Abstract
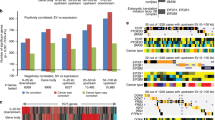
Mini-chromosome maintenance (Mcm) proteins are part of the replication-licensing complex that is loaded onto chromatin during the G1-phase of the cell cycle and required for initiation of DNA replication in the subsequent S-phase. Mcm proteins are typically loaded in excess of the number of locations that are used during S-phase. Nonetheless, partial depletion of Mcm proteins leads to cancers and stem cell deficiencies. Mcm2 deficient mice, on a 129Sv genetic background, display a high rate of thymic lymphoblastic lymphoma. Here array comparative genomic hybridization is used to characterize the genetic damage accruing in these tumors. The predominant events are deletions averaging less than 0.5 Mbp, considerably shorter than observed in prior studies using alternative mouse lymphoma models or human tumors. Such deletions facilitate identification of specific genes and pathways responsible for the tumors. Mutations in many genes that have been implicated in human lymphomas are recapitulated in this mouse model. These features, and the fact that the mutation underlying the accelerated genetic damage does not target a specific gene or pathway a priori, are valuable features of this mouse model for identification of tumor suppressor genes. Genes affected in all tumors include Pten, Tcfe2a, Mbd3 and Setd1b. Notch1 and additional genes are affected in subsets of tumors. The high frequency of relatively short deletions is consistent with elevated recombination between nearby stalled replication forks in Mcm2-deficient mice.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Abeysinghe SS, Chuzhanova N, Krawczak M, Ball EV, Cooper DN . (2003). Translocation and gross deletion breakpoints in human inherited disease and cancer I: nucleotide composition and recombination-associated motifs. Hum Mutat 22: 229.
Aftab S, Semenec L, Chu JS, Chen N . (2008). Identification and characterization of novel human tissue-specific RFX transcription factors. BMC Evol Biol 8: 226.
Bai T, Seebald JL, Kim KE, Ding HM, Szeto DP, Chang HC . (2010). Disruption of zebrafish cyclin G-associated kinase (GAK) function impairs the expression of Notch-dependent genes during neurogenesis and causes defects in neuronal development. BMC Dev Biol 10: 7.
Blow JJ, Dutta A . (2005). Preventing re-replication of chromosomal DNA. Nat Rev 6: 476–486.
Blow JJ, Ge XQ, Jackson DA . (2011). How dormant origins promote complete genome replication. Trends Biochem Sci 36: 405–414.
Brackertz M, Boeke J, Zhang R, Renkawitz R . (2002). Two highly related p66 proteins comprise a new family of potent transcriptional repressors interacting with MBD2 and MBD3. J Biol Chem 277: 40958–40966.
Buckler JL, Liu X, Turka LA . (2008). Regulation of T-cell responses by PTEN. Immunol Rev 224: 239–248.
Chuang CH, Wallace MD, Abratte C, Southard T, Schimenti JC . (2010). Incremental genetic perturbations to MCM2-7 expression and subcellular distribution reveal exquisite sensitivity of mice to DNA replication stress. PLoS Genet 6: e1001110.
Collins-Underwood JR, Mullighan CG . (2010). Genomic profiling of high-risk acute lymphoblastic leukemia. Leukemia 24: 1676–1685.
De Keersmaecker K, Real PJ, Gatta GD, Palomero T, Sulis ML, Tosello V et al. (2010). The TLX1 oncogene drives aneuploidy in T cell transformation. Nat Med 16: 1321–1327.
Ge XQ, Blow JJ . (2010). Chk1 inhibits replication factory activation but allows dormant origin firing in existing factories. J Cell Biol 191: 1285–1297.
Ge XQ, Jackson DA, Blow J . (2007). Dormant origins licensed by excess Mcm2-7 are required for human cells to survive replicative stress. Genes Dev 21: 3331–3341.
Hess JL . (2004). Mechanisms of transformation by MLL. Crit Rev Eukaryot Gene Exp 14: 235–254.
Ibarra A, Schwob E, Méndez J . (2008). Excess MCM proteins protect human cells from replicative stress by licensing backup origins of replication. Proc Natl Acad Sci USA 105: 8956–8961.
Jeannet R, Mastio J, Macias-Garcia A, Oravecz A, Ashworth T, Geimer Le Lay AS et al. (2010). Oncogenic activation of the Notch1 gene by deletion of its promoter in Ikaros-deficient T-ALL. Blood 116: 5443–5454.
Katzav S . (2009). Vav1: a hematopoietic signal transduction molecule involved in human malignancies. Int J Biochem Cell Biol 41: 1245–1248.
Kim J, Guermah M, McGinty RK, Lee JS, Tang Z, Milne TA et al. (2009). RAD6-mediated transcription-coupled H2B ubiquitylation directly stimulates H3K4 methylation in human cells. Cell 137: 459–471.
Kuiper RP, Schoenmakers EF, van Reijmersdal SV, Hehir-Kwa JY, van Kessel AG, van Leeuwen FN et al. (2007). High-resolution genomic profiling of childhood ALL reveals novel recurrent genetic lesions affecting pathways involved in lymphocyte differentiation and cell cycle progression. Leukemia 21: 1258–1266.
Kunnev D, Rusiniak ME, Kudla A, Freeland A, Cady GK, Pruitt SC . (2010). DNA damage response and tumorigenesis in Mcm2-deficient mice. Oncogene 29: 3630–3638.
LeBrun DP . (2003). E2A basic helix-loop-helix transcription factors in human leukemia. Front Biosci 8: s206–s222.
