002 (2011).
de la Puente-Redondo, V. A., del Blanco, N. G., Gutierrez-Martin, C. B., Garcia-Pena, F. J. & Rodriguez Ferri, E. F. Comparison of different PCR approaches for typing of Francisella tularensis strains. J Clin Microbiol 38, 1016–1022 (2000).
Siu, L. K. et al. Molecular Typing and Virulence Analysis of Serotype K1 Klebsiella pneumoniae Strains Isolated from Liver Abscess Patients and Stool Samples from Noninfectious Subjects in Hong Kong, Singapore, and Taiwan. Journal of Clinical Microbiology 49, 3761–3765, doi: 10.1128/JCM.00977-11 (2011).
Fang, C. T. et al. Klebsiella pneumoniae genotype K1: an emerging pathogen that causes septic ocular or central nervous system complications from pyogenic liver abscess. Clin Infect Dis 45, 284–293, doi: 10.1086/519262 (2007).
Author information
Authors and Affiliations
Contributions
Dr. Reham Wasfi, Dr. Walid F. Elkhatib, and Dr. Hossam M. Ashour contributed to the design of the study, performance of experiments, analysis of the results, and writing of the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Wasfi, R., Elkhatib, W. & Ashour, H. Molecular typing and virulence analysis of multidrug resistant Klebsiella pneumoniae clinical isolates recovered from Egyptian hospitals. Sci Rep 6, 38929 (2016). https://doi.org/10.1038/srep38929
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep38929
This article is cited by
-
Review and analysis of the overlapping threats of carbapenem and polymyxin resistant E. coli and Klebsiella in Africa
Antimicrobial Resistance & Infection Control (2023)
-
The notable relatedness between ESBL producing Enterobacteriaceae isolated from clinical samples and asymptomatic fecal carriers
BMC Infectious Diseases (2023)
-
Crippling of Klebsiella pneumoniae virulence by metformin, N-acetylcysteine and secnidazole
BMC Microbiology (2023)
-
Modulating the transcriptomic profile of multidrug-resistant Klebsiella pneumoniae biofilm formation by antibiotics in combination with zinc sulfate
Annals of Clinical Microbiology and Antimicrobials (2023)
-
Virulence analysis and antibiotic resistance of Klebsiella pneumoniae isolates from hospitalised patients in Poland
Scientific Reports (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.