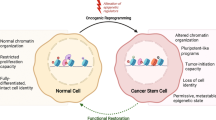
Abstract
Tumours are comprised of a highly heterogeneous population of cells, of which only a small subset of stem-like cells possess the ability to regenerate tumours in vivo. These cancer stem cells (CSCs) represent a significant clinical challenge as they are resistant to conventional cancer therapies and play essential roles in metastasis and tumour relapse. Despite this realization and great interest in CSCs, it has been difficult to develop CSC-targeted treatments due to our limited understanding of CSC biology. Here, we present evidence that specific histone deacetylases (HDACs) play essential roles in the CSC phenotype. Utilizing a novel CSC model, we discovered that the HDACs, HDAC1 and HDAC7, are specifically over-expressed in CSCs when compared to non-stem-tumour-cells (nsTCs). Furthermore, we determine that HDAC1 and HDAC7 are necessary to maintain CSCs, and that over-expression of HDAC7 is sufficient to augment the CSC phenotype. We also demonstrate that clinically available HDAC inhibitors (HDACi) targeting HDAC1 and HDAC7 can be used to preferentially target CSCs. These results provide actionable insights that can be rapidly translated into CSC-specific therapies.
Similar content being viewed by others
Introduction
The cancer stem cell (CSC) model posits that each tumour is composed of a hierarchy of cells, of which only a small subset are CSCs that possess the ability to regenerate tumours in vivo.1 The remaining tumour bulk is primarily comprised of non-stem-tumour-cells (nsTCs) that descend from CSCs but are themselves incapable of initiating or propagating tumours.
Multiple signalling pathways involved in the regulation of CSCs have been identified.1 However, the regulation of CSCs is unlike the reversible short-term changes in cellular phenotype induced by various extracellular factors, or the permanent changes induced by mutations. The hierarchical differentiation of CSCs to nsTCs is long-lasting over many cell generations but it is also reversible, that is more akin to tissue differentiation, which suggests that epigenomic factors such as histone modifications may also be involved in the regulation of the CSC phenotype.2
HDACs are chromatin-modifying enzymes that are involved in regulation of many aspects of cell biology including tissue differentiation, autophagy, apoptosis, migration, mitosis and angiogenesis.3 There are 11 different HDAC genes with distinct tissue-specific expression. The HDAC proteins are grouped by their homology to their orthologues in yeast; class I (HDAC 1-3, 8); class II a (HDAC 4, 5, 7, 9), class II b (6 and 10), and class V (HDAC 11).3 While the total pan-HDAC activity, including all family members, have been implicated in differentiation, pluripotent and embryonic stem cell regulation,4, 5, 6, 7, 8, 9, 10 the role of specific HDACs in the regulation of CSCs has not been fully explored.
The exploration of the epigenetic factors that may be involved in the regulation of CSCs has been hampered by experimental and technical challenges, which we partly address in this study. The CSCs can be isolated with fluorescence-activated cell sorting (FACS), using antibodies that recognize putative CSC cell surface markers.1, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22 However, the rare nature of CSCs in solid tumours, coupled with the methodological challenges of isolating and expanding them in vitro, has impeded the identification of the precise epigenetic regulators responsible for maintaining CSC phenotype.
It is worth pointing out that FACS enrichment of CSCs from primary tumours is not an innocuous process, because it exposes the tumour cells to many changes in temperature, CO2 and pH levels. Since isolation of CSCs by FACS requires single cell suspensions as starting material, the solid tumour fragments are first digested with proteases (such as collagenase and trypsin) at 37 °C for several hours depending on the tissue type (1–8 h). Next, the resulting single cell suspension is incubated with CSC-specific antibodies at 4 °C for 1–2 h, followed by FACS enrichment at 20 °C for another 1–2 h. At the end of this process, in our experience 101–3 fold enrichment of CSCs relative to nsTCs can be achieved by combining two to three CSC surface markers. However, if one is working with a cell line or tumour type with a low 10−5 to 6 CSC frequency, this would represent at best only one in a hundred cells in the enriched population are indeed CSCs. Therefore, one can isolate relatively CSC-enriched populations with FACS; however, these are not CSC-pure populations by any means. Lastly, even when CSC populations are successfully isolated with FACS, their phenotypic instability limits their utility, because CSCs rapidly differentiate into nsTC in standard culture.14
In summary, while FACS enrichment is used as a benchmark method to study CSCs, the lack of absolute purity, rapid dilution and potential alterations in CSC phenotype are some of the drawbacks of this approach. In this manuscript we describe a model system that addresses some of these methodological challenges, leading to identification of several HDACs involved in the regulation of CSC phenotype in several model systems.
Results
BPLER cell lines as a model of CSCs
The technical challenges associated with isolation and study of CSCs can be partly circumvented by utilizing a set of genetically engineered cell lines that functionally ‘phenocopy’ CSC behaviours. Previously we derived three independent pairs of genetically matched breast cancer cell lines, BPLER2, 3, 4 and HMLER.2, 3, 4, 23 The BPLER lines were derived from normal human breast precursor epithelial cells that have a mixed luminal-myoepithelial phenotype and maintain telomerase activity in vitro.23, 24 In contrast, HMLERs are derived from normal human mammary epithelial cells that exhibit a more differentiated myoepithelial phenotype and do not have telomerase activity.23, 24 Both high levels of hTERT activity and lack of lineage commitment have been associated with stem cells in normal tissues. Consistent with the difference in their cell-of-origin, we now demonstrate that BPLER cells recapitulate the mRNA expression profile and the biological properties of CSCs. In contrast, HMLER cells as a whole resemble nsTCs (Supplementary Figures 1–7).
Importantly, the frequency of CSCs in unsorted bulk BPLER cultures is equivalent to the CSC population obtained after FACS enrichment of standard cell lines and is stably maintained in routine 2D culture. Hence, while as few as five unsorted BPLER cells are capable of forming tumours in nude mice, the genetically matched HMLERs require injection of 103–5 cells to form tumours in vivo (Figure 1a). The CSC-like BPLERs also form significantly higher number of tumour spheres compared to matched HMLER lines (Figure 1b), which is a measure of CSC frequency in vitro.25, 26, 27, 28, 29
BPLER and HMLER cells differ in their in vivo and in vitro tumor initiating cell (TIC) frequency, and in CSC surface marker expression. (a) In serial dilution xenograft assays BPLER lines display 2–4 orders-of-magnitude (102–104) greater in vivo TIC frequency than paired HMLER cell lines. BPLER2/HMLER2, BPLER3/HMLER3, BPLER4/HMLER4 isogenic cell lines were derived from normal human breast primary cells isolated from three different donors as previously described23 (See Supplementary Figure 2a for further details). (b) BPLER lines (blue bars) demonstrate greater capacity for sphere formation than HMLER cells (red bars) in 3D sphere formation assays used as an in vitro measurement of CSC self-renewal. Data presented as a mean +/− s.d. of sphere counts from triplicate wells (P<0.05). Results are representative of at least three independent experiments. (c) BPLER lines express higher levels of the CSC markers CD326 (EpCAM/ESA), CD166 (ALCAM), and BMI-1* than HMLER. Additionally, BPLER lines express the CSC-specific CD44 isoform (CD44v-250 kDa), while HMLER express the standard CD44 isoform (CD44s-80 kDa). Western blot of whole-cell lysates. β-Actin represents loading control. *BMI-1-matched β-actin in Supplementary Fig ure 3f. (d) FACS-enriched BPLER CSCs, sorted for high expression of individual CSC markers (CD44, CD166, CD326), or a combination of all three markers, demonstrate enhanced mammosphere formation when compared to BPLER with low CSC-marker expression (black bars, CSC marker high; white bars, CSC marker low). Data presented as a mean +/− s.d. of sphere counts performed in triplicate (P<0.05). Results are representative of at least three independent experiments.
In addition, the BPLER CSC phenotype is stable, maintaining high endogenous levels of the CSC-associated markers CD166,1, 11, 12, 13, 14, 15, 16 CD326 (EpCAM, ESA),18, 19, 20 CD44v22 and BMI-12 throughout extended periods of culture (Figure 1c), and the CSC attributes of BPLER can be even further enhanced by FACS enrichment using individual CSC markers, in particular, CD44 and CD166 (Figure 1d). The unique stability of the CSC phenotype in the BPLER model enabled us to carry out the CSC-specific experiments described below, which would not have been possible with conventional cell lines.
Lastly, we found that the BPLER cells co-cluster with the triple-negative breast cancer (TNBC) cell lines (Supplementary Figure 4), TNBC patient tumours (Supplementary Figures 5 and 6), express a CSC-like mRNA expression signature identified in other cell lines (Supplementary Table 1), and recapitulate the accurate human breast adenocarcinoma morphology in xenograft tumours (Supplementary Figures 7 and 8). These results cumulatively indicate that the BPLER model at least partially phenocopies naturally occurring CSCs.
Discovery of CSC over-expression of HDAC1 and HDAC7 in BPLER CSCs
We hypothesized that specific members of the HDAC family, chromatin-modifying enzymes involved in epigenetic regulation of differentiation, might be involved in the regulation of the CSC phenotype in BPLERs. Since the CSC-like BPLER and nsTC-like HMLER cell line pairs are derived from the same patient and transformed with identical oncogenes, they are ostensibly isogenic, which makes them a good model system to survey epigenetic differences between CSC and nsTC. In order to initially examine these potential epigenetic differences, the paired BPLER/HMLER lines were treated with Trichostatin A (TSA), a pan-HDACi that inhibits all the HDAC family members. These dose-response studies reveal that BPLER proliferation is significantly more sensitive to TSA inhibition as compared to matched HMLERs in 2D culture (Figure 2a).
