Abstract
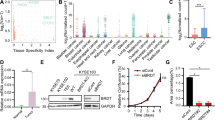
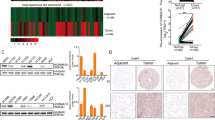
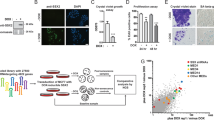
Cancer cells frequently express genes normally active in male germ cells. ATAD2 is one of them encoding a conserved factor harbouring an AAA type ATPase domain and a bromodomain. We show here that ATAD2 is highly expressed in testis as well as in many cancers of different origins and that its high expression is a strong predictor of rapid mortality in lung and breast cancers. These observations suggest that ATAD2 acts on upstream and basic cellular processes to enhance oncogenesis in a variety of unrelated cell types. Accordingly, our functional studies show that ATAD2 controls chromatin dynamics, genome transcriptional activities and apoptotic cell response. We could also highlight some of the important intrinsic properties of its two regulatory domains, including a functional cross-talk between the AAA ATPase domain and the bromodomain. Altogether, these data indicate that ATAD2 overexpression in somatic cells, by acting on basic properties of chromatin, may contribute to malignant transformation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Accession codes
References
Boussouar F, Rousseaux S, Khochbin S . (2008). A new insight into male genome reprogramming by histone variants and histone code. Cell Cycle 7: 3499–3502.
Ciro M, Prosperini E, Quarto M, Grazini U, Walfridsson J, McBlane F et al. (2009). ATAD2 Is a Novel Cofactor for MYC, Overexpressed and Amplified in Aggressive Tumors. Cancer Res 69: 8491–8498.
Col E, Caron C, Chable-Bessia C, Legube G, Gazzeri S, Komatsu Y et al. (2005). HIV-1 Tat targets Tip60 to impair the apoptotic cell response to genotoxic stresses. EMBO J 24: 2634–2645.
De Koning L, Savignoni A, Boumendil C, Rehman H, Asselain B, Sastre-Garau X et al. (2009). Heterochromatin protein 1alpha: a hallmark of cell proliferation relevant to clinical oncology. EMBO Mol Med 1: 178–191.
Fillingham J, Kainth P, Lambert JP, van Bakel H, Tsui K, Pena-Castillo L et al. (2009). Two-color cell array screen reveals interdependent roles for histone chaperones and a chromatin boundary regulator in histone gene repression. Mol Cell 35: 340–351.
Gradolatto A, Rogers RS, Lavender H, Taverna SD, Allis CD, Aitchison JD et al. (2008). Saccharomyces cerevisiae Yta7 regulates histone gene expression. Genetics 179: 291–304.
Hartman HB, Yu J, Alenghat T, Ishizuka T, Lazar MA . (2005). The histone-binding code of nuclear receptor co-repressors matches the substrate specificity of histone deacetylase 3. EMBO Rep 6: 445–451.
Jambunathan N, Martinez AW, Robert EC, Agochukwu NB, Ibos ME, Dugas SL et al. (2005). Multiple bromodomain genes are involved in restricting the spread of heterochromatic silencing at the Saccharomyces cerevisiae HMR-tRNA boundary. Genetics 171: 913–922.
Ladurner AG, Inouye C, Jain R, Tjian R . (2003). Bromodomains mediate an acetyl-histone encoded antisilencing function at heterochromatin boundaries. Mol Cell 11: 365–376.
Lenz G, Wright G, Dave SS, Xiao W, Powell J, Zhao H et al. (2008). Stromal gene signatures in large-B-cell lymphomas. N Engl J Med 359: 2313–2323.
Perche PY, Vourc'h C, Konecny L, Souchier C, Robert-Nicoud M, Dimitrov S et al. (2000). Higher concentrations of histone macroH2A in the Barr body are correlated with higher nucleosome density. Curr Biol 10: 1531–1534.
Pivot-Pajot C, Caron C, Govin J, Vion A, Rousseaux S, Khochbin S . (2003). Acetylation-dependent chromatin reorganization by BRDT, a testis-specific bromodomain-containing protein. Mol Cell Biol 23: 5354–5365.
