Abstract
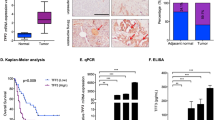
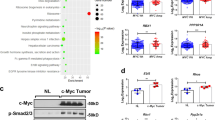
Hepatocyte growth factor (HGF)/Met signaling has critical roles in pancreatic ductal adenocarcinoma (PDA) development and progression and is considered a potential therapeutic target for this disease. However, the mechanism of aberrant activation of HGF/Met signaling and resistance to Met inhibition in PDA remains unclear. The mechanistic role of cross talk between Forkhead box M1 (FOXM1) and HGF/Met signaling in promotion of PDA growth and resistance to Met inhibition was examined using cell culture, molecular biology and mouse models; and the relevance of our experimental and mechanistic findings were validated using human PDA tissues. Met was markedly overexpressed in both PDA cell lines and pancreatic tumor specimens, and the expression of Met correlated directly with that of FOXM1 in human tumor specimens. Mechanistically, FOXM1 bound to the promoter region of the Met gene and transcriptionally increased the expression of Met. Increased expression of FOXM1 enhanced the activation of HGF/Met signaling and its downstream pathways, including retrovirus-associated DNA sequences/extracellular signal-regulated kinase 1/2, phosphoinositide 3-kinase/AKT and signal transducer and activator of transcription 3. Furthermore, activation of HGF/Met signaling increased the expression and transcriptional activity of FOXM1, and the cross talk between FOXM1 and HGF/Met signaling promoted PDA growth and resistance to Met inhibition. Collectively, our findings identified a positive feedback loop formed by FOXM1 and HGF/Met and revealed that this loop is a potentially effective therapeutic target for PDA.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Jemal A, Bray F, Center MM, Ferlay J, Ward E, Forman D . Global cancer statistics. CA Cancer J Clin 2011; 61: 69–90.
Siegel R, Ma J, Zou Z, Jemal A . Cancer statistics, 2014. CA Cancer J Clin 2014; 64: 9–29.
Hidalgo M . Pancreatic cancer. N Engl J Med 2010; 362: 1605–1617.
Birchmeier C, Gherardi E . Developmental roles of HGF/SF and its receptor, the c-Met tyrosine kinase. Trends Cell Biol 1998; 8: 404–410.
Zhang YW, Vande Woude GF . HGF/SF-met signaling in the control of branching morphogenesis and invasion. J Cell Biochem 2003; 88: 408–417.
Cui JJ . Targeting receptor tyrosine kinase MET in cancer: small molecule inhibitors and clinical progress. J Med Chem 2014; 57: 4427–4453.
De Bacco F, Luraghi P, Medico E, Reato G, Girolami F, Perera T et al. Induction of MET by ionizing radiation and its role in radioresistance and invasive growth of cancer. J Natl Cancer Inst 2011; 103: 645–661.
Neesse A, Michl P, Frese KK, Feig C, Cook N, Jacobetz MA et al. Stromal biology and therapy in pancreatic cancer. Gut 2011; 60: 861–868.
Logan-Collins J, Thomas RM, Yu P, Jaquish D, Mose E, French R et al. Silencing of RON receptor signaling promotes apoptosis and gemcitabine sensitivity in pancreatic cancers. Cancer Res 2010; 70: 1130–1140.
Zhu GH, Huang C, Qiu ZJ, Liu J, Zhang ZH, Zhao N et al. Expression and prognostic significance of CD151, c-Met, and integrin alpha3/alpha6 in pancreatic ductal adenocarcinoma. Dig Dis Sci 2011; 56: 1090–1098.
Kitajima Y, Ide T, Ohtsuka T, Miyazaki K . Induction of hepatocyte growth factor activator gene expression under hypoxia activates the hepatocyte growth factor/c-Met system via hypoxia inducible factor-1 in pancreatic cancer. Cancer Sci 2008; 99: 1341–1347.
Ketterer K, Kong B, Frank D, Giese NA, Bauer A, Hoheisel J et al. Neuromedin U is overexpressed in pancreatic cancer and increases invasiveness via the hepatocyte growth factor c-Met pathway. Cancer Lett 2009; 277: 72–81.
Otte JM, Kiehne K, Schmitz F, Folsch UR, Herzig KH . C-met protooncogene expression and its regulation by cytokines in the regenerating pancreas and in pancreatic cancer cells. Scand J Gastroenterol 2000; 35: 90–95.
