Key Points
-
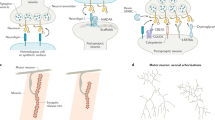
Synaptic activity regulates the expression of neuronal gene products that are important for neuronal survival and differentiation, synaptogenesis and complex behaviour. Diverse mechanisms modulate the activity of positive and negative transcriptional regulators.
-
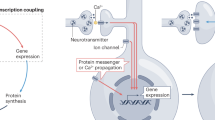
Members of the cyclic-AMP-response-element-binding protein (CREB) family of transcription factors are among the most widely studied activity-dependent transcription factors in the brain. It is unclear how synaptic activity in dendrites activates CREB in the nucleus. Protein complexes associated with plasma-membrane calcium channels seem to be crucial for transducing calcium influx into changes in gene transcription. The selective association of signalling molecules with distinct calcium channels might explain how the route of calcium entry can influence the ability of calcium to activate gene transcription.
-
CREB is activated by activity-dependent phosphorylation at Ser133, which works, at least in part, through the inducible recruitment of the transcriptional coactivator CREB-binding protein (CBP). Neuronal stimuli that cause the influx of calcium lead to the phosphorylation of CREB at additional residues, including Ser142 and Ser143, which block the binding of the KIX domain of CBP to CREB Ser133. So, CREB might also mediate transcription through alternative mechanisms, independently of CBP recruitment.
-
Stimuli induce diverse biological responses, in part by initiating distinct programmes of gene expression. In neurons, many stimuli induce the expression of c-Fos, but only stimuli that lead to the elevation of intracellular calcium induce Bdnf expression. Transcription of Bdnf is cooperatively regulated by a number of factors, including the transcription factor CaRF (calcium-response factor). CaRF is activated in a calcium- and neuron-selective manner, and might confer stimulus and cell-type selectivity on the induction of its target genes.
-
Calcium can also directly regulate transcription factors. Calcium binding to the transcriptional repressor DREAM (downstream-response-element-antagonist modulator) inhibits the ability of DREAM to bind DNA, relieving the repression of target genes such as prodynorphin. Dream-knockout mice show altered pain perception and enhanced expression of dynorphin, indicating that DREAM is a crucial transcriptional regulator of prodynorphin in vivo.
-
Neuronal activity also activates transcription by relocalizing latent transcription factors from the cytoplasm to the nucleus. NFAT (nuclear factor of activated T cells) and NF-κB (nuclear factor-κB) are expressed in the nervous system and relocate to the nucleus in response to neuronal stimuli. Although the role of these factors in neural function is unclear, their distribution outside the nucleus, where they can respond to local stimuli, indicates a potential involvement in processes such as axon guidance and synaptic modulation.
-
Neuronal stimuli modulate the chromatin architecture of target promoters, thereby regulating transcriptional initiation. Chromatin structure can be altered by post-translational modifications that are mediated by inducibly recruited coactivator or corepressor molecules. Among the best-characterized chromatin modifications is the reversible acetylation of histone proteins. This modification is controlled by the opposing actions of cellular histone acetyl-transferases (HATs) and histone deacetylases (HDACs). The activity of both HATs and HDACs is regulated by calcium signalling, and further work will be required to determine the physiological role of these transcriptional regulators in neuronal function.
-
Future advances in our understanding of how signal-specific transcriptional programmes are generated and maintained in the developing and mature nervous system should benefit from the increased use of new approaches, such as bioinformatics, chromatin immunoprecipitation and gene-expression profiling. The development and use of new imaging techniques, together with improved targeted animal models, should expand our understanding of the physiological roles of activity-dependent transcription in brain development and function.
Abstract
Synaptic activity regulates the expression of a set of neuronal gene products that are important for neuronal survival and differentiation, synaptogenesis and, ultimately, complex behaviour. Activity-dependent signalling pathways induce neuronal gene transcription by modulating the function of both transcriptional activators and repressors, and recent studies have revealed significant diversity in the mechanisms that control the activity of these transcriptional regulators. Investigators have begun to elucidate the distinct functions of individual activity-regulated transcription factors, and to explore how these factors cooperate to provide stimulus specificity in the initiation of neuronal transcriptional programmes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Greenberg, M. E., Ziff, E. B. & Greene, L. A. Stimulation of neuronal acetylcholine receptors induces rapid gene transcription. Science 234, 80–83 (1986). The first demonstration that neurotransmitters can rapidly induce gene transcription.
Huh, G. S. et al. Functional requirement for class I MHC in CNS development and plasticity. Science 290, 2155–2159 (2000).