Rousseaux S, Khochbin S . (2009). New hypotheses for large-scale epigenome alterations in somatic cancer cells: a role for male germ-cell-specific regulators. Epigenomics 1: 153–161.
Rubin DM, Glickman MH, Larsen CN, Dhruvakumar S, Finley D . (1998). Active site mutants in the six regulatory particle ATPases reveal multiple roles for ATP in the proteasome. EMBO J 17: 4909–4919.
Sobel RE, Cook RG, Perry CA, Annunziato AT, Allis CD . (1995). Conservation of deposition-related acetylation sites in newly synthesized histones H3 and H4. Proc Natl Acad Sci USA 92: 1237–1241.
Sung P, Higgins D, Prakash L, Prakash S . (1988). Mutation of lysine-48 to arginine in the yeast RAD3 protein abolishes its ATPase and DNA helicase activities but not the ability to bind ATP. EMBO J 7: 3263–3269.
Tackett AJ, Dilworth DJ, Davey MJ, O'Donnell M, Aitchison JD, Rout MP et al. (2005). Proteomic and genomic characterization of chromatin complexes at a boundary. J Cell Biol 169: 35–47.
Tseng RJ, Armstrong KR, Wang X, Chamberlin HM . (2007). The bromodomain protein LEX-1 acts with TAM-1 to modulate gene expression in C.elegans. Mol Genet Genomics 278: 507–518.
Yu J, Li Y, Ishizuka T, Guenther MG, Lazar MA . (2003). A SANT motif in the SMRT corepressor interprets the histone code and promotes histone deacetylation. EMBO J 22: 3403–3410.
Zou JX, Guo L, Revenko AS, Tepper CG, Gemo AT, Kung HJ et al. (2009). Androgen-induced coactivator ANCCA mediates specific androgen receptor signaling in prostate cancer. Cancer Res 69: 3339–3346.
Zou JX, Revenko AS, Li LB, Gemo AT, Chen HW . (2007). ANCCA, an estrogen-regulated AAA+ ATPase coactivator for ERalpha, is required for coregulator occupancy and chromatin modification. Proc Natl Acad Sci USA 104: 18067–18072.
Acknowledgements
This work was supported by the ‘ANR blanc-Episperm’, INCa-DHOS’ and ‘ARC-ARECA’ research programs. CL and SM are supported by the PhD fellowship program of Région Rhône-Alpes and the post-doc fellowship program of INCa (AO 2006), respectively. AD salary is paid by the INCa–DHOS funds. Transcriptomic analyses of lung cancer patients were obtained thanks to the Ligue Contre le Cancer ‘Carte d’identité des Tumeurs’ (CIT) program. We acknowledge the support of the Nice-Sophia Antipolis Transcriptome Platform of the Marseille–Nice Genopole, in which the microarray experiments were carried out. Special thanks are due to Géraldine Rios for microarray production. We also acknowledge the support of the microscopy platform, Albert Bonniot Institute, Grenoble, and thank Catherine Souchier for her precious help in the FRAP experiments.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Oncogene website
Rights and permissions
About this article
Cite this article
Caron, C., Lestrat, C., Marsal, S. et al. Functional characterization of ATAD2 as a new cancer/testis factor and a predictor of poor prognosis in breast and lung cancers. Oncogene 29, 5171–5181 (2010). https://doi.org/10.1038/onc.2010.259
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2010.259
Keywords
This article is cited by
-
A targetable MYBL2-ATAD2 axis governs cell proliferation in ovarian cancer
Cancer Gene Therapy (2023)
-
Effect of chronic intermittent hypoxia-induced HIF-1α/ATAD2 expression on lung cancer stemness
Cellular & Molecular Biology Letters (2022)
-
ATAD2 interacts with C/EBPβ to promote esophageal squamous cell carcinoma metastasis via TGF-β1/Smad3 signaling
Journal of Experimental & Clinical Cancer Research (2021)
-
A genome-scale CRISPR Cas9 dropout screen identifies synthetically lethal targets in SRC-3 inhibited cancer cells
Communications Biology (2021)
-
Evaluation of ATAD2 as a Potential Target in Hepatocellular Carcinoma
Journal of Gastrointestinal Cancer (2021)