Abstract
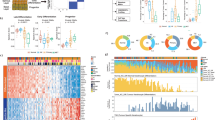
Basal cell carcinomas (BCC), which are the most common form of skin malignancy, are invariably associated with the deregulation of the Sonic Hedgehog (Shh) signalling pathway. As such, BCC represent a unique model for the study of interactions of the Shh pathway with other genes and pathways. We constructed a tissue microarray (TMA) of 75 paired BCC and normal skin and analysed the expression of β-catenin and RUNX3, nuclear effectors of the wingless-Int (Wnt) and bone morphogenetic protein/transforming growth factor-β pathways, respectively. In line with previous reports, we observed varying subcellular expression pattern of β-catenin in BCC, with 31 cases (41%) showing nuclear accumulation. In contrast, all the BCC cases tested by the TMA showed RUNX3 protein uniformly overexpressed in the nuclei of the cancer cells. Analysis by Western blotting and DNA sequencing indicates that the overexpressed protein is normal and full-length, containing no mutation in the coding region, implicating RUNX3 as an oncogene in certain human cancers. Our results indicate that although the deregulation of Wnt signalling could contribute to the pathogenesis of a subset of BCC, RUNX3 appears to be a universal downstream mediator of a constitutively active Shh pathway in BCC.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Andl T, Reddy ST, Gaddapara T, Millar SE . (2002). Dev Cell 2: 643–653.
Bae SC, Choi JK . (2004). Oncogene 23: 4336–4340.
Behrens J, von Kries JP, Kuhl M, Bruhn L, Wedlich D, Grosschedl R et al. (1996). Nature 382: 638–642.
Blyth K, Cameron ER, Neil JC . (2005). Nat Rev Cancer 5: 376–387.
Boonchai W, Walsh M, Cummings M, Chenevix-Trench G . (2000). Arch Dermatol 136: 937–938.
Cameron ER, Neil JC . (2004). Oncogene 23: 4308–4314.
Chan EF, Gat U, McNiff JM, Fuchs E . (1999). Nat Genet 21: 410–413.
Chen YG, Meng AM . (2004). Cell Res 14: 441–449.
Dahmane N, Lee J, Robins P, Heller P, Ruiz i Altaba A . (1997). Nature 389: 876–881.
Daya-Grosjean L, Couve-Privat S . (2005). Cancer Lett 225: 181–192.
El-Bahrawy M, El-Masry N, Alison M, Poulsom R, Fallowfield M . (2003). Br J Dermatol 148: 964–970.
Gianakopoulos PJ, Skerjanc IS . (2005). J Biol Chem 280: 21022–21028.
Goodrich LV, Johnson RL, Milenkovic L, McMahon JA, Scott MP . (1996). Genes Dev 10: 301–312.
Gregorieff A, Clevers H . (2005). Genes Dev 19: 877–890.
Huber O, Korn R, McLaughlin J, Ohsugi M, Herrmann BG, Kemler R . (1996). Mech Dev 59: 3–10.
Ito K, Liu Q, Salto-Tellez M, Yano T, Tada K, Ida H et al. (2005). Cancer Res 65: 7743–7750.
Ito Y, Miyazono K . (2003). Curr Opin Genet Dev 13: 43–47.
Kawai S, Sugiura T . (2001). Bone 29: 54–61.
Kim WJ, Kim EJ, Jeong P, Quan C, Kim J, Li QL et al. (2005). Cancer Res 65: 9347–9354.
Ku JL, Kang SB, Shin YK, Kang HC, Hong SH, Kim IJ et al. (2004). Oncogene 23: 6736–6742.
Li QL, Ito K, Sakakura C, Fukamachi H, Inoue K, Chi XZ et al. (2002). Cell 109: 113–124.
Logan CY, Nusse R . (2004). Annu Rev Cell Dev Biol 20: 781–810.
Marigo V, Tabin CJ . (1996). Proc Natl Acad Sci USA 93: 9346–9351.
Mori T, Nomoto S, Koshikawa K, Fujii T, Sakai M, Nishikawa Y et al. (2005). Liver Int 25: 380–388.
Reddy S, Andl T, Bagasra A, Lu MM, Epstein DJ, Morrisey EE et al. (2001). Mech Dev 107: 69–82.
Rubin AI, Chen EH, Ratner D . (2005). N Engl J Med 353: 2262–2269.
Rubinfeld B, Robbins P, El-Gamil M, Albert I, Porfiri E, Polakis P . (1997). Science 275: 1790–1792.
Saldanha G, Ghura V, Potter L, Fletcher A . (2004). Br J Dermatol 151: 157–164.
Salto-Tellez M, Lee SC, Chiu LL, Lee CK, Yong MC, Koay ES . (2004). Clin Chem 50: 1082–1086.
Unden AB, Zaphiropoulos PG, Bruce K, Toftgard R, Stahle-Backdahl M . (1997). Cancer Res 57: 2336–2340.
van Wijnen AJ, Stein GS, Gergen JP, Groner Y, Hiebert SW, Ito Y et al. (2004). Oncogene 23: 4209–4210.
Weeraratna AT . (2005). Cancer Metast Rev 24: 237–250.
Yamazaki F, Aragane Y, Kawada A, Tezuka T . (2001). Br J Dermatol 145: 771–777.
Yanada M, Yaoi T, Shimada J, Sakakura C, Nishimura M, Ito K et al. (2005). Oncol Rep 14: 817–822.
Yanagida M, Osato M, Yamashita N, Liqun H, Jacob B, Wu F et al. (2005). Oncogene 24: 4477–4485.
Ziegler A, Leffell DJ, Kunala S, Sharma HW, Gailani M, Simon JA et al. (1993). Proc Natl Acad Sci USA 90: 4216–4220.
Acknowledgements
This work was supported by the Agency for Science, Technology and Research (A*STAR), Singapore. M Salto-Tellez and SH Tan are recipients of Singapore Cancer Syndicate Grants MN005 and BS002, Agency for Science, Technology and Research, Singapore.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Salto-Tellez, M., Peh, B., Ito, K. et al. RUNX3 protein is overexpressed in human basal cell carcinomas. Oncogene 25, 7646–7649 (2006). https://doi.org/10.1038/sj.onc.1209739
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1209739
Keywords
This article is cited by
-
Small peptide inhibitor from the sequence of RUNX3 disrupts PAK1–RUNX3 interaction and abrogates its phosphorylation-dependent oncogenic function
Oncogene (2021)
-
RUNX3 suppresses metastasis and stemness by inhibiting Hedgehog signaling in colorectal cancer
Cell Death & Differentiation (2020)
-
RUNX3 is oncogenic in natural killer/T-cell lymphoma and is transcriptionally regulated by MYC
Leukemia (2017)
-
Inhibition of Hedgehog-dependent tumors and cancer stem cells by a newly identified naturally occurring chemotype
Cell Death & Disease (2016)
-
The RUNX family: developmental regulators in cancer
Nature Reviews Cancer (2015)