Kandel, E. R. The molecular biology of memory storage: a dialogue between genes and synapses. Science 294, 1030–1038 (2001).
Nestler, E. J. Molecular basis of long-term plasticity underlying addiction. Nature Rev. Neurosci. 2, 119–128 (2001).
Spitzer, N. C., Lautermilch, N. J., Smith, R. D. & Gomez, T. M. Coding of neuronal differentiation by calcium transients. Bioessays 22, 811–817 (2000).
Brown, J. R., Ye, H., Bronson, R. T., Dikkes, P. & Greenberg, M. E. A defect in nurturing in mice lacking the immediate early gene fosB. Cell 86, 297–309 (1996).
Reppert, S. M. & Weaver, D. R. Molecular analysis of mammalian circadian rhythms. Annu. Rev. Physiol. 63, 647–676 (2001).
Travnickova-Bendova, Z., Cermakian, N., Reppert, S. M. & Sassone-Corsi, P. Bimodal regulation of mPeriod promoters by CREB-dependent signaling and CLOCK/BMAL1 activity. Proc. Natl Acad. Sci. USA 99, 7728–7733 (2002).
Gau, D. et al. Phosphorylation of CREB Ser142 regulates light-induced phase shifts of the circadian clock. Neuron 34, 245–253 (2002). This paper provides evidence for a physiological role of the phosphorylation of CREB at Ser142 in transcriptional activation, by the introduction of a Ser142Ala mutation into the genomic CREB locus.
Takasu, M. A., Dalva, M. B., Zigmond, R. E. & Greenberg, M. E. Modulation of NMDA receptor-dependent calcium influx and gene expression through EphB receptors. Science 295, 491–495 (2002).
Lanahan, A. & Worley, P. Immediate–early genes and synaptic function. Neurobiol. Learn. Mem. 70, 37–43 (1998). PubMed |
Hevroni, D. et al. Hippocampal plasticity involves extensive gene induction and multiple cellular mechanisms. J. Mol. Neurosci. 10, 75–98 (1998).
O'Brien, R. J. et al. Synaptic clustering of AMPA receptors by the extracellular immediate–early gene product Narp. Neuron 23, 309–323 (1999).
Corriveau, R. A., Huh, G. S. & Shatz, C. J. Regulation of class I MHC gene expression in the developing and mature CNS by neural activity. Neuron 21, 505–520 (1998).
Nedivi, E., Wu, G. -Y. & Cline, H. T. Promotion of dendritic growth by CPG15, an activity-induced signaling molecule. Science 281, 1863–1866 (1998).
Poo, M. -M. Neurotrophins as synaptic modulators. Nature Rev. Neurosci. 2, 24–32 (2001).
Carafoli, E., Santella, L., Branca, D. & Brini, M. Generation, control, and processing of cellular calcium signals. Crit. Rev. Biochem. Mol. Biol. 36, 107–260 (2001).
Krichevsky, A. M. & Kosik, K. S. Neuronal RNA granules: a link between RNA localization and stimulation-dependent translation. Neuron 32, 683–696 (2001).
Xie, J. & Black, D. L. A CaMK IV responsive RNA element mediates depolarization-induced alternative splicing of ion channels. Nature 410, 936–939 (2001).
West, A. E. et al. Calcium regulation of neuronal gene expression. Proc. Natl Acad. Sci. USA 98, 11024–11031 (2001).
Cruzalegui, F. H. & Bading, H. Calcium-regulated protein kinase cascades and their transcription factor targets. Cell. Mol. Life Sci. 57, 402–410 (2000).
Mayr, B. & Montminy, M. Transcriptional regulation by the phosphorylation-dependent factor CREB. Nature Rev. Mol. Cell Biol. 2, 599–609 (2001).
Shaywitz, A. J. & Greenberg, M. E. CREB: a stimulus-induced transcription factor activated by a diverse array of extracellular signals. Annu. Rev. Biochem. 68, 821–861 (1999).
Lonze, B. E. & Ginty, D. D. Function and regulation of CREB family transcription factors in the nervous system. Neuron 35, 605–623 (2002).
Shieh, P. B., Hu, S. -C., Bobb, K., Timmusk, T. & Ghosh, A. Identification of a signaling pathway involved in calcium regulation of BDNF expression. Neuron 20, 727–740 (1998).
Tao, X., Finkbeiner, S., Arnold, D. B., Shaywitz, A. J. & Greenberg, M. E. Ca2+ influx regulates BDNF transcription by a CREB family transcription factor-dependent mechanism. Neuron 20, 709–726 (1998).