Clark KL, Halay ED, Lai E, Burley SK . Co-crystal structure of the HNF-3/fork head DNA-recognition motif resembles histone H5. Nature 1993; 364: 412–420.
Huang C, Du J, Xie K . FOXM1 and its oncogenic signaling in pancreatic cancer pathogenesis. Biochim Biophys Acta 2014; 1845: 104–116.
Huang C, Qiu Z, Wang L, Peng Z, Jia Z, Logsdon CD et al. A novel FoxM1-caveolin signaling pathway promotes pancreatic cancer invasion and metastasis. Cancer Res 2012; 72: 655–665.
Bao B, Wang Z, Ali S, Kong D, Banerjee S, Ahmad A et al. Over-expression of FoxM1 leads to epithelial-mesenchymal transition and cancer stem cell phenotype in pancreatic cancer cells. J Cell Biochem 2011; 112: 2296–2306.
Cui J, Shi M, Xie D, Wei D, Jia Z, Zheng S et al. FOXM1 promotes the warburg effect and pancreatic cancer progression via transactivation of LDHA expression. Clin Cancer Res 2014; 20: 2595–2606.
Zhao S, Cao L, Freeman JW . Knockdown of RON receptor kinase delays but does not prevent tumor progression while enhancing HGF/MET signaling in pancreatic cancer cell lines. Oncogenesis 2013; 2: e76.
Sanders DA, Gormally MV, Marsico G, Beraldi D, Tannahill D, Balasubramanian S . FOXM1 binds directly to non-consensus sequences in the human genome. Genome Biol 2015; 16: 130.
Zhang Y, Zhang N, Dai B, Liu M, Sawaya R, Xie K et al. FoxM1B transcriptionally regulates vascular endothelial growth factor expression and promotes the angiogenesis and growth of glioma cells. Cancer Res 2008; 68: 8733–8742.
Wierstra I, Alves J . Despite its strong transactivation domain, transcription factor FOXM1c is kept almost inactive by two different inhibitory domains. Biol Chem 2006; 387: 963–976.
Dai B, Kang SH, Gong W, Liu M, Aldape KD, Sawaya R et al. Aberrant FoxM1B expression increases matrix metalloproteinase-2 transcription and enhances the invasion of glioma cells. Oncogene 2007; 26: 6212–6219.
Lam AK, Ngan AW, Leung MH, Kwok DC, Liu VW, Chan DW et al. FOXM1b, which is present at elevated levels in cancer cells, has a greater transforming potential than FOXM1c. Front Oncol 2013; 3: 11.
Wang H, Teh MT, Ji Y, Patel V, Firouzabadian S, Patel AA et al. EPS8 upregulates FOXM1 expression, enhancing cell growth and motility. Carcinogenesis 2010; 31: 1132–1141.
Mencalha AL, Binato R, Ferreira GM, Du Rocher B, Abdelhay E . Forkhead box M1 (FoxM1) gene is a new STAT3 transcriptional factor target and is essential for proliferation, survival and DNA repair of K562 cell line. PLoS One 2012; 7: e48160.
Christensen JG, Schreck R, Burrows J, Kuruganti P, Chan E, Le P et al. A selective small molecule inhibitor of c-Met kinase inhibits c-Met-dependent phenotypes in vitro and exhibits cytoreductive antitumor activity in vivo. Cancer Res 2003; 63: 7345–7355.
Chiu WT, Huang YF, Tsai HY, Chen CC, Chang CH, Huang SC et al. FOXM1 confers to epithelial-mesenchymal transition, stemness and chemoresistance in epithelial ovarian carcinoma cells. Oncotarget 2015; 6: 2349–2365.
Avan A, Maftouh M, Funel N, Ghayour-Mobarhan M, Boggi U, Peters GJ et al. MET as a potential target for the treatment of upper gastrointestinal cancers: characterization of novel c-Met inhibitors from bench to bedside. Curr Med Chem 2014; 21: 975–989.
Wierstra I, Alves J . FOXM1, a typical proliferation-associated transcription factor. Biol Chem 2007; 388: 1257–1274.
Wang Z, Ahmad A, Li Y, Banerjee S, Kong D, Sarkar FH . Forkhead box M1 transcription factor: a novel target for cancer therapy. Cancer Treat Rev 2010; 36: 151–156.