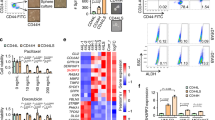
CSC-Like BPLER cells are associated with high HDAC1 and HDAC7 expression and sensitivity to pan-HDAC inhibitors. (a) The pan-HDAC inhibitor TSA preferentially inhibits BPLER proliferation (blue line), compared to HMLER (red line). The results are representative of at least three independent experiments, presented as a percentage of vehicle treated control, the error bars represent standard deviation of the mean (P<0.005). (b) Short-term (24 h) pretreatment with TSA (0.35 μM), preferentially inhibits BPLER sphere formation (3D) in drug-free medium with no effect on 2D proliferation in either BPLER or HMLER. In contrast, pretreatment with Taxol (50 nM) and 5-Fluorouracil (1.0 μM), preferentially inhibit 2D proliferation as compared to 3D sphere formation. The number of viable colonies from triplicate wells were determined by 2-(4-iodophenyl)-3-(4-nitrophenyl)-5-phenyl-2H-tetrazolium chloride (INT) staining. Results are representative of at least three independent experiments presented as percentage of vehicle treated control (P<0.05). BPLER: 2D proliferation (white bars with blue outline) vs 3D sphere formation (blue bars). HMLER: 2D proliferation (white bars with red outline) vs 3D sphere formation (red bars). The error bars represent standard deviation of the mean. (c) BPLER cell lines express higher levels of HDAC1 and HDAC7 proteins compared to matched HMLER lines. Western blot of whole-cell lysates. β-Actin represents loading control. (d) Heatmap of the mRNA expression profile of HDAC1-11 does not reveal any consistent differences between BPLER and HMLER lines (red, increased expression; green, decreased expression). (e) Treatment of BPLER cells for 48 h with TSA (0.35 μM) leads to downregulation of HDAC1, HDAC7, CD44 and CD166 protein expression in BPLER cells. Western blot of whole-cell lysates. β-Actin represents loading control. (f) Double immunofluorescence staining of BPLER cells simultaneously with HDAC7 and CD44 antibodies demonstrate that HDAC7 and CD44 are co-expressed. DAPI (blue), HDAC7 (red) CD44 (green). Scale bar 25 μM.
Next, we tested the effect of short-term (24 h) TSA treatment on subsequent 2D proliferation in routine culture vs 3D sphere formation, a surrogate assay for measuring CSC frequencies. Interestingly, we found that TSA pretreatment preferentially inhibits BPLER sphere formation (3D growth). In contrast, standard chemotherapeutics preferentially inhibit HMLERs and 2D proliferation (Figure 2b, Supplementary Figures 9a and b). Furthermore, these conventional chemotherapeutics (Taxol, 5-fluorouracil and Doxorubicin) preferentially induce apoptosis in HMLERs, but not BPLERs (Supplementary Figure 9c), consistent with the chemotherapeutic resistance typically associated with CSCs. Finally, TSA pretreatment preferentially inhibits 3D growth of standard breast cancer cell lines as compared to 2D proliferation (Supplementary Figure 9d). These findings demonstrate that short-term treatment with a pan-HDACi has lasting effects on sphere-initiating capacity in multiple models, and this effect is not simply due to a non-specific inhibition of cell proliferation.
These results prompted us to evaluate whether specific HDAC family members are expressed at higher levels in CSC-like BPLERs compared to nsTC-like HMLERs. We found that only two of the 11 HDAC family members, HDAC1 and HDAC7, were consistently expressed at significantly higher levels in BPLER lines at the protein level (Figure 2c). Interestingly, there is no consistent difference at the mRNA level between BPLER and HMLER lines for any of the HDACs 1–11 (Figure 2d), suggesting that these differences are maintained at the protein level in CSCs. The regulation of HDACs at the protein level may be one reason why the association between CSCs and HDAC1/7 has not been identified in previous mRNA profiling experiments. Furthermore, we found that TSA treatment of BPLER lines downregulates HDAC1 and HDAC7 protein expression, as well as CD44 and CD166, suggesting that HDAC-dependent mechanisms may regulate these CSC markers (Figure 2e). Consistent with this, immunofluorescent (IF) staining of unsorted BPLER cells revealed that CD44high/CD166high-CSC subpopulations express higher levels of HDAC7 (and HDAC1, data not shown) compared to CD44low or CD166low cells (Figure 2f).
Verification of HDAC1 and HDAC7 over-expression in primary tumour and cell line CSCs
In order to verify these results in other models, we used FACS to isolate CSCs. Consistent with previous reports, we observed that CD44high/CD166high populations have greater sphere-forming capacity compared to (CD44low/CD166low) counterparts (Figure 3a)(1, 11, 12, 13, 14). CD44high/CD166high CSCs from multiple breast (SUM159, MDA-MB-231, MCF7), ovarian (SKOV3, OV90) and colon (HT29) tumour cell lines. These FACS-enriched CSC subpopulations demonstrate increased HDAC enzymatic activity (Figure 3b), and in agreement with the BPLER/HMLER system, consistently expressed significantly higher HDAC1 and HDAC7 levels compared to paired nsTC populations (Figure 3c, Supplementary Figure 10a). Consistent with these findings, IF staining confirmed that CD44high/CD166high cells express higher levels of HDAC7 compared to CD44low/CD166low cells (Figure 3d, Supplementary Figures 10b and c). While several other HDAC family members have variable expression patterns in sorted CSCs, among these HDAC1 and HDAC7 are most consistently associated with CSCs in all cell lines examined.
CSCs isolated from standard breast and ovarian cancer cell lines express higher protein levels of HDAC1 and HDAC7, and have increased HDAC enzymatic activity. (a) The higher sphere forming capacity of CD44high/CD166high CSCs (black bars) compared to CD44low/CD166low nsTC cells (white bars) is observed in multiple breast (SUM159, MDA-231, and MCF7) and ovarian (SKOV3 and OV90) cell lines. The data are presented as the mean of sphere counts from triplicate wells, the error bars represent standard deviation of the mean (P<0.01). The results are representative of at least three independent experiments. (b) The higher HDAC enzyme activity is observed CD44high/CD166high CSCs (black bars) compared to CD44low/CD166low nsTCs (white bars) in multiple breast (SUM159, MDA-231, and MCF7) and ovarian (SKOV3 and OV90) and colon (HT29) cancer cell lines. Results are representative of at least three independent experiments. The error bars represent standard deviation of the mean (P<0.01). (c) HDAC1 and HDAC7 protein expression is significantly higher in CD44high/CD166high CSCs (+/+) from multiple breast (BPLER, MDA-MB-231, MCF7, SUM159) and ovarian (SKOV3) cancer cell lines compared to nsTC (−/−, CD44low/CD166low). Merged image of two Western blots run with whole-cell lysates, gel#1(BPLER, MDA-MB-231, MCF7, SUM159) and gel#2(SKOV3). β-Actin represents loading control. (d) HDAC7high (red) and CD44high (green) co-expression in ovarian cancer cell line SKOV3, demonstrated with double IF staining. Scale bar 25 μM. (e) Double IF staining identifies co-expression of CSC-markers (CD44, CD326, CD166, ALDH1, CD29) with HDAC7 in standard breast cell lines (MDA-MB-231, MCF7, SUM159), and primary human ovarian (OCI-P5x and OCI-E1p) and breast cancer cells (BCI-1009 and BCI-1133). (Y) represents consistent positive correlation of HDAC7 and CSC marker expression in surveyed lines. (y) represents positive, but inconsistent correlation. (N) represents lack of correlation. (n/a), not analyzed. (n/e) CSC marker not expressed in cell line. IF staining was repeated a minimum of two times for each line. See Supplementary Figure 10 for additional markers and cell lines.
In addition to CD44 and CD166, we verified the association between HDAC7 overexpression and CSC phenotype using six additional CSC markers (ALDH1a1, ALDH1a3, CD29, CD44v, CD49f and CD326) in a panel of seven additional breast and ovarian cancer lines. These experiments identified a broad correlation between high HDAC7 protein expression with a variety of well-established CSC markers (Figure 3e, Supplementary Figure 10d). These results were also confirmed in tumour cells freshly isolated from seven primary human ovarian and breast cancers,30 and in formalin-fixed paraffin-embedded sections of primary human breast cancer tissues (Supplementary Figure 11). In sum, we observed a correlation between eight CSC markers and high HDAC7 protein expression in seven cell lines and seven primary tumours using IF staining. These results establish an association between high HDAC1/7 protein and CSCs in multiple cell line models and primary tumour cells.