Ley TJ, Ding L, Walter MJ, McLellan MD, Lamprecht T, Larson DE et al. (2010). DNMT3A mutations in acute myeloid leukemia. N Engl J Med 363: 2424–2433.
Maser RS, Choudhury B, Campbell PJ, Feng B, Wong KK, Protopopov A et al. (2007). Chromosomally unstable mouse tumours have genomic alterations similar to diverse human cancers. Nature 447: 966–971.
Meyer C, Kowarz E, Hofmann J, Renneville A, Zuna J, Trka J et al. (2009). New insights to the MLL recombinome of acute leukemias. Leukemia 23: 1490–1499.
Mullighan CG, Goorha S, Radtke I, Miller CB, Coustan-Smith E, Dalton JD et al. (2007). Genome-wide analysis of genetic alterations in acute lymphoblastic leukaemia. Nature 446: 758–764.
Nishiyama A, Xin L, Sharov AA, Thomas M, Mowrer G, Meyers E et al. (2009). Uncovering early response of gene regulatory networks in ESCs by systematic induction of transcription factors. Cell Stem Cell 5: 420–433.
Orr SJ, Gaymes T, Ladon D, Chronis C, Czepulkowski B, Wang R . (2010). Reducing MCM levels in human primary T cells during the G(0)-->G(1) transition causes genomic instability during the first cell cycle. Oncogene 29: 3803–3814.
Palomero T, Sulis ML, Cortina M, Real PJ, Barnes K, Ciofani M et al. (2007). Mutational loss of PTEN induces resistance to NOTCH1 inhibition in T-cell leukemia. Nat Med 13: 1203–1210.
Pruitt SC, Bailey KJ, Freeland A . (2007). Reduced Mcm2 expression results in severe stem/progenitor cell deficiency and cancer. Stem Cells 25: 3121–3132.
Ramírez J, Hagman J . (2009). The Mi-2/NuRD complex: a critical epigenetic regulator of hematopoietic development, differentiation and cancer. Epigenetics 4: 532–536.
Rebollo A, Schmitt C . (2003). Ikaros, Aiolos and Helios: transcription regulators and lymphoid malignancies. Immunol Cell Biol 81: 171–175.
Remke M, Pfister S, Kox C, Toedt G, Becker N, Benner A et al. (2009). High-resolution genomic profiling of childhood T-ALL reveals frequent copy-number alterations affecting the TGF-beta and PI3K-AKT pathways and deletions at 6q15-16.1 as a genomic marker for unfavorable early treatment response. Blood 114: 1053–1062.
Santos J, Cerveira N, Correia C, Lisboa S, Pinheiro M, Torres L et al. (2010). Coexistence of alternative MLL-SEPT9 fusion transcripts in an acute myeloid leukemia with t(11;17)(q23;q25). Cancer Genet Cytogenet 197: 60–64.
Shima N, Alcaraz A, Liachko I, Buske TR, Andrews CA, Munroe RJ et al. (2007). A viable allele of Mcm4 causes chromosome instability and mammary adenocarcinomas in mice. Nat Genet 39: 93–98.
Tefferi A . (2010). Novel mutations and their functional and clinical relevance in myeloproliferative neoplasms: JAK2, MPL, TET2, ASXL1, CBL, IDH and IKZF1. Leukemia 24: 1128–1138.
Weng AP, Ferrando AA, Lee W, Morris 4th JP, Silverman LB, Sanchez-Irizarry C et al. (2004). Activating mutations of NOTCH1 in human T cell acute lymphoblastic leukemia. Science 306: 269–271.
Williams RT, Sherr CJ . (2008). The INK4-ARF (CDKN2A/B) locus in hematopoiesis and BCR-ABL-induced leukemias. Cold Spring Harb Symp Quant Biol 73: 461–467.
Yang Q, Kardava L, St Leger A, Martincic K, Varnum-Finney B, Bernstein ID et al. (2008). E47 controls the developmental integrity and cell cycle quiescence of multipotential hematopoietic progenitors. J Immunol 181: 5885–5894.
Zani VJ, Asou N, Jadayel D, Heward JM, Shipley J, Nacheva E et al. (1996). Molecular cloning of complex chromosomal translocation t(8;14;12)(q24.1;q32.3;q24.1) in a Burkitt lymphoma cell line defines a new gene (BCL7A) with homology to caldesmon. Blood 87: 3124–3134.
Acknowledgements
This work was supported by grants from the NIH-NCI, the Ellison Medical Foundation and NYSTEM to SCP. Cost of animal maintenance and flow cytometry was supported in part by an NCI-CCS grant to RPCI.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Oncogene website
Supplementary information
Rights and permissions
About this article
Cite this article
Rusiniak, M., Kunnev, D., Freeland, A. et al. Mcm2 deficiency results in short deletions allowing high resolution identification of genes contributing to lymphoblastic lymphoma. Oncogene 31, 4034–4044 (2012). https://doi.org/10.1038/onc.2011.566
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2011.566
Keywords
This article is cited by
-
Unwinding Helicase MCM Functionality for Diagnosis and Therapeutics of Replication Abnormalities Associated with Cancer: A Review
Molecular Diagnosis & Therapy (2024)
-
Examining transcriptional changes to DNA replication and repair factors over uveal melanoma subtypes
BMC Cancer (2018)
-
The transcription factor Rfx7 limits metabolism of NK cells and promotes their maintenance and immunity
Nature Immunology (2018)
-
An atypical 12q24.31 microdeletion implicates six genes including a histone demethylase KDM2B and a histone methyltransferase SETD1B in syndromic intellectual disability
Human Genetics (2016)