Abstract
Mutations in MECP2 are a cause of Rett syndrome. Recently, a new isoform of MeCP2 was described, which has an alternative N-terminus, transcribed from exon 1. We screened exon 1 and the promoter region of MECP2 in 97 mutation-negative Rett syndrome cases. We found two sequence variants, but there was no evidence that they are pathogenic. Mutations in exon 1 and the promoter of MECP2 are not a common cause of Rett syndrome.
Similar content being viewed by others
Introduction
Two recent studies have described a new isoform of MeCP2 and shown that it is the predominant form of the protein in the human brain.1,2 The new isoform has an alternative N-terminus, transcribed from exon 1 of MECP2. Exon 1 was previously thought to be noncoding and is therefore not included in genetic tests for Rett syndrome.
Mutations in MECP2 can cause Rett Syndrome,3 a neurodevelopmental X-linked disorder that predominantly affects female populations and is the most common genetic cause of profound intellectual disability in female subjects. Typically, there is a period of near-normal development followed by a regression period with loss of social, motor and communication skills. Other features of Rett syndrome include autonomic disturbance, scoliosis, feeding difficulties and epilepsy. Most cases of Rett syndrome are sporadic, with the majority being caused by a de novo mutation on the paternal copy of MECP2.4
Two cases have been described in which Rett syndrome was caused by mutation or deletion of MECP2 exon 1 (from a sample of 19 classical Rett MECP2 mutation-negative cases).1 An additional case has been described where exons 1 and 2 were deleted.5 It is possible that exon 1 mutations account for a significant proportion of the ∼15% of classical Rett cases who do not have mutations in exons 2, 3 or 4 of MECP2. We assessed the sequence variation in exon 1 and the promoter region of MECP2 in 243 individuals: 97 cases of Rett syndrome with no mutation found in exons 2, 3 or 4 of MECP2 and 146 controls.
Methods
Samples
DNA samples were obtained from 97 Rett syndrome patients who have previously had a negative MECP2 test (by sequencing exons 2, 3 and 4). Patients were defined as classical Rett (n = 37) or atypical Rett (n = 60) according to the criteria described in Kerr et al.6 The control DNA samples were obtained from 146 anonymous, healthy adults.
MECP2 exon 1 sequencing
PCR was carried out using the primers EX1_F 5′-gcactcggtgcatctgtggacagag-3′ and EX1_R 5′-catccgccagccgtgtcgtccgac-3′ in a [NH4]2SO4-based buffer with 3.7 mM. MgCl2, 750 μM dNTPs, 0.5 M betaine, 5% DMSO and Platinum Taq (Invitrogen). The annealing temperature was 65°C. Sequencing was performed with a BigDye v3.1 sequencing kit (Applied Biosystems) according to the manufacturer's instructions except for the addition of 1 M betaine. The template was sequenced from each end using the PCR primers and from the middle using two internal primers: INT1_F 5′-caattgacggcatcgccgctgagac-3′ and INT1_R 5′-cgtcattggctgtgatggc-3′.
Multiplex ligation-dependent probe amplification
The girls with classical Rett syndrome were tested for deletions using a MECP2 Multiplex ligation-dependent probe amplification (MLPA) kit (MRC Holland) according to the instructions provided.
Assay for −219_220insC
This 1 bp insertion creates a Bsa I restriction site. Samples were amplified with primers EX1_F and EX1_R. The PCR products were digested overnight at 50°C with 5 U Bsa I (New England Biolabs) and run on a 2% agarose gel. Normal DNA remains undigested (size 672 bp), DNA with the insertion is digested to 270- and 402- bp fragments.
Assay for 3_4insGCCGCC
Samples were amplified using primer EX1_R and a fluorescent INT1_F primer; the PCR products were run on an ABI 3100 (Applied Biosystems). The normal PCR product size is 419 bp and with the insertion it is 425 bp.
X-inactivation
Aliquots of DNA were pre-digested with the methylation-sensitive enzymes HpaII and McrBC (New England Biolabs). The HUMARA locus was then amplified using fluorescent primers and the allele peak areas analysed using an ABI 3100 and Genotyper software (Applied Biosystems).
Results
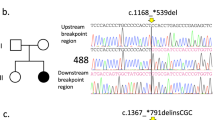
We sequenced MECP2 exon 1 and approximately 400 bp of the promoter region in a total of 97 individuals with Rett syndrome who do not have mutations in exon 2, 3 or 4. We found two changes from the normal sequence and we assessed their frequency by genotyping 146 controls (Table 1, Figure 1 Supplementary Figure). The first variant is a 1- bp insertion at position −220 relative to the start codon in exon 1 (g.-219_220insC, Figure 1a). Note the reference sequence, AF030876, has the insertion (GenBank/EMBL/DDBJ). The second variant is a 6- bp insertion in the [GCC]6 trinucleotide repeat at the start of exon 1 (c.3_4insGCCGCC, Figure 1b).