Re-identification of HDAC inhibitors through a BPLER siRNA lethality screen
Reassuringly, HDAC sensitivity of the BPLER lines was confirmed in an independent genome-wide high-throughput unbiased siRNA lethality screen.31 A genome-wide siRNA screen was used to identify 154 genes that are specifically lethal when inhibited in BPLER cells but not in HMLER cells.31 This signature correlates with poor survival and early onset of metastasis in breast cancer patients independent of tumour subtype.31 We interrogated the mRNA expression of these 154 genes in the Broad Institute connectivity map (CMAP), which contains mRNA expression data from cancer cell lines exposed to 1309 compounds in over 7000 experiments.31 This analysis identified 37 compounds that preferentially reduced the expression of 154 BPLER-lethality genes in other cell lines (Supplementary Table 2). Interestingly, the HDACi, TSA, was the top candidate that emerged out of this in silico screen, and Vorinostat (SAHA) was the fourth most significant (Figure 4a). Next, we evaluated the dose-response of BPLER vs HMLER cells to the top 10 compounds, and found that only the pan-HDACis, TSA and SAHA have differential efficacy towards BPLER (Figure 4b). Interestingly, the other eight in silico candidate compounds did not exhibit any differential effects on BPLER vs HMLER cells (Figure 4b, Supplementary Figure 12). Hence, we found the same differential HDAC sensitivity between BPLER and HMLER cells, both by a hypothesis-driven candidate-based approach and a high-throughput unbiased approach. Remarkably, the two HDAC inhibitors were the only two candidate hits that were confirmed from the high-throughput screen.
Confirmation of CSC sensitivity to pan-HDAC inhibitors in other model systems. (a) The black lines within the red and green regions indicate different CMAP experiments that display significant upregulation (red) or downregulation (green) of the 154 BPLER lethality genes upon treatment with the indicated drugs. Black lines in the gray area indicate CMAP experiments with no significant variation. (b) The percent viability of BPLER and HMLER cells that were treated with TSA, Vorinostat, Loperamide and Triamterene at the indicated doses. A vehicle-treated control was used to estimate relative percent cell viability for each treatment. BPLER (blue line), HMLER cells (red line). The error bars represent standard deviation of the mean (P<0.005). (c) HDAC1, HDAC7 and CD166 proteins are expressed significantly higher in the CD44high/CD24low+ CSCs (CD24L) compared to CD44high/CD24Neg ns-TS (CD24N) MDA-MB-231 cells. Western blot of whole-cell lysates. β-Actin represents loading control. (d) The mRNA expression heatmap of CD44high/CD24low+ CSCs (CD24L) compared to CD44high/CD24Neg nsTC (CD24N) MDA-MB-231 cells, with high (red) and low (green) expression. The difference in expression for any HDAC family members between the two populations was less than 1.1 fold. (e) The sensitivity of CD24-negative (CD24N) vs CD24-low (CD24L) MDA-MB-231 cells to 60 epigenetic compounds was measured. The upward sloping curve marked with blue stars indicate 12 of drugs that preferentially inhibit the proliferation CD44high/CD24low+ MDA-MB-231 CSCs. including Trichostatin (a1), Apicidin (a6), Scriptaid (a8), Vorinistat (a12), M-344 (b8), Fluoro-SAHA (b12), Oxamflatin (c10), BML-281 (d6), Rocilinostat (e5), CUDC-907 (e6), CUDC-101 (e7). Dimethyl sulfoxide controls are shown in the last two rows. The red star identifies one compound (GSK126) that selectively killed CD24neg cells.
Re-identification of HDAC inhibitors in a small molecule CSC lethality screen
In order to confirm the HDAC sensitivity of CSC-like BLERs in other CSC models, we used FACs-isolated CD44high/CD24low+ MDA-MB-231 CSCs. This subpopulation of cells have been previously shown to possess greater sphere-forming potential, increased capacity for tumour-initiation and metastasis in xenograft models, and higher expression of stem-like and metastatic mRNA signatures compared to CD44high/CD24neg cells.17, 32 Consistent with the BPLER results, we found that the CD44high/CD24low+ CSCs express significantly higher HDAC1, HDAC7 and CD166 proteins compared to their CD44high/CD24neg counterparts (Figure 4c). Also reminiscent of the BPLER results, this difference was only present at the protein level, with no significant difference in HDAC mRNA level (Figure 4d).
Having confirmed the association between CSCs and HDAC1/7 in this second cell line model, we used the CD44high/CD24low+ CSCs to screen a library of 60 epigenetic-targeting-compounds. This analysis identified 12 compounds that preferentially inhibit CD44high/CD24low+ MDA-MB-231 CSC population in cell viability assays. Interestingly, 11 of the 12 positive hits turned out to be HDACis, including TSA and Vorinostat (Figure 4e). Also, consistent with BPLER/HMLER, GSK126, which specifically inhibited the less tumorigenic, CD44high/CD24neg MDA-MB-231 population in this assay also preferentially inhibits HMLERs (data not shown). In conclusion, these results establish an association between high HDAC1/7 protein, HDAC activity and CSCs in an independent cell line model.
HDAC1 and HDAC7 are necessary to maintain the CSC phenotype
Given the high levels of HDAC1 and HDAC7 expression in CSC subpopulations, we sought to determine whether these HDACs were necessary to maintain the CSC phenotype. Using shRNA or CMV-driven constructs, we knocked down or overexpressed each HDAC in multiple breast and ovarian cell lines and examined the effects on the CSC phenotype. Experimental lines were selected based on endogenous HDAC1/7 expression levels (Supplementary Figures 13 and 14).
Consistent with a role for HDACs in CSC regulation, HDAC1 depletion decreases CSC marker expression, and inhibits sphere formation (3D) and 2D cell proliferation (2D) (Figures 5a and b). These results are confirmed with additional shRNA constructs in multiple cell lines (Supplementary Figure 13). Furthermore, HDAC1-shRNA expression results in significant reduction in xenograft tumour size and ~4-fold reduction in tumour-initiating cell (TIC) frequency compared to control cells expressing a scramble-shRNA construct (Figures 5c and d).
Knockdown of HDAC1 or HDAC7 alters the CSC phenotype in breast and ovarian cancer cell lines. (a) The knockdown of HDAC1 with shRNA reduces HDAC1, CD44 and CD166 protein levels in breast (MDA-MB-468/MCF7) and ovarian (SKOV3) cancer cell lines compared to a scramble shRNA control. Western blot of whole-cell lysates. β-Actin represents loading control. (b) HDAC1 knock-down with shRNA decreases 3D sphere formation (dark blue bars) more significantly than 2D proliferation (light blue bars) as compared a scramble shRNA-expressing control (white bars) in breast (MDA-MB-468, and MCF7) and ovarian (SKOV3) carcinoma cell lines. The data are presented as a percentage of the scramble shRNA controls. The error bars represent standard deviation of the mean from three replicates (*P<0.05)(**P<0.005). The results are representative of at least three independent experiments. Similar results were observed with additional HDAC1 shRNA constructs and in additional cell lines (Supplementary Figures 5b–d). (c) HDAC1 knockdown decreases tumour size in SUM159 breast cancer xenografts expressing two different stable HDAC1-shRNAs (shH1#1 and shH1#2) as compared to control scramble shRNA (shCntrl). Mean xenograft size measured from mice injected with 10 000 cells and plotted over time weeks) (P<0.05). (d) HDAC1 knockdown decreases tumour frequency in MDA-MB-231 xenografts expressing stable HDAC1-shRNA as compared to a scramble shRNA control line. TIC frequency calculated by limiting dilution analysis using http://bioinf.wehi.edu.au/software/elda/. TIC frequency of control cells (1.05 × 10−5), HDAC1 shRNA#1 (4.42 × 10−5) (P<0.005) and HDAC1 shRNA#2 (4.62 × 10−5) (P<0.005). (e) The knockdown of HDAC7 with shRNA reduces HDAC7, CD44 and CD166 protein levels compared to a scramble shRNA control in breast (MDA-MB-468, and MCF7) and ovarian (SKOV3) carcinoma cell line within 72 h western blot of whole-cell lysates. β-Actin represents loading control. (f) HDAC7 knockdown with shRNA decreases 3D sphere formation (dark green bars) more significantly than 2D proliferation (light green bars) as compared to a scramble shRNA-expressing control (white bars) in breast (MDA-MB-468, and MCF7) and ovarian (SKOV3) carcinoma cell lines. The results from one shRNA construct is shown, similar results were observed with additional HDAC7 shRNA constructs and in additional cell lines (Supplementary Figure 5). The cells are counted after trypan blue (2D) or INT staining (3D), and the results are presented as a percentage of the scramble shRNA control. The error bars represent standard deviation of the mean from three replicates (*P<0.05) (**P<0.01). The results are representative of at least three independent experiments.
Short-term HDAC7 depletion also decreases CSC marker expression, and inhibits 3D proliferation to a significantly greater extent than 2D in multiple cell lines, confirmed with several shRNA constructs (Figures 5e and f, Supplementary Figure 14). The specific inhibition of sphere formation, with lesser effects on 2D cell growth, suggests that reduction of sphere formation by CSCs with HDAC7 depletion is not simply due to a non-specific effect on regular cell proliferation. We found that HDAC7 protein is upregulated within several passages in HDAC7-shRNA cells with all three HDAC7-shRNA constructs (Supplementary Figures 15a and b). Therefore, long-term in vivo tumorigenesis analysis with stable cell lines expressing HDAC7-shRNA was not possible. The short-term shRNA results were also confirmed using transient siRNA transfection (Supplementary Figures 15c and d).