Sasaki, M. et al. Dynamic regulation of neuronal NO synthase transcription by calcium influx through a CREB family transcription factor-dependent mechanism. Proc. Natl Acad. Sci. USA 97, 8617–8622 (2000).
Mitsuda, N. et al. Activated cAMP-response element-binding protein regulates neuronal expression of presenilin-1. J. Biol. Chem. 276, 9688–9698 (2001).
Riccio, A., Ahn, S., Davenport, C. M., Blendy, J. A. & Ginty, D. D. Mediation by a CREB family transcription factor of NGF-dependent survival of sympathetic neurons. Science 286, 2358–2361 (1999).
Bonni, A. et al. Cell survival promoted by the Ras–MAPK signaling pathway by transcription-dependent and -independent mechanisms. Science 286, 1358–1362 (1999).
Lonze, B. E., Riccio, A., Cohen, S. & Ginty, D. D. Apoptosis, axonal growth defects, and degeneration of peripheral neurons in mice lacking CREB. Neuron 34, 371–385 (2002).
Hummler, E. et al. Targeted mutation of the CREB gene: compensation within the CREB/ATF family of transcription factors. Proc. Natl Acad. Sci. USA 91, 5647–5651 (1994).
Mantamadiotis, T. et al. Disruption of CREB function in brain leads to neurodegeneration. Nature Genet. 31, 47–54 (2002). References 31 and 33 use CREB-deficient mice to show a crucial role for CREB in neuronal survival in vivo.
Bleckmann, S. C. et al. Activating transcription factor 1 and CREB are important for cell survival during early mouse development. Mol. Cell. Biol. 22, 1919–1925 (2002).
Barco, A., Alarcon, J. M. & Kandel, E. R. Expression of constitutively active CREB protein facilitates the late phase of long-term potentiation by enhancing synaptic capture. Cell 108, 689–703 (2002).
Pittenger, C. et al. Reversible inhibition of CREB/ATF transcription factors in region CA1 of the dorsal hippocampus disrupts hippocampus-dependent spatial memory. Neuron 34, 447–462 (2002).
Ahn, S., Ginty, D. D. & Linden, D. J. A late phase of cerebellar long-term depression requires activation of CaMKIV and CREB. Neuron 23, 559–568 (1999).
Sheng, M., Thompson, M. A. & Greenberg, M. E. CREB: a Ca2+-regulated transcription factor phosphorylated by calmodulin-dependent kinases. Science 252, 1427–1430 (1991).
Gonzalez, G. A. & Montminy, M. R. Cyclic AMP stimulates somatostatin gene transcription by phosphorylation of CREB at serine 133. Cell 59, 675–680 (1989).
Ho, N. et al. Impaired synaptic plasticity and cAMP response element binding protein activation in Ca2+/calmodulin-dependent protein kinase type IV/Gr-deficient mice. J. Neurosci. 20, 6459–6472 (2000).
Ribar, T. J. et al. Cerebellar defects in Ca2+/calmodulin kinase IV-deficient mice. J. Neurosci. 20, RC107 (2000).
Wu, G. -Y., Deisseroth, K. & Tsien, R. W. Activity-dependent CREB phosphorylation: convergence of a fast, sensitive calmodulin kinase pathway and a slow, less sensitive mitogen-activated protein kinase pathway. Proc. Natl Acad. Sci. USA 98, 2808–2813 (2001).
Dolmetsch, R. E., Pajvani, U., Fife, K., Spotts, J. M. & Greenberg, M. E. Signaling to the nucleus by an L-type calcium channel–calmodulin complex through the MAP kinase pathway. Science 294, 333–339 (2001). This paper shows that the association of the signalling molecule calmodulin with the L-VGCC is required for calcium entry through the L-VGCC to activate CREB-dependent transcription.
Hardingham, G. E., Arnold, F. J. L. & Bading, H. Nuclear calcium signaling controls CREB-mediated gene expression triggered by synaptic activity. Nature Neurosci. 4, 261–267 (2001).
Deisseroth, K., Heist, E. K. & Tsien, R. W. Translocation of calmodulin to the nucleus supports CREB phosphorylation in hippocampal neurons. Nature 392, 198–202 (1998).
Bading, H., Ginty, D. D. & Greenberg, M. E. Regulation of gene expression in hippocampal neurons by distinct calcium signaling pathways. Science 260, 181–186 (1993).
