Abstract
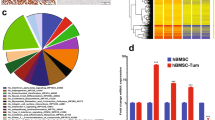
Cell identity is determined by its gene expression programs. The ability of a cell to change its identity and produce cell types outside its lineage is achieved by the activity of transcription controllers capable of reprogramming differentiation gene networks. The synovial sarcoma (SS)-associated protein, SYT–SSX2, reprograms myogenic progenitors and human bone marrow-derived mesenchymal stem cells (BMMSCs) by dictating their commitment to a pro-neural lineage. It fulfills this function by directly targeting an extensive array of neural-specific genes as well as genes of developmental pathway mediators. Concomitantly, the ability of both myoblasts and BMMSCs to differentiate into their normal myogenic and adipogenic lineages was compromised. SS is believed to arise in mesenchymal stem cells where formation of the t(X/18) translocation product, SYT–SSX, constitutes the primary event in the cancer. SYT–SSX is therefore believed to initiate tumorigenesis in its target stem cell. The data presented here allow a glimpse at the initial events that likely occur when SYT–SSX2 is first expressed, and its dominant function in subverting the nuclear program of the stem cell, leading to its aberrant differentiation, as a first step toward transformation. In addition, we identified the fibroblast growth factor receptor gene, Fgfr2, as one occupied and upregulated by SYT–SSX2. Knockdown of FGFR2 in both BMMSCs and SS cells abrogated their growth and attenuated their neural phenotype. These results support the notion that the SYT–SSX2 nuclear function and differentiation effects are conserved throughout sarcoma development and are required for its maintenance beyond the initial phase. They also provide the stem cell regulator, FGFR2, as a promising candidate target for future SS therapy.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Alfaro MP, Pagni M, Vincent A, Atkinson J, Hill MF, Cates J et al. (2008). The Wnt modulator sFRP2 enhances mesenchymal stem cell engraftment, granulation tissue formation and myocardial repair. Proc Natl Acad Sci USA 105: 18366–18371.
Asp P, Blum R, Vethantham V, Parisi F, Micsinai M, Cheng J et al. (2011). Genome-wide remodeling of the epigenetic landscape during myogenic differentiation. Proc Natl Acad Sci USA 108: E149–E158.
Barco R, Hunt LB, Frump AL, Garcia CB, Benesh A, Caldwell RL et al. (2007). The synovial sarcoma SYT–SSX2 oncogene remodels the cytoskeleton through activation of the ephrin pathway. Mol Biol Cell 18: 4003–4012.
Barco R, Garcia CB, Eid JE . (2009). The synovial sarcoma-associated SYT–SSX2 oncogene antagonizes the polycomb complex protein Bmi1. PLoS One 4: e5060.
Chadashvili T, Peterson DA . (2006). Cytoarchitecture of fibroblast growth factor receptor 2 (FGFR-2) immunoreactivity in astrocytes of neurogenic and non-neurogenic regions of the young adult and aged rat brain. J Comp Neurol 498: 1–15.
Charytonowicz E, Cordon-Cardo C, Matushansky I, Ziman M . (2009). Alveolar rhabdomyosarcoma: is the cell of origin a mesenchymal stem cell? Cancer Lett 279: 126–136.
Colter DC, Sekiya I, Prockop DJ . (2001). Identification of a subpopulation of rapidly self-renewing and multipotential adult stem cells in colonies of human marrow stromal cells. Proc Natl Acad Sci USA 98: 7841–7845.
Edgar R, Domrachev M, Lash AE . (2002). Gene expression omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res 30: 207–210.
Ever L, Gaiano N . (2005). Radial ‘glial’ progenitors: neurogenesis and signaling. Curr Opin Neurobiol 15: 29–33.
Farnham PJ . (2009). Insights from genomic profiling of transcription factors. Nat Rev Genet 10: 605–616.
Gurdon JB, Melton DA . (2008). Nuclear reprogramming in cells. Science 322: 1811–1815.
Haldar M, Hancock JD, Coffin CM, Lessnick SL, Capecchi MR . (2007). A conditional mouse model of synovial sarcoma: insights into a myogenic origin. Cancer Cell 11: 375–388.