HDAC7 is sufficient to augment the CSC phenotype
These knockdown experiments indicate that HDAC1 and HDAC7 are necessary for the maintenance of the CSC phenotype. We next examined whether HDAC1 or HDAC7 overexpression are also sufficient to augment the CSC phenotype. We found that HDAC7 overexpression upregulated CSC markers, increased sphere formation two- to sixfold, and significantly enhanced sphere size, without any effect on 2D proliferation, as compared to control cells expressing the empty vector (Figures 6a–c, Supplementary Figure 16). Furthermore, HDAC7 overexpression in MCF7 cells upregulates 334 pro-metastatic or CSC-associated genes, alters gene expression of CSC-associated metabolic pathways,33 and downregulates expression of established HDAC7 targets34 and microRNAs associated with CSC phenotype (Supplementary Data 1, Supplementary Tables 3–5). Finally, in vivo limiting dilution analysis demonstrates that HDAC7 over-expression increases TIC frequency approximately twofold (Figure 6d). These findings indicate that HDAC7 is necessary and sufficient for augmenting the CSC phenotype in these breast cancer cell lines.
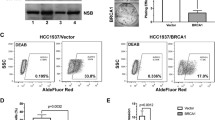
Overexpression of HDAC7 alters the CSC phenotype in breast and ovarian cancer cell lines. (a) HDAC7 over-expression (H7) increases CD44 and CD166 protein expression in MCF7 and SUM159, and CD44v(*) in SUM159 and HCC1937, compared to control cells expressing empty vector (EV). Western blot of whole-cell lysates. β-Actin represents loading control. (b) HDAC7 overexpression increases 3D sphere formation (dark green bars) with minimal effect on 2D proliferation (light green bars), as compared to an EV-expressing control (white bars), in breast (MCF7/HCC1937) and ovarian (CaOV3) cell lines. 2D growth assays were counted after trypan blue staining to assess viable cells counts. 3D sphere assays were counted after INT staining. The data is presented as a percentage of the EV-expressing control. The error bars represent standard deviation of the mean from triplicates (*P<0.05) (**P<0.01). The results are representative of at least three independent experiments. See Supplementary Figure 16 for additional cell lines. (c) Image of a representative well corresponding to the counts of 3D spheres in panel b. These images illustrate that both the number and size of the spheres are increased in MCF7 and HCC1937 cells over-expressing HDAC7-GFP as compared to an EV-expressing control. (d) HDAC7 overexpression increases TIC frequency in MDA-MB-231 xenografts expressing HDAC7-CMV as compared to EV-control. TIC frequency calculated by limiting dilution analysis using http://bioinf.wehi.edu.au/software/elda/. TIC frequency of control cells (1.14 × 10−5) and HDAC7 over-expressing cells (H7-CMV) (5.56 × 10−4)(P<0.03).
Transcriptional self-repression of HDAC1 has been previously reported.35 Consistent with this, we were not able to overexpress HDAC1 in multiple cell lines due to downregulation of endogenous HDAC1 levels (Supplementary Figures 17a and b). Consequently, while we did observe a small increase in 2D proliferation (up to twofold) in some cell lines expressing the HDAC1-CMV construct, there was no increase in sphere formation (Supplementary Figure 17c).
Inhibition of HDACs with class-specific drugs inhibits CSC phenotype
Motivated by the potential clinical implications of these findings we explored the effect of isoform-specific HDACis on CSC proliferation. While pan-HDACis (SAHA and TSA) have been used to target CSCs,36, 37, 38, 39 these HDACis indiscriminately target all 11 HDAC family members, resulting in toxicity that limits their clinical use.40 Consequently, we hypothesized that HDAC1/7-specific HDACis might prove more selective in inhibiting CSCs compared to pan-HDAC inhibitors. Although drugs that inhibit single HDAC isoforms are not currently available, we were able to obtain class selective HDACi.
Of particular interest, we discovered that MS275 (Entinostat) and MGCD0103 (Mocetinostat), members of the benzamide class of HDACi known to specifically inhibit HDAC1/2/3 enzyme activity,41, 42 dramatically downregulated HDAC7 protein levels, with limited effects on the protein levels of other HDAC family members (Figure 7). Based on our knockdown experiments implicating both HDAC1 and HDAC7 as necessary for the maintenance of the CSC phenotype, we found that both drugs preferentially target CSCs, as a 4-day treatment killed >85 % of BPLER population (as compared to vehicle treated controls), while having no effect on HMLER proliferation (Figure 7a). This differential effect was apparent as early as 24 h post-treatment. Furthermore, 24-h pretreatment of SUM159 with MS275 decreases the number of CD44hi/CD166hi, CD44hi/CD133hi and CD44hi/CD326hi cells as seen by FACS analysis (Supplementary Figure 18).
Isoform-specific HDACis that inhibit HDAC1 and HDAC7 can be used to selectively target CSCs. (a) The HDAC class I specific drugs that also downregulate HDAC7 (MS275 and MGCD) significantly inhibit BPLER proliferation (blue bar), with minimal effect on HMLERs (red bar). The class II specific HDACi MC1568 and MC1575 also preferentially inhibit BPLER proliferation compared to matched HMLER lines. In contrast, the HDACi that do not target HDAC1 or HDAC7 (Droxinostat and Apicidin) preferentially inhibit HMLER. The cells were treated with MS275 (1 μM), MGCD0103 (1 μM), MC1568 (1 μM), MC1575 (1 μM), Apicidin (0.1 μM) and Droxinostat (5 μM). Similar results were observed with two additional matched BPLER/HMLER pairs, in at least three independent experiments. The viable cells were assessed with trypan blue staining and counted after four days of treatment. Inhibition in proliferation is represented as a percentage of vehicle treated control (white bar). The error bars represent the standard deviation of the mean of triplicate samples (P<0.005). (b) Isoform-specific HDACis that target HDAC1 and/or HDAC7 (MS-275 and MGCD0103) preferentially inhibit 3D sphere formation (black bars) compared to 2D proliferation (grey bars). The breast cancer (BPLER, SUM159, MCF7, ZR751) and ovarian cancer (SKOV3) cell lines were pre-treated for 24 h with MS-275 (1 μM), MGCD0103 (1 μM) or MC1568 (1 μM). The next day, 2D and 3D cultures were established in drug-free medium. In contrast, pre-treatment with Droxinostat (5 μM) has minimal effects on 2D or 3D cell proliferation. The number of viable cells for 2D growth was determined by trypan blue staining. The number of 3D spheres was determined by INT staining. Inhibition in proliferation is represented as a percentage of vehicle treated control (white bar). The error bars represent the standard deviation of the mean of triplicate samples (*P<0.05) (**P<0.01). (c) The level of HDAC7 protein expression is significantly downregulated by short-term (24 h) treatment with MS-275 or MGCD0103 at 1 μM (low) or 2 μM (high) doses, but not with MC1568 or Droxinostat at 1 μM (low) or 5 μM (high) doses in SUM159 cells. Similar results are observed with BPLER, MCF7 and MDA-MB-231 cells (data not shown). Western blots of whole-cell lysates. β-Actin (*) was used as a loading control. (d) MS-275 pretreatment preferentially inhibits 3D sphere formation (black bars) compared to 2D proliferation (grey bars) in a panel of primary human ovarian cancer cell lines, including; OCI-C5x (clear cell), OCI-P7a (papillary serous), OCI-P9a1 (papillary serous) and OCI-P5x (papillary serous). The lines were pre-treated for 24 h with 1 μM MS-275, and plated into 2D and 3D cultures the following day in drug-free medium. The number of viable cells for 2D growth was determined by trypan blue staining. The number of 3D spheres was determined by INT staining. Inhibition in proliferation is represented as a percentage of vehicle treated control (white bar). The error bars represent the standard deviation of the mean of triplicate samples (*P<0.05) (**P<0.01).
In order to rule out MS275 and MGCD0103 effects on HDAC2 and HDAC3, we treated BPLER with the HDAC2/3/8-selective Apicidin and HDAC3/6/8-selective Droxinostat,43, 44 and determined that neither drug downregulates HDAC7 protein levels (Figure 7c). As anticipated from our results, neither drug preferentially inhibits BPLER proliferation (Figure 7a). These results indicate that the selective effects of benzamides on BPLER appear to be due to the combined inhibition of HDAC1 and HDAC7, and not related to HDAC3 or HDAC8.
While single isoform HDAC7-specific inhibitors are not yet available, HDACi that enzymatically inhibit HDAC4/5/7/9 have been developed (MC1568 and MC1575). Although these drugs inhibit HDAC7 enzymatically, they do not alter HDAC7 protein levels (Figure 7c). Interestingly, these drugs preferentially reduce 2D BPLER proliferation, with no effect on HMLER lines (Figure 7a), but have no effect on sphere formation (Figure 7b and data not shown), suggesting combined pharmacological inhibition of HDAC1 and downregulation of HDAC7 protein is important for CSC-specificity.
To confirm these findings in other models we treated several standard breast and ovarian cancer cell lines with the HDACi panel using low-dose, short-term (24 h) pretreatments determined by dose-response studies to have minimal effects on subsequent 2D growth. Compellingly, these studies reiterate that, while pretreatment with MS275 or MGCD0103 significantly reduces or eliminates sphere formation, pretreatment with MC1568 or MC1575 has limited effects on sphere formation in most lines (Figure 7c and data not shown). Similarly, pretreatment with Droxinostat or Apicidin had minimal effects on sphere formation (Figure 7c and data not shown). In addition to standard cell lines, we repeated these experiments in a panel of primary human ovarian cancer cell lines recently established by our lab,45 and observed that MS275 pretreatment also inhibits sphere formation in all primary lines tested (Figure 7d). Consistent with our results with BPLER cells, these findings suggest that MS275 or MGCD0103 inhibition of sphere-formation is not due to a generic inhibition of cell proliferation, and support the concept that these HDACis specifically inhibit CSCs. Furthermore, we found that MS275 treatment of BPLER cells reduces HDAC7 chromatin binding at stem cell associated transcription factor genes (Supplementary Figure 19), alters Histone 3 (H3) and H3K27 acetylation of stem cell transcription factor genes and reduces expression of several stem cell factors such as myc (Supplementary Figures 20 and 21). Lastly, we found that treatment of a primary ovarian cell line (OCI-P5x), established from a human high-grade serous ovarian carcinoma, inhibited mouse xenograft tumour formation in vivo (Figures 8a–c), and reduced the tumour sphere formation capacity of the explanted cells from these xenograft tumours. (Figure 8d).