Deisseroth, K., Bito, H. & Tsien, R. W. Signaling from synapse to nucleus: postsynaptic CREB phosphorylation during multiple forms of hippocampal synaptic plasticity. Neuron 16, 89–101 (1996).
Zuhlke, R. D., Pitt, G. S., Deisseroth, K., Tsien, R. W. & Reuter, H. Calmodulin supports both inactivation and facilitation of L-type calcium channels. Nature 399, 159–162 (1999).
Lee, A. et al. Ca2+/calmodulin binds to and modulates P/Q-type calcium channels. Nature 399, 155–159 (1999).
Peterson, B. Z., DeMaria, C. D., Adelman, J. P. & Yue, D. T. Calmodulin is the Ca2+ sensor for Ca2+-dependent inactivation of L-type calcium channels. Neuron 22, 549–558 (1999).
Cullen, P. J. & Lockyer, P. J. Integration of calcium and Ras signalling. Nature Rev. Mol. Cell Biol. 3, 339–348 (2002).
Hardingham, G. E., Arnold, F. J. & Bading, H. A calcium microdomain near NMDA receptors: on switch for ERK-dependent synapse-to-nucleus communication. Nature Neurosci. 4, 565–566 (2001).
Sala, C., Rudolph-Correia, S. & Sheng, M. Developmentally regulated NMDA receptor-dependent dephosphorylation of cAMP response element-binding protein (CREB) in hippocampal neurons. J. Neurosci. 20, 3529–3536 (2000).
Hardingham, G. E., Fukunaga, Y. & Bading, H. Extrasynaptic NMDARs oppose synaptic NMDARs by triggering CREB shut-off and cell death pathways. Nature Neurosci. 5, 405–414 (2002). An important paper that shows opposing roles of synaptic and extrasynaptic NMDARs in CREB activation and cell survival.
Kwok, R. P. S. et al. Nuclear protein CBP is a coactivator for the transcription factor CREB. Nature 379, 223–226 (1994).
Vo, N. & Goodman, R. H. CREB-binding protein and p300 in transcriptional regulation. J. Biol. Chem. 276, 13505–13508 (2001).
Impey, S. et al. Phosphorylation of CBP mediates transcriptional activation by neural activity and CaM kinase IV. Neuron 34, 235–244 (2002). The authors identify Ser301 as a site of calcium-inducible phosphorylation on the coactivator CBP in hippocampal neurons.
Wu, X. & McMurray, C. T. Calmodulin kinase II attenuation of gene transcription by preventing cAMP response element-binding protein (CREB) dimerization and binding of the CREB-binding protein. J. Biol. Chem. 276, 1735–1741 (2001).
Parker, D. et al. Analysis of an activator:coactivator complex reveals an essential role for secondary structure in transcriptional activation. Mol. Cell 2, 353–359 (1998).
Kornhauser, J. M. et al. CREB transcriptional activity in neurons is regulated by multiple, calcium-specific phosphorylation events. Neuron 34, 221–233 (2002).
Fimia, G. M., De Cesare, D. & Sassone-Corsi, P. CBP-independent activation of CREM and CREB by the LIM-only protein ACT. Nature 398, 165–169 (1999).
Bartel, D. P., Sheng, M., Lau, L. F. & Greenberg, M. E. Growth factors and membrane depolarization activate distinct programs of early response gene expression: dissociation of fos and jun induction. Genes Dev. 3, 304–313 (1989).
Mayr, B. M., Canettieri, G. & Montminy, M. R. Distinct effects of cAMP and mitogenic signals on CREB-binding protein recruitment impart specificity to target gene activation via CREB. Proc. Natl Acad. Sci. USA 98, 10936–10941 (2001).
Buonanno, A. & Fields, R. D. Gene regulation by patterned electrical activity during neural and skeletal muscle development. Curr. Opin. Neurobiol. 9, 110–120 (1999).
Tao, X., West, A. E., Chen, W. G., Corfas, G. & Greenberg, M. E. A calcium-responsive transcription factor, CaRF, that regulates neuronal activity-dependent expression of BDNF. Neuron 33, 383–395 (2002).
Burgoyne, R. D. & Weiss, J. L. The neuronal calcium sensor family of Ca2+-binding proteins. Biochem. J. 353, 1–12 (2001).
Carrion, A. M., Mellström, B. & Narranjo, J. R. Protein kinase A-dependent derepression of the human prodynorphin gene via differential binding to an intragenic silencer element. Mol. Cell. Biol. 18, 6921–6929 (1998).
Carrion, A. M., Link, W. A., Ledo, F., Mellström, B. & Naranjo, J. R. DREAM is a Ca2+-regulated transcriptional repressor. Nature 398, 80–84 (1999).
