Abstract
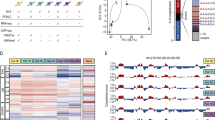
The mechanism by which the p53 family of proteins coordinately regulates select target genes after various types of cell stress is not well understood. To further define factors that dictate regulation of target genes, we examined the binding of p53, ΔNp63α and RNA polymerase II (pol II) to the regulatory regions of select target genes in primary human epidermal keratinocytes (HEKs) using chromatin immunoprecipitation. In rapidly proliferating cells, we observed constitutive binding of ΔNp63α and varying levels of p53 binding, to consensus sites in target genes involved in cell cycle arrest, DNA repair and apoptosis. Following genotoxic stress, p53 occupancy increased whereas ΔNp63α occupancy decreased at the majority of binding sites examined. Microarray analysis of transcripts isolated from HEKs ectopically expressing p53 and ΔNp63α revealed an inverse regulation of select target genes by the two family members. Collectively, our results suggest that ΔNp63α can function as a repressor of select p53 target genes involved in growth arrest, DNA repair and apoptosis, and that the location of the p53 consensus binding site(s) in a target gene may dictate whether pol II is constitutively bound in proliferating cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Appella E, Anderson CW . (2001). Post-translational modifications and activation of p53 by genotoxic stresses. Eur J Biochem 268: 2764–2772.
Bakkers J, Hild M, Kramer C, Furutani-Seiki M, Hammerschmidt M . (2002). Zebrafish DeltaNp63 is a direct target of Bmp signaling and encodes a transcriptional repressor blocking neural specification in the ventral ectoderm. Dev Cell 2: 617–627.
Barbieri CE, Perez CA, Johnson KN, Ely KA, Billheimer D, Pietenpol JA . (2005). IGFBP-3 is a direct target of transcriptional regulation by DeltaNp63alpha in squamous epithelium. Cancer Res 65: 2314–2320.
Barbieri CE, Tang LJ, Brown KA, Pietenpol JA . (2006). Loss of p63 leads to increased cell migration and up-regulation of genes involved in invasion and metastasis. Cancer Res 66: 7589–7597.
Beretta C, Chiarelli A, Testoni B, Mantovani R, Guerrini L . (2005). Regulation of the cyclin-dependent kinase inhibitor p57Kip2 expression by p63. Cell Cycle 4: 1625–1631.
Bode AM, Dong Z . (2004). Post-translational modification of p53 in tumorigenesis. Nat Rev Cancer 4: 793–805.
Dohn M, Zhang S, Chen X . (2001). p63alpha and DeltaNp63alpha can induce cell cycle arrest and apoptosis and differentially regulate p53 target genes. Oncogene 20: 3193–3205.
Dumaz N, Meek DW . (1999). Serine15 phosphorylation stimulates p53 transactivation but does not directly influence interaction with HDM2. EMBO J 18: 7002–7010.
El-Deiry WS, Tokino T, Velculescu VE, Levy DB, Parsons R, Trent JM et al. (1993). WAF1, a potential mediator of p53 tumor suppression. Cell 75: 817–825.
Espinosa JM, Verdun RE, Emerson BM . (2003). p53 functions through stress- and promoter-specific recruitment of transcription initiation components before and after DNA damage. Mol Cell 12: 1015–1027.
Harms K, Nozell S, Chen X . (2004). The common and distinct target genes of the p53 family transcription factors. Cell Mol Life Sci 61: 822–842.
Hermeking H, Lengauer C, Polyak K, He T-C, Zhang L, Thiagalingam S et al. (1997). 14-3-3 sigma is a p53-regulated inhibitor of G2/M progression. Mol Cell 1: 3–11.
Ihrie RA, Marques MR, Nguyen BT, Horner JS, Papazoglu C, Bronson RT et al. (2005). Perp is a p63-regulated gene essential for epithelial integrity. Cell 120: 843–856.
Jacobs WB, Govoni G, Ho D, Atwal JK, Barnabe-Heider F, Keyes WM et al. (2005). p63 is an essential proapoptotic protein during neural development. Neuron 48: 743–756.
Kaeser MD, Iggo RD . (2002). Chromatin immunoprecipitation analysis fails to support the latency model for regulation of p53 DNA binding activity in vivo. Proc Natl Acad Sci USA 99: 95–100.