MS275 inhibits xenograft tumour growth. The OCI-P5X ovarian cancer cell line that express luciferase was treated either with vehicle (Ctrl) or MS275 at 0.5 μM for 24 h and injected into the right and left flanks of nu/nu Balb/C mice at serial dilutions (106, 105, 104 and 103 cells/site). (a) The tumour growth was monitored with IVIS imaging once a week by intraperitoneal injection of 150 mg/kg D-luciferin 10 min before scanning. The representative image shows tumour specific bioluminescence signal in mice at week 6 after OCI-P5X cell injection. The same scale bar was used between ctrl and MS275 groups at each dilution for comparison. (b) Quantification of bioluminescence intensities in MS275 and control (vehicle) groups 6 weeks after injection. Black dots: Ctrl group. Black circles: MS275. Black line in the middle of the cluster represents the average signal in each group. (c) The number of gross tumours that formed at each injection site in MS275 and control (vehicle) groups; 10 sites of injection (5 mice per group, 2 sites/mouse). (d) The number of tumours spheres that are formed by OCI-P5x explants that were treated with MS275 and control (vehicle). White bar gross tumours that formed at each injection site in MS275 (black bar) and control (white bar). The images show representative wells photographed with regular and fluorescent microscopy (scale bar=2000 μm).
Discussion
In summary, our results show that the paired isogenic BPLER (CSC) and HMLER (nsTC) cell line pairs can provide a suitable model system to uncover epigenetic differences between CSC and nsTC. From a mechanistic perspective, our results suggest that HDAC1 and HDAC7 may be important for maintaining both normal stem cells and the CSC phenotype. Interestingly, HDAC1 and HDAC7 have been implicated in the regulation of normal pluripotency-associated genes4, 5, 6, 7, 8, 46 and colony formation in normal embryonic stem cells.9, 10 In malignant cells the class IIA HDACs are upregulated in breast and colon cancers compared to other cancers, highly expressed and associated with poor prognosis in astrocytoma, medulloblastoma and in childhood acute lymphoblastic leukemia.47, 48, 49 HDAC1/7 are shown to be repressed by the CSC-suppressor MiR-34a,50 which correlate with prognosis in gliomas and osteosarcomas.51, 52 In addition, it was found that Vorinostat downregulates HDAC7 in bladder and prostate carcinomas.53
The class I HDACs are upregulated in gastric, colorectal, esophageal, prostate and pancreatic cancer, high HDAC1 expression is associated with poor prognosis in colorectal cancer, pancreatic cancer, hepatocellular carcinoma, and lung cancer.54, 55, 56
So far, the limited clinical studies have been completed with MS275 and MGCD0103 predominantly in leukemia and lymphoma patients.57 However, as the potential CSC-specific actions of these drugs were previously unknown, CSC biomarkers were not used in patient selection or for measuring patient response. As it appears that the regulation of HDAC1 and HDAC7 levels is predominantly occurring at the protein level, it would be feasible to use immunohistochemistry to stratify patients based on HDAC expression levels for future clinical trials. Furthermore, our results indicate that in addition to inhibiting HDAC1, MS275 and MGCD0103 significantly downregulate HDAC7 protein levels. Thus, it might be possible to take advantage of this observation in monitoring patient response by comparing tumour biopsies before and after treatment. The significant reduction of HDAC7 in post-treatment biopsies might retrospectively distinguish patients in whom effective doses were achieved. A similar strategy might be used to individualize optimum dosing for each patient prospectively.
In standard trial design with cytostatic and cytotoxic drugs, the reduction in tumour size has been one of the main metrics for monitoring patient response. However, as suggested previously, CSC targeting drugs may not cause a rapid reduction in tumour size.58 This is because CSCs are thought to comprise a small portion of the tumour bulk and the remaining nsTC can proliferate for some time even when CSCs are completely eliminated. Hence, the effects of CSC targeting drugs may take a long time to result in a clinically measurable reduction in tumour size. The lag in clinical response will depend on how many population doublings nsTC can go through without being replenished by CSCs, which may be patient and tumour type dependent. These considerations make the use of biomarkers such as HDAC1 and HDAC7 for patient selection and monitoring response particularly compelling as a supplement to tumour size measurements.
Our results also indicate that these drugs should be tested more extensively in solid tumours, perhaps in combination with conventional chemotherapeutics to target the nsTC comprising the tumour bulk. This strategy may address some of the considerations regarding the lag time in tumour response with CSC-targeting drugs outlined above. Furthermore, it is thought that CSCs might be particularly relevant in recurrence and metastasis.58 Hence, a reduction in long-term recurrence might be an important metric, in addition to tumour size, to assess the role of CSC targeting drugs. To our knowledge the CSC-specific strategies outlined above were not used for patient selection or assessing response in previous studies. This is expected because the CSC-specific actions of MS275 and MGCD0103 we describe here were previously not fully recognized. Hence, the response in ongoing and previous studies to HDAC inhibitors should be viewed with caution; it is possible that better results might be achieved with careful attention to CSC-associated activities of this class of drugs. We hope that this report will stimulate efforts for more innovative clinical trials and development of more selective HDAC1 and HDAC7 inhibitors that may be even more effective than those currently available.
Materials and methods
Cell culture
The BPE, BPLER and OCI cells and their culture media BMI-P, BMI-T and OCMI are available from the Live Tumor Culture Core (LTCC) LTCC@med.miami.edu, at Sylvester Comprehensive Cancer Center (SCCC), Miller School of Medicine, University of Miami http://sylvester.org/shared-resources/Live-Tumor-Culture-Core. The BPE and BPLER cells were cultured in the BMI-P and BMI-T medium, respectively. The primary ovarian cancer cells (OCI) were cultured in OCMI medium as previously described,45 while the primary breast cells (BCI) were cultured in BCMI (unpublished). The HMLER cells are cultured in MEGM medium (Lonza, Allendale, NJ, USA) as previously described.23 The standard cancer cell lines were cultured in ATCC recommended media, at 37 °C and 5 % CO2 on standard plates (Falcon, New York, NY, USA). All standard cancer cell lines were recently short tandem repeat profiled and tested for mycoplasma contamination prior to experimentation. The BPE, BPLER, OCI and BCI cells are cultured on Primaria plates (BD Biosciences, San Jose, CA, USA) as previously described.23, 45
Serial dilution in vivo (BPLER/HMLER)
Single-cell suspensions were prepared in a WIT:Matrigel (1:1) mixture and injected in 25 μl (orthotopic) or 100 μl (subcutaneous) volumes. Female athymic nude mice (Balb/c nu/nu, Taconic) were γ-irradiated (400 rad) 12 h prior to subcutaneous injections. Sample size was determined by degree of dilution as follows: nine mice/experimental group for 105 and 104 dilution conditions, 12 mice/experimental set for 103, 102 and 101 dilution conditions. Animals were randomly assigned to each treatment group. Injections of tumorigenic cells into mammary fat pads were performed in 8-week-old female Nod/Scid mice that were anesthetized with intraperitoneal Avertin. Investigators were not blinded in the animal studies. All animal studies were approved and performed in accordance with relevant institutional IACUC committee guidelines.
Mammosphere formation
All mammospheres assays were carried out in BMI-T medium supplemented with 2 % B27 (Invitrogen, Carlsbad, CA, USA), 20 ng/ml EGF, 20 ng/ml bFGF (BD Biosciences), 4 μg/ml heparin (Sigma, St Louis, MO, USA) and 0.5% methyl cellulose. Six-well culture plates are coated with poly hema (20 mg/ml), and 20 000 cells are plated in 4 ml per well. After 2 weeks, the colonies are stained with 2-(4-iodophenyl)-3-(4-nitrophenyl)-5-phenyl-2H-tetrazolium chloride (INT), and counted on a Gelcount instrument (Oxford Optronix, Abingdon, UK). All experiments were carried out with triplicates and repeated a minimum of three times. The error bars represent the standard error of the mean of the three replicates.
Flow cytometry and cell sorting
Cells were harvested from sub-confluent culture plates and cell pellets were washed, resuspended in HBSS buffer containing 0.2 % BSA, and stained with fluorophore-conjugated monoclonal antibodies per manufacturer’s instruction. Antibodies used for cell sorting or analysis included: anti-CD44, APC conjugated (BD Pharmingen, San Jose, CA, USA; #559942); anti-CD44, FITC conjugated (Abcam, Cambridge, UK; #ab19622); CD133/1, PE conjugated (Miltenyi Biotech, Bergisch Gladbach, Germany; #130-080-801); CD166, PE conjugated (BD Pharmingen, #559263); CD326, FITC conjugated (AbD # MCA1870F); and CD326, PerCP-Cy5.5 conjugated (BD Pharmingen, #347199). Flow cytometry analysis was performed using an Accuri C6 cytometer (Accuri cytometers Inc., San Jose, CA, USA). Sorted cells were collected in heat-inactivated fetal serum or HBSS buffer containing 2 % BSA. Purity of sorted cells was between 90 and 95 %.