Deletion analysis by MLPA was successful in 20 classical Rett girls. One girl was found to have a deletion of exon 4 and she was therefore excluded from this study. A further six of the classical Rett girls have previously had deletions in exon 2, 3 and 4 excluded by quantitative fluorescent PCR (data not shown). We were unable to exclude deletions in the 12 remaining classical Rett girls due to variable DNA quality. Based on our experience in diagnostic testing for MECP2 mutations, we would expect to find a deletion in approximately 5% of similar MECP2-negative Rett patients. Deletions were excluded in 29 of the atypical Rett girls by MLPA.
Discussion
The 1- bp insertion (g. -219_220insC) appears to be a rare polymorphism since it was seen in 3% of Rett cases and 3% of controls (Table 1). Three of the controls with the insertion were male and are therefore hemizygous for MECP2. This confirms that the 1- bp insertion is not pathogenic.
The 6- bp insertion (c.3_4insGCCGCC) is predicted to result in insertion of an extra two alanine residues in the polyalanine sequence at the N-terminus of the recently described MeCP2 splice variant. It was found in one girl with classical Rett syndrome and her unaffected mother. Skewed X-inactivation can sometimes mask the pathogenic effect of a MECP2 mutation,7 but in this case X-inactivation was random in lymphocyte DNA from both the girl and her mother (X-inactivation was 54% in the mother and 60% in the girl). It is possible that there is skewed X-inactivation in the brain of the girl or the mother, but it is not feasible to test this. It seems unlikely that c.3_4insGCCGCC is pathogenic, although we cannot rule out this possibility because we did not find this variant in the controls.
Exon 1 of MECP2 contains two trinucleotide repeat tracts, [GCC]6 and [GGA]5. It has been suggested that expansion of these repeat tracts could be pathogenic, similar to other triplet repeat disorders such as fragile X syndrome.2 We observed a small expansion (c.3_4insGCCGCC) in the first repeat tract. This change is probably not pathogenic, but it demonstrates that expansions can occur. It is interesting to note that the mutation described by Mnatzakanian et al1 was also in one of the repeat tracts (an 11- bp deletion in the [GGA]5 repeat).
We conclude that mutations in exon 1 and the promoter region of MECP2 are not a common cause of Rett syndrome. In a substantial number of classical Rett cases, the cause remains unknown.
References
Mnatzakanian GN, Lohi H, Munteanu I et al: A previously unidentified MECP2open reading frame defines a new protein isoform relevant to Rett syndrome. Nat Genet 2004; 36: 339–341.
Kriaucionis S, Bird A : The major form of MeCP2 has a novel N-terminus generated by alternative splicing. Nucleic Acids Res 2004; 32: 1818–1823.
Amir RE, Van den Veyver IB, Wan M, Tran CQ, Francke U, Zoghbi HY : Rett syndrome is caused by mutations in X-linked MECP2, encoding methyl-CpG-binding protein 2. Nat Genet 1999; 23: 185–188.
Trappe R, Laccone F, Cobilanschi J et al: MECP2 mutations in sporadic cases of Rett syndrome are almost exclusively of paternal origin. Am J Hum Genet 2001; 68: 1093–1101.
Erlandson A, Samuelsson L, Hagberg B, Kyllerman M, Vujic M, Wahlström J : Multiplex ligation-dependent probe amplification (MLPA) detects large deletions in the MECP2 gene of Swedish Rett syndrome patients. Genet Test 2003; 7: 329–332.
Kerr AM, Nomura Y, Armstrong D et al: Guidelines for reporting clinical features in cases with MECP2 mutations. Brain Dev 2001; 23: 208–211.
Wan M, Sung Jae Lee S, Zhang X et al: Rett syndrome and beyond: recurrent spontaneous and familial MECP2 mutations at CpG hotspots. Am J Hum Genet 1999; 65: 1520–1529.
Acknowledgements
We thank the patients, families and clinicians who helped in this study. This work was funded by The Health Foundation, a healthcare charity.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on European Journal of Human Genetics website (http://www.nature.com/ejhg).
Supplementary information
Rights and permissions
About this article
Cite this article
Evans, J., Archer, H., Whatley, S. et al. Variation in exon 1 coding region and promoter of MECP2 in Rett syndrome and controls. Eur J Hum Genet 13, 124–126 (2005). https://doi.org/10.1038/sj.ejhg.5201270
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.ejhg.5201270
Keywords
This article is cited by
-
Rett Syndrome and MeCP2
NeuroMolecular Medicine (2014)
-
Mutation analysis of the MECP2 gene in patients of Slavic origin with Rett syndrome: novel mutations and polymorphisms
Journal of Human Genetics (2007)
-
MECP2 and CDKL5 gene mutation analysis in Chinese patients with Rett syndrome
Journal of Human Genetics (2007)
-
Expression profiling of clonal lymphocyte cell cultures from Rett syndrome patients
BMC Medical Genetics (2006)
-
Rett syndrome: new clinical and molecular insights
European Journal of Human Genetics (2006)