Cheng, H. Y. et al. DREAM is a critical transcriptional repressor for pain modulation. Cell 108, 31–43 (2002). An important paper that confirms the role of DREAM as a regulator of Pdyn gene expression in vivo by the generation and analysis of DREAM-deficient animals.
Buxbaum, J. D. et al. Calsenilin: a calcium-binding protein that interacts with the presenilins and regulates the levels of a presenilin fragment. Nature Med. 4, 1177–1181 (1998).
An, W. F. et al. Modulation of A-type potassium channels by a family of calcium sensors. Nature 403, 553–556 (2000).
Osawa, M. et al. Calcium-regulated DNA binding and oligomerization of the neuronal calcium-sensing protein, calsenilin/DREAM/KChIP3. J. Biol. Chem. 276, 41005–41013 (2001).
Carrion, A. M., Link, W. A., Ledo, F., Mellstrom, B. & Naranjo, J. R. DREAM is a Ca2+-regulated transcriptional repressor. Nature 398, 80–84 (1999).
Ledo, F., Carrion, A. M., Link, W. A., Mellstrom, B. & Naranjo, J. R. DREAM–αCREM interaction via leucine-charged domains derepresses downstream regulatory element-dependent transcription. Mol. Cell. Biol. 20, 9120–9126 (2000).
Ledo, F., Kremer, L., Mellström, B. & Naranjo, J. R. Ca2+-dependent block of CREB–CBP transcription by repressor DREAM. EMBO J. 21, 4583–4592 (2002).
Crabtree, G. R. Generic signals and specific outcomes: signaling through Ca2+, calcineurin, and NF-AT. Cell 96, 611–614 (1999).
Ho, A. M., Jain, J., Rao, A. & Hogan, P. G. Expression of the transcription factor NFATp in a neuronal cell line and in the murine nervous system. J. Biol. Chem. 269, 28181–28186 (1994).
Graef, I. A. et al. L-type calcium channels and GSK-3 regulate the activity of NF-ATc4 in hippocampal neurons. Nature 401, 703–708 (1999).
Genazzani, A. A., Carafoli, E. & Guerini, D. Calcineurin controls inositol 1,4,5-trisphosphate type 1 receptor expression in neurons. Proc. Natl Acad. Sci. USA 96, 5797–5801 (1999).
Berridge, M. J. Neuronal calcium signaling. Neuron 21, 13–26 (1998).
Feske, S., Giltnane, J., Dolmetsch, R., Staudt, L. M. & Rao, A. Gene regulation mediated by calcium signals in T-lymphocytes. Nature Immunol. 2, 316–324 (2001).
Ghosh, S., May, M. J. & Kopp, E. B. NF-κB and Rel proteins: evolutionarily conserved mediators of immune responses. Annu. Rev. Immunol. 16, 225–260 (1998).
Kaltschmidt, C., Kaltschmidt, B. & Baeuerle, P. A. Brain synapses contain inducible forms of the transcription factor NF-κB. Mech. Dev. 43, 135–147 (1993).
Kaltschmidt, C., Kaltschmidt, B., Neumann, H., Wekerle, H. & Baeuerle, P. A. Constitutive NF-κB activity in neurons. Mol. Cell. Biol. 14, 3981–3992 (1994).
Meberg, P. J., Kinney, W. R., Valcourt, E. G. & Routtenberg, A. Gene expression of the transcription factor NF-κB in hippocampus: regulation by synaptic activity. Brain Res. Mol. Brain Res. 38, 179–190 (1996).
Guerrini, L., Blasi, F. & Denis-Donini, S. Synaptic activation of NF-κB by glutamate in cerebellar granule neurons in vitro. Proc. Natl Acad. Sci. USA 92, 9077–9081 (1995).
Kaltschmidt, C., Kaltschmidt, B. & Baeuerle, P. A. Stimulation of ionotropic glutamate receptors activates transcription factor NF-κB in primary neurons. Proc. Natl Acad. Sci. USA 92, 9618–9622 (1995).
Wellmann, H., Kaltschmidt, B. & Kaltschmidt, C. Retrograde transport of transcription factor NF-κB in living neurons. J. Biol. Chem. 276, 11821–11829 (2001).
Mattson, M. P., Culmsee, C., Yu, Z. & Camandola, S. Roles of nuclear factor κB in neuronal survival and plasticity. J. Neurochem. 74, 443–456 (2000).