Western blotting
Whole-cell lysates, harvested with RIPA buffer containing protease and phosphatase inhibitors, were run on 4–15 % SDS-PAGE gradient gels (Biorad, Hercules, CA, USA). Blots were blocked in 5 % milk, then probed with antibodies; CD326 (Cell Signaling, Danvers, MA, USA; #2929 S), CD166 (Abcam#ab49496), CD44(2c5) (R&D#BBA10), BMI-1 (Cell Signaling#2830 S), HDAC1 (Cell Signaling#5356), HDAC2 (Cell Signaling#5113), HDAC3 (Cell Signaling#3949), HDAC4 (Cell Signaling#7628), HDAC5 (Cell Signaling#2082), HDAC6 (Novus, Littleton, CO, USA; #91805), HDAC7 (Abcam#ab12174), HDAC8 (Novus#91807), HDAC9 (Abcam#ab18970), HDAC10 (Novus#91801). Blots were developed using SuperSignal West Pico or Femto ECL (ThermoScientific, Waltham, MA, USA). All experiments were repeated a minimum of three times.
Inhibition of 2D cell proliferation with continuous drug treatment
To assess the effect of long-term drug treatment on 2D proliferation of BPLER and HMLER cells, they were plated at a density of 250 000 cells/well in six-well Primaria plates. After 24 h, drug-containing media was added to each well. Cells were treated continuously for 4 days, at which point cell counts and viability (based on Trypan blue exclusion) were quantified using a Nexelcom Cellometer, and assessed as a percentage of control cells. All treatments were done in triplicates, and repeated a minimum of three times.
Inhibition of 2D/3D cell proliferation with drug pretreatment
To assess effect of short-term drug pretreatment on 2D/3D growth, cells are plated in six-well plates and allowed to recover for 24 h. At this point, drug-containing media was added to each well. After 24 h of treatment, cells were then were plated for mammospheres or 2D proliferation assays. 2D proliferation was determined after 3 days by viable cell counts, and assessed as a percentage of the dimethyl sulfoxide control. All treatments were carried out in triplicate, and repeated a minimum of three times. The error bars represent the standard error of the mean of the three replicates. The small molecule drugs that are used to treat the cells for various experiments are: TSA (Selleck, Boston, MA, USA) Doxirubicin (Selleck), Paclitaxel (Selleck), 5-Fluorouracil (Sigma), SAHA (Selleck), GSK126 (Cayman Chem), MS275 (Selleck), MGCD0103 (Selleck), MC1568 (Selleck), MC1575 (Gift of Dr. Antonello Mai), Apicidin (Sigma), Droxinostat (Selleck). All drugs are prepared in dimethyl sulfoxide, which was used as vehicle control in each assay.
mRNA expression analysis
mRNA analysis as previously described23 using Affymatrix human 133 2.0.
Apoptosis
The cells lines were treated with drugs for 24 h at the following doses: 5-FU (50 μM), Taxol (50 nM), Doxirubicin (5 μM). Cells were then assayed for apoptosis with Annexin V-PE apoptosis kit (Abcam#ab14155) as per the manufacturer’s protocol. All experiments were carried out in triplicates, and repeated a minimum of three times.
Immunofluorescence
The cells were plated on to eight-well chamber slides, fixed with 4 % paraformaldehyde for 15 min, washed three times with PBS/100 mM Glycine, and then permeabilized for 15 min with 0.5 % Tween-20. Slides were then blocked for 1 h in 10 % Goat serum/0.1 % BSA/PBS, incubated for 1.5 h with primary antibody, followed by 45 min incubation with fluorophore-conjugated secondary antibody. Slides were mounted with Fluoro-Gel II, with DAPI (EM Sciences, Hatfield, PA, USA) for nuclear staining. Cells were imaged on an Evos fluorescence microscope (AMG & Life Tech, Millcreek, WA, USA). Antibodies: HDAC7 (Abcam#ab12174), CD44 (R&D#BBA10), CD166 (AbD Serotech, Raleigh, NC, USA; #MCA1926T). All experiments were repeated a minimum of three times.
HDAC activity assay
The cells were harvested and sorted by flow cytometry as described above. After three washes with cold HBSS buffer cells were lysed in cold lysis buffer and centrifuged at 4 °C. The protein concentration of the cell lysates were quantified by Coomassie Plus Reagent (Thermoscientific #1856210). HDAC activity assay (Cayman Chemical Co., Ann Artbor, MI, USA; #10011563) was performed according to the manufacture’s protocol. The data are presented as fluorescence units. All experiments were carried out in triplicates in each of at least three independent experiments.
shRNA knockdown of HDAC1 and HDAC7
The cell lines were transduced with GIPZ lentiviral shRNAmir constructs purchased from Open Biosystems (Lafayette, CO, USA)/ThermoScientific (Catalog # V3LHS_351665, V2LHS_96400, V2LHS_96401). HDAC1 cell lines were transduced with MISSION shRNA purchased from Sigma (Catalog# TRCN0000195467, TRCN0000195672, TRCN0000195103, TRCN0000195672). The cell lines that stably express the shRNAs were selected for puromycin resistance. All experiments were performed in triplicate, and repeated a minimum of three times.
siRNA knockdown of HDAC7
HDAC7 siRNA (Sigma#EHU078781) was transfected into cell lines with Mission Transfection Reagent (Sigma#S1452) as per manufacturer’s protocol. Transfected cells plated in triplicate in 2D/3D assays and lysates harvested at 72 h post-transfection. All experiments were repeated a minimum of three times.
Over-expression of HDAC1 and HDAC7
HDAC1-Flag (provided by Eric Verdin through Addgene, Cambridge, MA, USA; plasmid#13820) and HDAC7 (purchased from Origene, Rockville, MD, USA; cat. #RC215233) were cloned into pENTR vectors. These constructs were recombined into pLenti CMV DEST lentiviral vectors (provided by Eric Campeau through Addgene). Lentivirus was packaged in 293T cells using the manufacturer’s recommended protocols. Stable lines were selected with G418 (HDAC7) or Blast (HDAC1). All experiments were carried out in triplicate, and repeated a minimum of three times.
Serial dilution in vivo (knockdown/overexpression)
MDA-MB-231-luciferase cells were transduced with lentivirus and selected for stable expression of HDAC7-CMV, empty vector control (EV), HDAC1 shRNA (Construct #1 and Contruct #2), or a scramble shRNA control. HDAC7-overexpression, or HDAC1 knockdown was confirmed by western blot. Serial dilutions of MDA-MB-231-luciferase cells were injected (1 000 000, 100 000, 10 000, 1000, or 100 cells suspended in 100 μl of HBSS:Matrigel) into the mammary fat pad of 5-week-old, female, Balb/C nude mice (Charles River, Wilmington, MA, USA) at the rate of three injections per mouse. The mice were randomly assigned to each treatment group. Tumours were followed by weekly IVIS analysis beginning at 5 weeks. The mice that died post-injection and prior to IVIS imaging were eliminated from analysis. The tumours were also visually scored (+/−) and each tumour was measured at its maximum diameter. Tumours from 1 000 000 and 100 000 dilution groups were excised at 1 cm diameter and animals were examined for metastasis. All others tumours were excised per Animal Care and Use Committee or at 100 days if no tumour arose. Tumour luciferase signals were analysed by Living Image In Vivo imaging software (Perkin Elmer, Waltham, MA, USA) and plotted using GraphPad Prism 5.0. Investigators were not blinded in the animal studies. All animal studies were approved and performed in accordance with relevant institutional IACUC committee guidelines.
Serial dilution in vivo (MS-275 treatment)
The primary ovarian cancer cell line OCI-P5X that expresses luciferase-GFP (OCI-P5X Fluc-IRES-GFP) was plated on T-75 flasks and treated with either 0.5 μM MS275 or vehicle control for 24 h. After trypsinization, serial dilutions were injected (1 000 000, 100 000, 10 000, or 1000 cells suspended in 100 μl of 1:1 HBSS: Matrigel) into the left and right flanks of 5-week-old Balb/C nude mice (Charles River). Animals were randomly assigned to each treatment group. Tumours were followed by weekly IVIS analysis beginning at 1 week. Tumour luciferase signals were analysed by Living Image In Vivo imaging software (Perkin Elmer) and plotted using GraphPad Prism 5.0. Investigators were not blinded in the animal studies. All animal studies were approved and performed in accordance with relevant institutional IACUC committee guidelines.
Statistical analysis
Cell proliferation, sphere formation and HDAC activities were compared by two-tailed Student’s t-test, using GraphPad Prism 5.0, GraphPad Software with P<0.05 considered significant. Limiting dilution analysis was done with http://bioinf.wehi.edu.au/software/elda/index.html.59
References
Ablett MP, Singh JK, Clarke RB . Stem cells in breast tumours: are they ready for the clinic? Eur J Cancer [Research Support, Non-US Gov’t Review] 2012; 48: 2104–2116.
Vincent A, Van Seuningen I . On the epigenetic origin of cancer stem cells. Biochimica et biophysica acta 2012; 1826: 83–88.