Haldar M, Hedberg ML, Hockin MF, Capecchi MR . (2009). A CreER-based random induction strategy for modeling translocation-associated sarcomas in mice. Cancer Res 69: 3657–3664.
Huang W, Yang S, Shao J, Li YP . (2007). Signaling and transcriptional regulation in osteoblast commitment and differentiation. Front Biosci 12: 3068–3092.
Ishibe T, Nakayama T, Aoyama T, Nakamura T, Toguchida J . (2008). Neuronal differentiation of synovial sarcoma and its therapeutic application. Clin Orthop Relat Res 466: 2147–2155.
Ishibe T, Nakayama T, Okamoto T, Aoyama T, Nishijo K, Shibata KR et al. (2005). Disruption of fibroblast growth factor signal pathway inhibits the growth of synovial sarcomas: potential application of signal inhibitors to molecular target therapy. Clin Cancer Res 11: 2702–2712.
Katoh M . (2008). Cancer genomics and genetics of FGFR2. Int J Oncol 33: 233–237.
Katoh Y, Katoh M . (2009). FGFR2-related pathogenesis and FGFR2-targeted therapeutics. Int J Mol Med 23: 307–311.
Kawai A, Naito N, Yoshida A, Morimoto Y, Ouchida M, Shimizu K et al. (2004). Establishment and characterization of a biphasic synovial sarcoma cell line, SYO-1. Cancer Lett 204: 105–113.
Khan J, Bittner ML, Saal LH, Teichmann U, Azorsa DO, Gooden GC et al. (1999). cDNA microarrays detect activation of a myogenic transcription program by the PAX3-FKHR fusion oncogene. Proc Natl Acad Sci USA 96: 13264–13269.
Ladanyi M . (2001). Fusions of the SYT and SSX genes in synovial sarcoma. Oncogene 20: 5755–5762.
Lobo NA, Shimono Y, Qian D, Clarke MF . (2007). The biology of cancer stem cells. Annu Rev Cell Dev Biol 23: 675–699.
Lubieniecka JM, de Bruijn DR, Su L, van Dijk AHA, Subramanian S, van de Rijn M et al. (2008). Histone deacetylase inhibitors reverse SS18–SSX2-mediated polycomb silencing of the tumor suppressor early growth response 1 in synovial sarcoma. Cancer Res 68: 4303–4310.
Mackall CL, Meltzer PS, Helman LJ . (2004). Focus on sarcomas. Cancer Cell 2: 175–178.
Maric D, Fiorio Pla A, Chang YH, Barker JL . (2007). Self-renewing and differentiating properties of cortical neural stem cells are selectively regulated by basic fibroblast growth factor (FGF) signaling via specific FGF receptors. J Neurosci 27: 1836–1852.
Martens JA, Winston F . (2003). Current advances in understanding chromatin remodeling by Swi/Snf complexes. Curr Opin Genet Dev 13: 136–142.
Mateos-Langerak J, Cavalli G . (2008). Polycomb group proteins and long-range gene regulation. Adv Genet 61: 45–66.
Miraoui H, Marie PJ . (2010). Fibroblast growth factor receptor signaling crosstalk in skeletogenesis. Sci Signal 3: re9.
Muruganandan S, Roman AA, Sinal CJ . (2009). Adipocyte differentiation of bone marrow-derived mesenchymal stem cells: crosstalk with the osteoblastogenic program. Cell Mol Life Sci 66: 236–253.
Nagai M, Tanaka S, Tsuda M, Endo S, Kato H, Sonobe H et al. (2001). Analysis of transforming activity of human synovial sarcoma-associated chimeric protein SYT–SSX1 bound to chromatin remodeling factor hBRM/hSNF2α. Proc Natl Acad Sci USA 98: 3843–3848.
Naka N, Takenaka S, Araki N, Miwa T, Nobuyuki H, Yoshioka K et al. (2010). Synovial sarcoma is a stem cell malignancy. Stem Cells 28: 1119–1131.
