Key Points
-
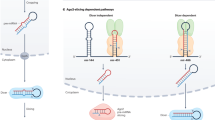
microRNAs (miRNAs) are ∼21 nucleotide transcripts that are predicted to regulate the translation of up to a third of the mRNAs in a cell. The miRNA biogenesis pathway begins with a primary miRNA transcript that is cleaved to an approximately 70 nucleotide precursor. This precursor is transported to the cytoplasm where the mature miRNA is generated by a second cleavage event and subsequently targeted to a specific mRNA in an RNA silencing complex.
-
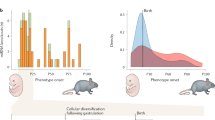
miRNAs are abundant in the nervous system where they have key roles in development, neural plasticity, and probably a host of other yet-to-be-discovered functions. In development, specific miRNAs have been implicated in patterning, cell specification and axonal pathfinding.
-
A set of miRNAs are turned on during stem cell differentiation when neural precursors become neurons. Among the properties of these miRNAs is the ability to suppress non-neural mRNAs broadly and to shift the fate of precursor cell populations between neurons and glia.
-
Accumulated data have strongly implicated miRNAs in the regulation of apoptosis and indicated that they are likely to contribute to widespread apoptosis that takes place during neural development.
-
Key support for miRNA functions related to neural plasticity include the involvement of miR-134 in targeting LIM-domain kinase 1 in the dendrite to regulate changes in spine size, and the degradation of the helicase, armitage, as a regulator of activity-dependent effects on translation at synapses.
-
Fragile X syndrome is one disease in which miRNA dysfunction is likely to contribute to disease pathology. miRNAs and miRNA-processing proteins are associated with the fragile X mental retardation protein. In addition, many forms of cancer, including glioblastoma, are associated with miRNA dysfunction.
Abstract
A class of small, non-coding transcripts called microRNAs (miRNAs) that provide a crucial and pervasive layer of post-transcriptional gene regulation has recently emerged and become the focus of intense research. miRNAs are abundant in the nervous system, where they have key roles in development and are likely to be important mediators of plasticity. A highly conserved pathway of miRNA biogenesis is closely linked to the transport and translatability of mRNAs in neurons. Although there are nearly 500 known human miRNA sequences, each of only ∼21 nucleotides, which bind to multiple mRNA targets, the accurate prediction of miRNA targets seems to lie just beyond our grasp. Nevertheless, the identification of such targets promises to provide new insights into many facets of neuronal function.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Lewis, B. P. et al. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 120, 15–20 (2005).
Krichevsky, A. M. et al. A microRNA array reveals extensive regulation of microRNAs during brain development. RNA 9, 1274–1281 (2003).
Miska, E. et al. Microarray analysis of microRNA expression in the developing mammalian brain. Genome Biol. 5, R68 (2004).
Sempere, L. F. et al. Expression profiling of mammalian microRNAs uncovers a subset of brain-expressed microRNAs with possible roles in murine and human neuronal differentiation. Genome Biol. 5, R13 (2004).
Kim, J. et al. Identification of many microRNAs that copurify with polyribosomes in mammalian neurons. Proc. Natl Acad. Sci. USA 101, 360–365 (2004).
Lagos-Quintana, M. et al. Identification of tissue-specific microRNA's from mouse. Curr. Biol. 12, 735–739 (2002).
Zeng, Y. & Cullen, B. R. Recognition and cleavage of primary microRNA transcripts. Methods Mol. Biol. 342, 49–56 (2006).
Zeng, Y., Yi, R. & Cullen, B. R. Recognition and cleavage of primary microRNA precursors by the nuclear processing enzyme Drosha. EMBO J. 24, 138–148 (2005).
Yi, R. et al. Exportin-5 mediates the nuclear export of pre-microRNAs and short hairpin RNAs. Genes Dev. 17, 3011–3016 (2003).
Lund, E. et al. Nuclear export of microRNA precursors. Science 303, 95–98 (2004).
Forstemann, K. et al. Normal microRNA maturation and germ-line stem cell maintenance requires Loquacious, a double-stranded RNA-binding domain protein. PLoS Biol. 3, e236 (2005).
Saito, K. et al. Processing of pre-microRNAs by the Dicer-1-Loquacious complex in Drosophila cells. PLoS Biol. 3, e235 (2005).
Bartel, D. P. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116, 281–297 (2004).
He, L. & Hannon, G. L. MicroRNAs: small RNAs with a big role in gene regulation. Nature Rev. Genet. 5, 522–531 (2004).
Hutvagner, G. & Zamore, P. D. RNAi: nature abhors a double-strand. Curr. Opin. Genet. Dev. 12, 225–232 (2002).
Lim, L. P. et al. Microarray analysis shows that some microRNAs downregulate large numbers of target mRNAs. Nature, 433, 769–773 (2005). An important study that suggested the global control of miRNAs over cell identity and the first to show a widespread effect of miRNAs on mRNA stability.
Bagga, S. et al. Regulation by let-7 and lin-4 miRNAs results in target mRNA degradation. Cell 122, 553–563 (2005).
