Abstract
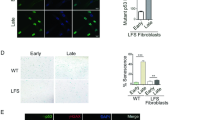
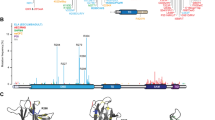
p53 is a tumour suppressor gene which functions as a transcription factor to upregulate genes for growth arrest and apoptosis following DNA damage. p53 mutations are associated with Li–Fraumeni and Li–Fraumeni like syndromes. Recently mutations of the oligomerization domain have been isolated from an LFS and an LFL family affecting respectively codon 344 (Leu to Pro) and 337 (Arg to Cys). The present study was designed to determine the affect of these mutations on the function of p53 protein. p53 344 Leu to Pro existed only in a monomeric form and could not bind to DNA. It was inactive at inducing apoptosis, transactivating luciferase from a bax promoter and inhibiting cell growth. In contrast, p53 337 Arg to Cys could form tetramers and could bind to DNA. However, p53 337 Arg to Cys was not fully active and could only induce apoptosis, transactivate luciferase from a bax promoter and inhibit cell growth with ≈amp;60% of the ability of wild-type p53. Both mutant proteins had reduced ability to bind to MDM2, p53 337 Arg to Cys being more reduced than p53 344 Leu to Pro. These results indicate that point mutations in the oligomerization domain can disrupt p53 function. In addition, the value of LFS and LFL families for the further understanding of the biological and biochemical properties of p53 is demonstrated.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Lomax, M., Barnes, D., Hupp, T. et al. Characterization of p53 oligomerization domain mutations isolated from Li–Fraumeni and Li–Fraumeni like family members. Oncogene 17, 643–649 (1998). https://doi.org/10.1038/sj.onc.1201974
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1201974
Keywords
This article is cited by
-
Therapeutic targeting of p53: all mutants are equal, but some mutants are more equal than others
Nature Reviews Clinical Oncology (2018)
-
Cotranslational protein assembly imposes evolutionary constraints on homomeric proteins
Nature Structural & Molecular Biology (2018)
-
Current understanding of the role and targeting of tumor suppressor p53 in glioblastoma multiforme
Tumor Biology (2013)
-
Occurrence of multipolar mitoses and association with Aurora-A/-B kinases and p53 mutations in aneuploid esophageal carcinoma cells
BMC Cell Biology (2011)
-
Conformational stability and activity of p73 require a second helix in the tetramerization domain
Cell Death & Differentiation (2009)