Nielsen TO, West RB, Linn SC, Alter O, Knowling MA, O'Connell JX et al. (2002). Molecular characterisation of soft tissue tumours: a gene expression study. Lancet 359: 1301–1307.
Odelberg SJ, Kohlhoff A, Keating MT . (2000). Dedifferentiation of mammalian myotubes induced by msx1. Cell 103: 1099–1109.
Pardo OE, Latigo J, Jeffery RE, Nye E, Poulsom R, Spencer-Dene B et al. (2009). The fibroblast growth factor receptor inhibitor PD173074 blocks small cell lung cancer growth in vitro and in vivo. Cancer Res 69: 8645–8651.
Pretto D, Barco R, Rivera J, Neel N, Gustavson MD, Eid JE . (2006). The synovial sarcoma translocation protein SYT–SSX2 recruits b-catenin to the nucleus and associates with it in an active complex. Oncogene 25: 3661–3669.
Pruitt KD, Tatusova T, Maglott DR . (2007). NCBI Reference Sequence (RefSeq): a curated non-redundant database of genomes, transcripts and proteins. Nucleic Acids Res 35 (Database issue): D61–D65.
Schuettengruber B, Chourrout D, Vervoort M, Leblanc B, Cavalli G . (2007). Genome regulation by polycomb and trithorax proteins. Cell 128: 735–745.
Sekiya I, Larson BL, Smith JR, Pochampally R, Cui JG, Prockop DJ . (2002). Expansion of human adult stem cells from bone marrow stroma: conditions that maximize the yields of early progenitors and evaluate their quality. Stem Cells 20: 530–541.
Villegas SN, Canham M, Brickman JM . (2010). FGF signaling as a mediator of lineage transitions—evidence from embryonic stem cell differentiation. J Cell Biochem 110: 10–20.
Watanabe Y, Kameoka S, Gopalakrishnan V, Aldape KD, Pan ZZ, Lang FP et al. (2004). Conversion of myoblasts to physiologically active neuronal phenotype. Genes Dev 18: 889–900.
Zhang Y, Liu T, Meyer CA, Eeckhoute J, Johnson DS, Bernstein BE et al. (2008). Model-based analysis of ChIP-Seq (MACS). Genome Biol 9: R137.
Zhao C, Deng W, Gage FH . (2008). Mechanisms and functional implications of adult neurogenesis. Cell 132: 645–660.
Acknowledgements
Funding was provided by the Alex's Lemonade Stand Foundation (Innovation Award) and the Sarcoma Foundation of America. Microarrays were performed at the Vanderbilt Functional Genomics Shared Resource core supported by the VICC (P30 CA68485), the VDCC (P30 DK58404) and the Vanderbilt Vision Center (P30 EY08126). Some of the materials used in this work were provided by the Texas A&M Health Science Center College of Medicine Institute for Regenerative Medicine at Scott & White through a grant from NCRR of the NIH (grant no. P40RR017447). We thank BV Mary and H Trinity for assistance; T Ito and M Ladanyi for SYO-1 cells and A Sandelin and S Wilhite for technical support.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Oncogene website
Supplementary information
Rights and permissions
About this article
Cite this article
Garcia, C., Shaffer, C., Alfaro, M. et al. Reprogramming of mesenchymal stem cells by the synovial sarcoma-associated oncogene SYT–SSX2. Oncogene 31, 2323–2334 (2012). https://doi.org/10.1038/onc.2011.418
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2011.418
Keywords
This article is cited by
-
Biological and clinical implications of FGFR aberrations in paediatric and young adult cancers
Oncogene (2023)
-
Primary Renal Synovial Sarcoma and Clinical and Pathological Findings: a Systematic Review
Current Urology Reports (2021)
-
The Biology of Synovial Sarcoma: State-of-the-Art and Future Perspectives
Current Treatment Options in Oncology (2021)
-
Synovial Sarcoma: A Complex Disease with Multifaceted Signaling and Epigenetic Landscapes
Current Oncology Reports (2020)
-
Synovial sarcoma is a gateway to the role of chromatin remodeling in cancer
Cancer and Metastasis Reviews (2015)