Seto E, Yoshida M . Erasers of histone acetylation: the histone deacetylase enzymes. Cold Spring Harb Perspect Biol [Review] 2014; 6: a018713.
Kidder BL, Palmer S . HDAC1 regulates pluripotency and lineage specific transcriptional networks in embryonic and trophoblast stem cells. Nucleic Acids Res 2012; 40: 2925–2939.
Liang J, Wan M, Zhang Y, Gu P, Xin H, Jung SY et al. Nanog and Oct4 associate with unique transcriptional repression complexes in embryonic stem cells. Nat Cell Biol 2008; 10: 731–739.
Dovey OM, Foster CT, Cowley SM . Histone deacetylase 1 (HDAC1), but not HDAC2, controls embryonic stem cell differentiation. Proc Natl Acad Sci USA 2010; 107: 8242–8247.
McCool KW, Xu X, Singer DB, Murdoch FE, Fritsch MK . The role of histone acetylation in regulating early gene expression patterns during early embryonic stem cell differentiation. J Biol Chem 2007; 282: 6696–6706.
Karantzali E, Schulz H, Hummel O, Hubner N, Hatzopoulos A, Kretsovali A . Histone deacetylase inhibition accelerates the early events of stem cell differentiation: transcriptomic and epigenetic analysis. Genome Biol 2008; 9: R65.
Zhu C, Chen Q, Xie Z, Ai J, Tong L, Ding J et al. The role of histone deacetylase 7 (HDAC7) in cancer cell proliferation: regulation on c-Myc. J Mol Med (Berl) 2011; 89: 279–289.
You JS, Kang JK, Seo DW, Park JH, Park JW, Lee JC et al. Depletion of embryonic stem cell signature by histone deacetylase inhibitor in NCCIT cells: involvement of Nanog suppression. Cancer Res 2009; 69: 5716–5725.
Jiao J, Hindoyan A, Wang S, Tran LM, Goldstein AS, Lawson D et al. Identification of CD166 as a surface marker for enriching prostate stem/progenitor and cancer initiating cells. PloS One [Research Support, N.I.H., Extramural Research Support, Non-US Gov’t Research Support, US Gov’t, Non-P.H.S.] 2012; 7: e42564.
Dalerba P, Dylla SJ, Park IK, Liu R, Wang X, Cho RW et al. Phenotypic characterization of human colorectal cancer stem cells. Proc Natl Acad Sci USA 2007; 104: 10158–10163.
Rowehl RA, Burke S, Bialkowska AB, Pettet DW 3rd, Rowehl L, Li E et al. Establishment of highly tumorigenic human colorectal cancer cell line (CR4) with properties of putative cancer stem cells. PloS One [Research Support, Non-US Gov’t] 2014; 9: e99091.
Fillmore CM, Kuperwasser C . Human breast cancer cell lines contain stem-like cells that self-renew, give rise to phenotypically diverse progeny and survive chemotherapy. Breast Cancer Res: BCR. [Research Support, N.I.H., Extramural Research Support, Non-US Gov’t] 2008; 10: R25.
Stuelten CH, Mertins SD, Busch JI, Gowens M, Scudiero DA, Burkett MW et al. Complex display of putative tumor stem cell markers in the NCI60 tumor cell line panel. Stem Cells [Research Support, N.I.H., Extramural Research Support, N.I.H., Intramural] 2010; 28: 649–660.
Kok M, Koornstra RH, Margarido TC, Fles R, Armstrong NJ, Linn SC et al. Mammosphere-derived gene set predicts outcome in patients with ER-positive breast cancer. J Pathol [Research Support, Non-US Gov’t] 2009; 218: 316–326.
Azzam DJ, Zhao D, Sun J, Minn AJ, Ranganathan P, Drews-Elger K et al. Triple negative breast cancer initiating cell subsets differ in functional and molecular characteristics and in gamma-secretase inhibitor drug responses. EMBO Mol Med [Research Support, N.I.H., Extramural Research Support, Non-US Gov’t Research Support, US Gov’t, Non-P.H.S.] 2013; 5: 1502–1522.
Civenni G, Walter A, Kobert N, Mihic-Probst D, Zipser M, Belloni B et al. Human CD271-positive melanoma stem cells associated with metastasis establish tumor heterogeneity and long-term growth. Cancer Res [Research Support, Non-US Gov’t] 2011; 71: 3098–3109.
Munz M, Baeuerle PA, Gires O . The emerging role of EpCAM in cancer and stem cell signaling. Cancer Res [Research Support, Non-US Gov’t Review] 2009; 69: 5627–5629.
Yamashita T, Ji J, Budhu A, Forgues M, Yang W, Wang HY et al. EpCAM-positive hepatocellular carcinoma cells are tumor-initiating cells with stem/progenitor cell features. Gastroenterology [Research Support, N.I.H., Extramural Research Support, Non-US Gov’t Research Support, US Gov’t, Non-P.H.S.] 2009; 136: 1012–1024.
Williams K, Motiani K, Giridhar PV, Kasper S . CD44 integrates signaling in normal stem cell, cancer stem cell and (pre)metastatic niches. Exp Biol Med (Maywood) [Research Support, N.I.H., Extramural Research Support, Non-US Gov’t Review] 2013; 238: 324–338.
Belsky DW, Moffitt TE, Baker TB, Biddle AK, Evans JP, Harrington H et al. Polygenic risk and the developmental progression to heavy, persistent smoking and nicotine dependence: evidence from a 4-decade longitudinal study. JAMA Psychiatry [Research Support, N.I.H., Extramural Research Support, Non-US Gov’t Research Support, US Gov’t, P.H.S.] 2013; 70: 534–542.
Ince TA, Richardson AL, Bell GW, Saitoh M, Godar S, Karnoub AE et al. Transformation of different human breast epithelial cell types leads to distinct tumor phenotypes. Cancer Cell 2007; 12: 160–170.
Sauder CA, Koziel JE, Choi M, Fox MJ, Grimes BR, Badve S et al. Phenotypic plasticity in normal breast derived epithelial cells. BMC Cell Biol 2014; 15: 20.
Singh SK, Hawkins C, Clarke ID, Squire JA, Bayani J, Hide T et al. Identification of human brain tumour initiating cells. Nature [Research Support, Non-US Gov’t] 2004; 432: 396–401.
Ponti D, Costa A, Zaffaroni N, Pratesi G, Petrangolini G, Coradini D et al. Isolation and in vitro propagation of tumorigenic breast cancer cells with stem/progenitor cell properties. Cancer Res [Research Support, Non-US Gov’t] 2005; 65: 5506–5511.
Ricci-Vitiani L, Lombardi DG, Pilozzi E, Biffoni M, Todaro M, Peschle C et al. Identification and expansion of human colon-cancer-initiating cells. Nature [Research Support, Non-US Gov’t] 2007; 445: 111–115.
Eramo A, Lotti F, Sette G, Pilozzi E, Biffoni M, Di Virgilio A et al. Identification and expansion of the tumorigenic lung cancer stem cell population. Cell Death Differ [Research Support, Non-US Gov’t] 2008; 15: 504–514.
Han ME, Jeon TY, Hwang SH, Lee YS, Kim HJ, Shim HE et al. Cancer spheres from gastric cancer patients provide an ideal model system for cancer stem cell research. Cell Mol Life Sci: CMLS. [Research Support, Non-US Gov’t] 2011; 68: 3589–3605.
Merritt MA, Bentink S, Schwede M, Iwanicki MP, Quackenbush J, Woo T et al. Gene expression signature of normal cell-of-origin predicts ovarian tumor outcomes. PLoS One 2013; 8: e80314.
Petrocca F, Altschuler G, Tan SM, Mendillo ML, Yan H, Jerry DJ et al. A genome-wide siRNA screen identifies proteasome addiction as a vulnerability of basal-like triple-negative breast cancer cells. Cancer Cell 2013; 24: 182–196.
Leth-Larsen R, Terp MG, Christensen AG, Elias D, Kuhlwein T, Jensen ON et al. Functional heterogeneity within the CD44 high human breast cancer stem cell-like compartment reveals a gene signature predictive of distant metastasis. Mol Med [Research Support, Non-US Gov’t] 2012; 18: 1109–1121.
Pasto A, Bellio C, Pilotto G, Ciminale V, Silic-Benussi M, Guzzo G et al. Cancer stem cells from epithelial ovarian cancer patients privilege oxidative phosphorylation, and resist glucose deprivation. Oncotarget [Research Support, Non-US Gov’t] 2014; 5: 4305–4319.
Kasler HG, Young BD, Mottet D, Lim HW, Collins AM, Olson EN et al. Histone deacetylase 7 regulates cell survival and TCR signaling in CD4/CD8 double-positive thymocytes. J Immunol [Research Support, N.I.H., Extramural] 2011; 186: 4782–4793.
Schuettengruber B, Simboeck E, Khier H, Seiser C . Autoregulation of mouse histone deacetylase 1 expression. Mol Cell Biol [Research Support, Non-US Gov’t] 2003; 23: 6993–7004.
Chikamatsu K, Ishii H, Murata T, Sakakura K, Shino M, Toyoda M et al. Alteration of cancer stem cell-like phenotype by histone deacetylase inhibitors in squamous cell carcinoma of the head and neck. Cancer Sci [Research Support, Non-US Gov’t] 2013; 104: 1468–1475.