Kaltschmidt, B., Uherek, M., Wellmann, H. & Volk, B. Inhibition of NF-κB potentiates amyloid β-mediated neuronal apoptosis. Proc. Natl Acad. Sci. USA 96, 9409–9414 (1999).
Grilli, M., Pizzi, M., Memo, M. & Spano, P. Neuroprotection by aspirin and sodium salicylate through blockade of NF-κB activation. Science 274, 1383–1385 (1996).
Beattie, E. C. et al. Control of synaptic strength by TNFα. Science 295, 2282–2285 (2002).
Berger, S. L. Histone modifications in transcriptional regulation. Curr. Opin. Genet. Dev. 12, 142–148 (2002).
Brown, C. E., Lechner, T., Howe, L. & Workman, J. L. The many HATs of transcription coactivators. Trends Biochem. Sci. 25, 15–19 (2000).
Ng, H. H. & Bird, A. Histone deacetylases: silencers for hire. Trends Biochem. Sci. 25, 121–126 (2000).
Chawla, S., Hardingham, G. E., Quinn, D. R. & Bading, H. CBP: a signal-regulated transcriptional coactivator controlled by nuclear calcium and CaM kinase IV. Science 281, 1505–1509 (1998).
Hu, S. C., Chrivia, J. & Ghosh, A. Regulation of CBP-mediated transcription by neuronal calcium signaling. Neuron 22, 799–808 (1999).
Black, B. L. & Olsen, E. N. Transcriptional control of muscle development by myocyte enhancer factor-2 (MEF2) proteins. Annu. Rev. Cell Dev. Biol. 14, 167–196 (1998).
Mao, Z., Bonni, A., Xia, F., Nadal-Vicens, M. & Greenberg, M. E. Neuronal activity-dependent cell survival mediated by transcription factor MEF2. Science 286, 785–790 (1999).
Okamoto, S., Krainc, D., Sherman, K. & Lipton, S. A. Antiapoptotic role of the p38 mitogen-activated protein kinase–myocyte enhancer factor 2 transcription factor pathway during neuronal differentiation. Proc. Natl Acad. Sci. USA 97, 7561–7566 (2000).
Miska, E. A. et al. HDAC4 deacetylase associates with and represses the MEF2 transcription factor. EMBO J. 18, 5099–5107 (1999).
Wang, A. H. et al. HDAC4, a human histone deacetylase related to yeast HDA1, is a transcriptional corepressor. Mol. Cell. Biol. 19, 7816–7827 (1999).
Lemercier, C. et al. mHDA1/HDAC5 histone deacetylase interacts with and represses MEF2A transcriptional activity. J. Biol. Chem. 275, 15594–15599 (2000).
Dressel, U. et al. A dynamic role for HDAC7 in MEF2-mediated muscle differentiation. J. Biol. Chem. 276, 17007–17013 (2001).
Zhou, X., Marks, P. A., Rifkind, R. A. & Richon, V. M. Cloning and characterization of a histone deacetylase, HDAC9. Proc. Natl Acad. Sci. USA 98, 10572–10577 (2001).
Sparrow, D. B. et al. MEF-2 function is modified by a novel co-repressor, MITR. EMBO J. 18, 5085–5098 (1999).
Zhou, X., Richon, V. M., Rifkind, R. A. & Marks, P. A. Identification of a transcriptional repressor related to the noncatalytic domain of histone deacetylases 4 and 5. Proc. Natl Acad. Sci. USA 97, 1056–1061 (2000).
Lu, J., McKinsey, T. A., Nicol, R. L. & Olson, E. N. Signal-dependent activation of the MEF2 transcription factor by dissociation from histone deacetylases. Proc. Natl Acad. Sci. USA 97, 4070–4075 (2000).
McKinsey, T. A., Zhang, C. L., Lu, J. & Olson, E. N. Signal-dependent nuclear export of a histone deacetylase regulates muscle differentiation. Nature 408, 106–111 (2000). The authors show that the calcium-dependent export of an HDAC can control muscle differentiation.
Grozinger, C. M. & Schreiber, S. L. Regulation of histone deacetylase 4 and 5 and transcriptional activity by 14-3-3-dependent cellular localization. Proc. Natl Acad. Sci. USA 97, 7835–7840 (2000).
McKinsey, T. A., Zhang, C. L. & Olson, E. N. Activation of the myocyte enhancer factor-2 transcription factor by calcium/calmodulin-dependent protein kinase-stimulated binding of 14-3-3 to histone deacetylase 5. Proc. Natl Acad. Sci. USA 97, 14400–14405 (2000).