Qi J, McTigue MA, Rogers A, Lifshits E, Christensen JG, Janne PA et al. Multiple mutations and bypass mechanisms can contribute to development of acquired resistance to MET inhibitors. Cancer Res 2011; 71: 1081–1091.
Sunaga N, Kaira K, Imai H, Shimizu K, Nakano T, Shames DS et al. Oncogenic KRAS-induced epiregulin overexpression contributes to aggressive phenotype and is a promising therapeutic target in non-small-cell lung cancer. Oncogene 2013; 32: 4034–4042.
Altomare DA, Zhang L, Deng J, Di Cristofano A, Klein-Szanto AJ, Kumar R et al. GSK690693 delays tumor onset and progression in genetically defined mouse models expressing activated Akt. Clin Cancer Res 2010; 16: 486–496.
Sen N, Che X, Rajamani J, Zerboni L, Sung P, Ptacek J et al. Signal transducer and activator of transcription 3 (STAT3) and survivin induction by varicella-zoster virus promote replication and skin pathogenesis. Proc Natl Acad Sci USA 2012; 109: 600–605.
Li L, Li Z, Kong X, Xie D, Jia Z, Jiang W et al. Down-regulation of microRNA-494 via loss of SMAD4 increases FOXM1 and β-catenin signaling in pancreatic ductal adenocarcinoma cells. Gastroenterology 2014; 147: 485–497.
Liu M, Dai B, Kang SH, Ban K, Huang FJ, Lang FF et al. FoxM1B is overexpressed in human glioblastomas and critically regulates the tumorigenicity of glioma cells. Cancer Res 2006; 66: 3593–3602.
Hu B, Guo P, Bar-Joseph I, Imanishi Y, Jarzynka MJ, Bogler O et al. Neuropilin-1 promotes human glioma progression through potentiating the activity of the HGF/SF autocrine pathway. Oncogene 2007; 26: 5577–5586.
Yeh CY, Shin SM, Yeh HH, Wu TJ, Shin JW, Chang TY et al. Transcriptional activation of the Axl and PDGFR-alpha by c-Met through a ras- and Src-independent mechanism in human bladder cancer. BMC Cancer 2011; 11: 139.
Cui J, Shi M, Xie D, Wei D, Jia Z, Zheng S et al. FOXM1 Promotes the Warburg Effect and Pancreatic Cancer Progression via Transactivation of LDHA Expression. Clin Cancer Res 2014; 20: 2595–2606.
Xue J, Lin X, Chiu WT, Chen YH, Yu G, Liu M et al. Sustained activation of SMAD3/SMAD4 by FOXM1 promotes TGF-beta-dependent cancer metastasis. J Clin Invest 2014; 124: 564–579.
Zhang N, Wei P, Gong A, Chiu WT, Lee HT, Colman H et al. FoxM1 promotes β-catenin nuclear localization and controls Wnt target-gene expression and glioma tumorigenesis. Cancer Cell 2011; 20: 427–442.
Acknowledgements
We thank Don Norwood for editorial assistance and Xuemei Wang, Associate Director of Quantitative Research at The University of Texas MD Anderson Cancer Center, for assistance with statistical analyses. This work was supported by grants R01-CA129956, R01-CA148954, R01CA152309, and R01CA172233 from the National Cancer Institute, National Institutes of Health (to K Xie).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Oncogene website
Rights and permissions
About this article
Cite this article
Cui, J., Xia, T., Xie, D. et al. HGF/Met and FOXM1 form a positive feedback loop and render pancreatic cancer cells resistance to Met inhibition and aggressive phenotypes. Oncogene 35, 4708–4718 (2016). https://doi.org/10.1038/onc.2016.14
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2016.14
This article is cited by
-
Distinctive molecular features of regenerative stem cells in the damaged male germline
Nature Communications (2022)
-
Identification of a prognostic ferroptosis-related lncRNA signature in the tumor microenvironment of lung adenocarcinoma
Cell Death Discovery (2021)
-
Coregulation of pathways in lung cancer patients with EGFR mutation: therapeutic opportunities
British Journal of Cancer (2021)
-
Imidazopyridine hydrazone derivatives exert antiproliferative effect on lung and pancreatic cancer cells and potentially inhibit receptor tyrosine kinases including c-Met
Scientific Reports (2021)
-
Upregulation of FOXM1 leads to diminished drug sensitivity in myeloma
BMC Cancer (2018)