Salvador MA, Wicinski J, Cabaud O, Toiron Y, Finetti P, Josselin E et al. The histone deacetylase inhibitor abexinostat induces cancer stem cells differentiation in breast cancer with low Xist expression. Clin Cancer Res: An Official Journal of the American Association for Cancer Research [Research Support, Non-US Gov’t] 2013; 19: 6520–6531.
Debeb BG, Lacerda L, Xu W, Larson R, Solley T, Atkinson R et al. Histone deacetylase inhibitors stimulate dedifferentiation of human breast cancer cells through WNT/beta-catenin signaling. Stem Cells [Research Support, N.I.H., Extramural Research Support, Non-US Gov’t] 2012; 30: 2366–2377.
Munster PN, Troso-Sandoval T, Rosen N, Rifkind R, Marks PA, Richon VM . The histone deacetylase inhibitor suberoylanilide hydroxamic acid induces differentiation of human breast cancer cells. Cancer Res [Research Support, US Gov’t, P.H.S.] 2001; 61: 8492–8497.
Galanis E, Jaeckle KA, Maurer MJ, Reid JM, Ames MM, Hardwick JS et al. Phase II trial of vorinostat in recurrent glioblastoma multiforme: a north central cancer treatment group study. J Clin Oncol: Official Journal of the American Society of Clinical Oncology [Clinical Trial, Phase II Research Support, N.I.H., Extramural Research Support, Non-US Gov’t] 2009; 27: 2052–2058.
Beckers T, Burkhardt C, Wieland H, Gimmnich P, Ciossek T, Maier T et al. Distinct pharmacological properties of second generation HDAC inhibitors with the benzamide or hydroxamate head group. Int J Cancer [Journal international du cancer] 2007; 121: 1138–1148.
Boissinot M, Inman M, Hempshall A, James SR, Gill JH, Selby P et al. Induction of differentiation and apoptosis in leukaemic cell lines by the novel benzamide family histone deacetylase 2 and 3 inhibitor MI-192. Leukemia Res [Research Support, Non-US Gov’t] 2012; 36: 1304–1310.
Khan N, Jeffers M, Kumar S, Hackett C, Boldog F, Khramtsov N et al. Determination of the class and isoform selectivity of small-molecule histone deacetylase inhibitors. Biochem J 2008; 409: 581–589.
Wood TE, Dalili S, Simpson CD, Sukhai MA, Hurren R, Anyiwe K et al. Selective inhibition of histone deacetylases sensitizes malignant cells to death receptor ligands. Mol Cancer Ther [Research Support, Non-US Gov’t] 2010; 9: 246–256.
Ince TA, Sousa AD, Jones MA, Harrell JC, Agoston ES, Krohn M et al. Characterization of twenty-five ovarian tumour cell lines that phenocopy primary tumours. Nat Commun 2015; 6: 7419.
Zhang L, Jin M, Margariti A, Wang G, Luo Z, Zampetaki A et al. Sp1-dependent activation of HDAC7 is required for platelet-derived growth factor-BB-induced smooth muscle cell differentiation from stem cells. J Biol Chem 2010; 285: 38463–38472.
Clocchiatti A, Di Giorgio E, Ingrao S, Meyer-Almes FJ, Tripodo C, Brancolini C . Class IIa HDACs repressive activities on MEF2-depedent transcription are associated with poor prognosis of ER(+) breast tumors. FASEB Journal: Official publication of the Federation of American Societies for Experimental Biology [Research Support, Non-US Gov’t] 2013; 27: 942–954.
Milde T, Oehme I, Korshunov A, Kopp-Schneider A, Remke M, Northcott P et al. HDAC5 and HDAC9 in medulloblastoma: novel markers for risk stratification and role in tumor cell growth. Clini Cancer Res: An Official Journal of the American Association for Cancer Research [Research Support, Non-US Gov’t] 2010; 16: 3240–3252.
Van Damme M, Crompot E, Meuleman N, Mineur P, Bron D, Lagneaux L et al. HDAC isoenzyme expression is deregulated in chronic lymphocytic leukemia B-cells and has a complex prognostic significance. Epigenetics: Official Journal of the DNA Methylation Society [Research Support, Non-US Gov’t] 2012; 7: 1403–1412.
Wu MY, Fu J, Xiao X, Wu J, Wu RC . MiR-34a regulates therapy resistance by targeting HDAC1 and HDAC7 in breast cancer. Cancer Lett 2014; 354: 311–319.
Wang Y, Jia LS, Yuan W, Wu Z, Wang HB, Xu T et al. Low miR-34a and miR-192 are associated with unfavorable prognosis in patients suffering from osteosarcoma. Am J Transl Res 2015; 7: 111–119.
Gao H, Zhao H, Xiang W . Expression level of human miR-34a correlates with glioma grade and prognosis. J Neurooncol 2013; 113: 221–228.
Dokmanovic M, Perez G, Xu W, Ngo L, Clarke C, Parmigiani RB et al. Histone deacetylase inhibitors selectively suppress expression of HDAC7. Mol Cancer Ther 2007; 6: 2525–2534.
Higashijima J, Kurita N, Miyatani T, Yoshikawa K, Morimoto S, Nishioka M et al. Expression of histone deacetylase 1 and metastasis-associated protein 1 as prognostic factors in colon cancer. Oncol Rep [Research Support, Non-US Gov’t] 2011; 26: 343–348.
Miyake K, Yoshizumi T, Imura S, Sugimoto K, Batmunkh E, Kanemura H et al. Expression of hypoxia-inducible factor-1alpha, histone deacetylase 1, and metastasis-associated protein 1 in pancreatic carcinoma: correlation with poor prognosis with possible regulation. Pancreas 2008; 36: e1–e9.
Minamiya Y, Ono T, Saito H, Takahashi N, Ito M, Mitsui M et al. Expression of histone deacetylase 1 correlates with a poor prognosis in patients with adenocarcinoma of the lung. Lung Cancer 2011; 74: 300–304.
Garcia-Manero G, Assouline S, Cortes J, Estrov Z, Kantarjian H, Yang H et al. Phase 1 study of the oral isotype specific histone deacetylase inhibitor MGCD0103 in leukemia. Blood [Clinical Trial, Phase I Research Support, Non-US Gov’t] 2008; 112: 981–989.
Carrasco E, Alvarez PJ, Prados J, Melguizo C, Rama AR, Aranega A et al. Cancer stem cells and their implication in breast cancer. Eur J Clin Invest [Research Support, Non-US Gov’t Review] 2014; 44: 678–687.
Hu Y, Smyth GK . ELDA: extreme limiting dilution analysis for comparing depleted and enriched populations in stem cell and other assays. J Immunol Methods [Comparative Study] 2009; 347: 70–78.
Acknowledgements
This work was funded by grants from: (1) Sylvester Comprehensive Cancer Center, Miller School of Medicine, University of Miami (TAI); (2) Breast Cancer Research Foundation, New York, NY (TAI); (3) Women’s Cancer Association (TAI); (4) DoD-CDMRP Ovarian Cancer Research Program (W81XWH-14-1-0160) Resource Development Grant OC130649 (TAI), and (5) National Cancer Institute, Roadmap Epigenomics project (R01-CA146445-01) to TAI, TIL and RAY. We thank Jayanta Debnath for his help with 3D morphogenesis assays, Dr Antonello Mai for providing MC1575, Chuck Harrell for help with mRNA expression array analysis, P.L. Guevara for help with FACs analysis, Noriyuki Kasahara and Sara Collins for transduction of OCI-P5x cell line with luciferase and F. Reinhardt for technical assistance.
Author contributions
Conception, design and study supervision (Tan A. Ince, Abigail E. Witt), development of methodology and cell culture medium (Tan A. Ince), acquisition of data (Abigail E. Witt, Chung-Wei Lee, Tong Ihn Lee, Diana J. Azzam, Fabio Petrocca, James Grosso, Evan B. Cohick, Bin Wang, Corrado Caslini, Michelle Jones and Adrienne B. Gropper), bioinformatic, computational, statistical analysis and interpretation (Tan A. Ince, Richard A Young, Fabio Petrocca, Andrea L. Richardson, Ramin Shiekhattar), writing, review and revision of the manuscript (Tan A. Ince and Abigail E. Witt).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Tan A. Ince discloses pending patent intellectual proprietary interest as the inventor of BMI and OCMI media.
Additional information
Supplementary Information accompanies this paper on the Oncogene website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Witt, A., Lee, CW., Lee, T. et al. Identification of a cancer stem cell-specific function for the histone deacetylases, HDAC1 and HDAC7, in breast and ovarian cancer. Oncogene 36, 1707–1720 (2017). https://doi.org/10.1038/onc.2016.337
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2016.337
This article is cited by
-
The impact of selective HDAC inhibitors on the transcriptome of early mouse embryos
BMC Genomics (2024)
-
Complexation of histone deacetylase inhibitor belinostat to Cu(II) prevents premature metabolic inactivation in vitro and demonstrates potent anti-cancer activity in vitro and ex vivo in colon cancer
Cellular Oncology (2024)
-
Histone modifications in drug-resistant cancers: From a cancer stem cell and immune evasion perspective
Experimental & Molecular Medicine (2023)
-
HDAC7/c-Myc signaling pathway promotes the proliferation and metastasis of choroidal melanoma cells
Cell Death & Disease (2023)
-
Epigenetic and molecular coordination between HDAC2 and SMAD3-SKI regulates essential brain tumour stem cell characteristics
Nature Communications (2023)