McKinsey, T. A., Zhang, C. L. & Olson, E. N. Identification of a signal-responsive nuclear export sequence in class II histone deacetylases. Mol. Cell. Biol. 21, 6312–6321 (2001).
McKinsey, T. A., Zhang, C. L. & Olson, E. N. MEF2: a calcium-dependent regulator of cell division, differentiation and death. Trends Biochem. Sci. 27, 40–47 (2002).
Zhang, C. L., McKinsey, T. A. & Olson, E. N. The transcriptional corepressor MITR is a signal-responsive inhibitor of myogenesis. Proc. Natl Acad. Sci. USA 98, 7354–7359 (2001).
Youn, H. D., Sun, L., Prywes, R. & Liu, J. O. Apoptosis of T cells mediated by Ca2+-induced release of the transcription factor MEF2. Science 286, 790–793 (1999).
Youn, H. D. & Liu, J. O. Cabin1 represses MEF2-dependent Nur77 expression and T cell apoptosis by controlling association of histone deacetylases and acetylases with MEF2. Immunity 13, 85–94 (2000).
Youn, H. D., Grozinger, C. M. & Liu, J. O. Calcium regulates transcriptional repression of myocyte enhancer factor 2 by histone deacetylase 4. J. Biol. Chem. 275, 22563–22567 (2000).
Sun, L. et al. Cabin 1, a negative regulator for calcineurin signaling in T lymphocytes. Immunity 8, 703–711 (1998).
Pfum, M. K. H., Tong, J. K., Lane, W. S. & Schreiber, S. L. Histone deacetylase 1 phosphorylation promotes enzymatic activity and complex formation. J. Biol. Chem. 276, 47733–47741 (2001).
David, G., Neptune, M. A. & DePinho, R. A. SUMO-1 modification of histone deacetylase 1 (HDAC1) modulates its biological activities. J. Biol. Chem. 277, 23658–23663 (2002).
Brosenitsch, T. A. & Katz, D. M. Physiological patterns of electrical stimulation can induce neuronal gene expression by activating N-type calcium channels. J. Neurosci. 21, 2571–2579 (2001).
Oertner, T. G. & Svoboda, K. Subliminal messages in hippocampal pyramidal neurons. J. Physiol. (Lond.) 543, 397 (2002).
Dudek, S. & Fields, R. D. Somatic action potentials are sufficient for late-phase LTP-related cell signaling. Proc. Natl Acad. Sci. USA 99, 3962–3967 (2002).
Impey, S. et al. Cross talk between ERK and PKA is required for Ca2+ stimulation of CREB-dependent transcription and ERK nuclear translocation. Neuron 21, 869–883 (1998).
Graef, I. A., Chen, F. & Crabtree, G. R. NFAT signaling in vertebrate development. Curr. Opin. Genet. Dev. 11, 505–512 (2001).
Acknowledgements
M.E.G. acknowledges the generous support of the F. M. Kirby Foundation to the Division of Neuroscience. This work was supported by a Mental Retardation Research Center grant and a National Institutes of Health grant to M.E.G., an American Cancer Society postdoctoral fellowship to A.E.W., and a Helen Hay Whitney postdoctoral fellowship to E.C.G. We thank P. Greer, A. Brunet, C. Cowan and J. Kornhauser for critical reading of the manuscript. We apologize to our colleagues whose original research could not be cited owing to space limitations.
Author information
Authors and Affiliations
Corresponding author
Related links
Related links
DATABASES
LocusLink
OMIM
FURTHER INFORMATION
Encyclopedia of Life Sciences
chromatin remodelling and histone modification in transcription regulation
protein phosphorylation and long-term synaptic plasticity
protein synthesis and long-term synaptic plasticity
Glossary
- CIRCADIAN RHYTHMS
-
Biological rhythms of physiology and behaviour that have a periodicity of 24 hours and are controlled by a clock mechanism in the suprachiasmatic nucleus of the brain.
- IMMEDIATE–EARLY GENES
-
Genes that are induced rapidly and transiently in the absence of de novo protein synthesis. Many immediate–early genes, such as c-Fos, control the transcription of other genes, and thereby provide the early stages in the control of the production of specific proteins.
- NEUROTROPHINS
-
A family of secreted molecules (BDNF, NGF, NT3 and NT4/5) that promote neuronal survival.
- DOMINANT NEGATIVE
-
Describes a mutant molecule that can form a heteromeric complex with the normal molecule, knocking out the activity of the entire complex.
- APOPTOSIS
-
A process of cell death that is characterized by the activation of a genetically encoded cell-suicide programme.