Kaeser MD, Iggo RD . (2004). Promoter-specific p53-dependent histone acetylation following DNA damage. Oncogene 23: 4007–4013.
Lambert PF, Kashanchi F, Radonovich MF, Shiekhattar R, Brady JN . (1998). Phosphorylation of p53 serine 15 increases interaction with CBP. J Biol Chem 273: 33048–33053.
Muller M, Wilder S, Bannasch D, Israeli D, Lehlbach K, Li-Weber M et al. (1998). p53 activates the CD95 (APO-1/Fas) gene in response to DNA damage by anticancer drugs. J Exp Med 188: 2033–2045.
Oda E, Ohki R, Murasawa H, Nemoto J, Shibue T, Yamashita T et al. (2000). Noxa, a BH3-only member of the Bcl-2 family and candidate mediator of p53-induced apoptosis. Science 288: 1053–1058.
Qian H, Wang T, Naumovski L, Lopez CD, Brachmann RK . (2002). Groups of p53 target genes involved in specific p53 downstream effects cluster into different classes of DNA binding sites. Oncogene 21: 7901–7911.
Sasaki Y, Naishiro Y, Oshima Y, Imai K, Nakamura Y, Tokino T . (2005). Identification of pigment epithelium-derived factor as a direct target of the p53 family member genes. Oncogene 24: 5131–5136.
Szak ST, Mays D, Pietenpol JA . (2001). Kinetics of p53 binding to promoter sites in vivo. Mol Cell Biol 21: 3375–3386.
Tan T, Chu G . (2002). p53 Binds and activates the xeroderma pigmentosum DDB2 gene in humans but not mice. Mol Cell Biol 22: 3247–3254.
Tanaka H, Arakawa H, Yamaguchi T, Shiraishi K, Fukuda S, Matsui K et al. (2000). A ribonucleotide reductase gene involved in a p53-dependent cell-cycle checkpoint for DNA damage. Nature 404: 42–49.
Weinberg RL, Veprintsev DB, Bycroft M, Fersht AR . (2005). Comparative binding of p53 to its promoter and DNA recognition elements. J Mol Biol 348: 589–596.
Westfall MD, Mays DJ, Sniezek JC, Pietenpol JA . (2003). The Delta Np63 alpha phosphoprotein binds the p21 and 14-3-3 sigma promoters in vivo and has transcriptional repressor activity that is reduced by Hay-Wells syndrome-derived mutations. Mol Cell Biol 23: 2264–2276.
Westfall MD, Pietenpol JA . (2004). p63: Molecular complexity in development and cancer. Carcinogenesis 25: 857–864.
Zauberman A, Flusberg D, Haupt Y, Barak Y, Oren M . (1995). A functional p53-responsive intronic promoter is contained within the human mdm2 gene. Nucleic Acids Res 23: 2584–2592.
Acknowledgements
We thank Joaquin Espinosa and Beverly Emerson for helpful suggestions on optimizing ChIP PCR conditions; Lucy Tang for her assistance with the Western blotting; Jennifer Rosenbluth for protein lysates; and Carmen Perez for her assistance with microarray data analysis. This work was supported by the National Institutes of Health Grants (CA70856 and CA105436 (JAP), ES00267 and CA68485 (Core services), as well as training grant T32 ES07028 (KLS)).
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on the Oncogene website (http://www.nature.com/onc).
Supplementary information
Rights and permissions
About this article
Cite this article
Schavolt, K., Pietenpol, J. p53 and ΔNp63α differentially bind and regulate target genes involved in cell cycle arrest, DNA repair and apoptosis. Oncogene 26, 6125–6132 (2007). https://doi.org/10.1038/sj.onc.1210441
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1210441
Keywords
This article is cited by
-
E47 upregulates ΔNp63α to promote growth of squamous cell carcinoma
Cell Death & Disease (2021)
-
The homeoprotein DLX3 and tumor suppressor p53 co-regulate cell cycle progression and squamous tumor growth
Oncogene (2016)
-
Characterization, expression and silencing by RNAi of p53 from Penaeus monodon
Molecular Biology Reports (2016)
-
PIR2/Rnf144B regulates epithelial homeostasis by mediating degradation of p21WAF1 and p63
Oncogene (2013)
-
ISG20L1 is a p53 family target gene that modulates genotoxic stress-induced autophagy
Molecular Cancer (2010)