- LONG-TERM POTENTIATION
-
(LTP). An enduring increase in postsynaptic responsiveness as a result of high-frequency (tetanic) stimulation of presynaptic neurons. It is measured both as the amplitude of excitatory postsynaptic potentials and as the magnitude of postsynaptic-cell population spike. LTP is most often studied in the hippocampus and is often considered to be the cellular basis of learning and memory.
- LONG-TERM DEPRESSION
-
(LTD). An enduring weakening of synaptic strength that is thought to interact with long-term potentiation (LTP) in the cellular mechanisms of learning and memory in structures such as the hippocampus and cerebellum. Unlike LTP, which is produced by brief high-frequency stimulation, LTD can be produced by long-term, low-frequency stimulation.
- IQ DOMAIN
-
A small structural domain that mediates interactions with calmodulin. The motif only loosely defines the amino-acid sequence at 5 of 11 possible residues. Different IQ domains bind calmodulin at varying intracellular Ca2+ concentrations or independently of Ca2+.
- TRANSCRIPTIONAL COACTIVATOR
-
A protein that assists in the recruitment of RNA polymerase II to promoters by associating with transcription factors that are bound to sequence-specific DNA elements.
- KID AND KIX DOMAINS
-
The protein interaction domains of CREB and CBP. The kinase-inducible domain (KID) of CREB contains Ser133, which is phosphorylated in response to many extracellular stimuli. Phosphorylation at Ser133 recruits CBP through binding of its KIX domain.
- CYTOKINES
-
In general terms, cytokines are proteins made by cells that affect the behaviour of other cells. They are produced mainly by the immune system.
- PC12 CELLS
-
Neuron-like cells that are derived from a malignant neural crest tumour (a phaeochromocytoma).
- EF HAND
-
A Ca2+-binding domain that was originally identified in parvalbumin. Also known as the helix–loop–helix domain. The loop can accommodate Ca2+ by coordination through several amino acids in a pentagonal pyramid.
- PRODYNORPHIN
-
The preprocessed form of the peptide dynorphin, which mediates pain sensation by binding to the κ-opioid receptor.
- REPORTER GENE
-
A gene that encodes an easily assayed product. It is coupled to the upstream sequence of another gene, and can then be transfected into cells to identify factors that activate response elements in the upstream region of the gene of interest.
- STERIC HINDRANCE
-
The prevention of a reaction between different molecules as a result of their sizes or spatial disposition.
- MHC CLASS I GENES
-
Major histocompatibility complex (MHC) class I genes are required for the presentation of antigen to T cells in the immune system; their function in the immunologically privileged CNS is largely unknown.
- CHROMATIN
-
A complex of DNA, histones and non-histone proteins that is found in the nucleus of the eukaryotic cell.
- MASS SPECTROMETRY
-
A technique in which a compound is bombarded with an electron beam of sufficient energy to fragment the molecule. The cations that are produced are accelerated in a vacuum through a magnetic field, and sorted on the basis of mass-to-charge ratio. The ratio is roughly equivalent to the molecular weight of the fragment.
- CHROMATIN IMMUNOPRECIPITATION
-
A technique that is used to identify transcription factors bound to chromatin.
Rights and permissions
About this article
Cite this article
West, A., Griffith, E. & Greenberg, M. Regulation of transcription factors by neuronal activity. Nat Rev Neurosci 3, 921–931 (2002). https://doi.org/10.1038/nrn987
Issue Date:
DOI: https://doi.org/10.1038/nrn987
This article is cited by
-
Astaxanthin and DHA Supplementation Modulates the Maternal Undernutrition-induced Impairment of Cognitive Behavior and Synaptic Plasticity in Adult Life of Offspring’s -Exploring the Molecular Mechanism
Molecular Neurobiology (2024)
-
α7 Nicotinic acetylcholine receptor: a key receptor in the cholinergic anti-inflammatory pathway exerting an antidepressant effect
Journal of Neuroinflammation (2023)
-
Epigenetic regulation of beta-endorphin synthesis in hypothalamic arcuate nucleus neurons modulates neuropathic pain in a rodent pain model
Nature Communications (2023)
-
GASP1 enhances malignant phenotypes of breast cancer cells and decreases their response to paclitaxel by forming a vicious cycle with IGF1/IGF1R signaling pathway
Cell Death & Disease (2022)
-
Impaired ion homeostasis as a possible associate factor in mucopolysaccharidosis pathogenesis: transcriptomic, cellular and animal studies
Metabolic Brain Disease (2022)