Jing, Q. et al. Involvement of microRNA in AU-rich element-mediated mRNA instability. Cell 120, 623–634 (2005).
Sasaki, T. et al. Identification of eight members of the Argonaute family in the human genome small star, filled. Genomics 82, 323–330 (2003).
Liu, J. et al. Argonaute2 is the catalytic engine of mammalian RNAi. Science 305, 1437–1441 (2004).
Song, J. J. et al. Crystal structure of Argonaute and its implications for RISC slicer activity. Science 305, 1434–1437 (2004).
Okamura, K. et al. Distinct roles for Argonaute proteins in small RNA-directed RNA cleavage pathways. Genes Dev. 18, 1655–1666 (2004).
Caudy, A. A. et al. Fragile X-related protein and VIG associate with the RNA interference machinery. Genes Dev. 16, 2491–2496 (2002).
Caudy, A. A. et al. A micrococcal nuclease homologue in RNAi effector complexes. Nature 425, 411–414 (2003).
Liu, Q. et al. R2D2, a bridge between the initiation and effector steps of the Drosophila RNAi pathway. Science 301, 1921–1925 (2003).
Mourelatos, Z. et al. miRNPs: a novel class of ribonucleoproteins containing numerous microRNAs. Genes Dev. 16, 720–728 (2002).
Tomari, Y. et al. RISC assembly defects in the Drosophila RNAi mutant armitage. Cell 116, 831–841 (2004).
Chu, C. Y. & Rana, T. M. Translation repression in human cells by microRNA-induced gene silencing requires RCK/p54. PLoS Biol. 4, e210 (2006).
Liu, J. et al. A role for the P-body component GW182 in microRNA function. Nature Cell Biol. 7, 1261–1266 (2005).
Sen, G. L. & Blau, H. M. Argonaute 2/RISC resides in sites of mammalian mRNA decay known as cytoplasmic bodies. Nature Cell Biol. 7, 633–636 (2005).
Pauley, K. M. et al. Formation of GW bodies is a consequence of microRNA genesis. EMBO Rep. 7, 904–910 (2006).
Eystathioy, T. et al. Clinical and serological associations of autoantibodies to GW bodies and a novel cytoplasmic autoantigen GW182. J. Mol. Med. 81, 811–818 (2003).
Ding, L. et al. The developmental timing regulator AIN-1 interacts with miRISCs and may target the argonaute protein ALG-1 to cytoplasmic P bodies in C. elegans. Mol. Cell 19, 437–447 (2005).
Chalfie, M., Horvitz, H. R. & Sulston, J. E. Mutations that lead to reiterations in the cell lineages of C. elegans. Cell 24, 59–69 (1981).
Ambros, V. A hierarchy of regulatory genes controls a larva-to-adult developmental switch in C. elegans. Cell 57, 49–57 (1989).
Zhang, B., Pan, X. & Anderson, T. A. MicroRNA: a new player in stem cells. J. Cell Physiol. 209, 266–269 (2006).
Kataoka, Y., Takeichi, M. & Uemura, T. Developmental roles and molecular characterization of a Drosophila homologue of Arabidopsis Argonaute1, the founder of a novel gene superfamily. Genes Cells 6, 313–325 (2001).
Pearson, J. C., Lemons, D. & McGinnis, W. Modulating Hox gene functions during animal body patterning. Nature Rev. Genet. 6, 893–904 (2005).
Aboobaker, A. A. et al. Drosophila microRNAs exhibit diverse spatial expression patterns during embryonic development. Proc. Natl Acad. Sci. USA 102, 18017–18022 (2005).
Mansfield, J. H. et al. MicroRNA-responsive 'sensor' transgenes uncover Hox-like and other developmentally regulated patterns of vertebrate microRNA expression. Nature Genet. 36, 1079–1083 (2004). A highly novel method to detect miRNA expression patterns in the whole organism.
Ronshaugen, M. et al. The Drosophila microRNA iab-4 causes a dominant homeotic transformation of halteres to wings. Genes Dev. 19, 2947–2952 (2005).
Garzon, R. et al. MicroRNA fingerprints during human megakaryocytopoiesis. Proc. Natl Acad. Sci. USA 103, 5078–5083 (2006).
Yekta, S., Shih, I. H. & Bartel, D. P. MicroRNA-directed cleavage of HOXB8 mRNA. Science 304, 594–596 (2004). Deepened interest in an important feature of the HOX gene loci, specifically the relationship between the physical location of a gene and its function.
Greer, J. M. & M. R. Capecchi Hoxb8 is required for normal grooming behavior in mice. Neuron 33, 23–34 (2002).
Giraldez, A. J. et al. MicroRNAs regulate brain morphogenesis in zebrafish. Science 308, 833–838 (2005).
Wienholds, E. et al. MicroRNA expression in zebrafish embryonic development. Science 309, 310–311 (2005).
Leaman, D. et al. Antisense-mediated depletion reveals essential and specific functions of microRNAs in Drosophila development. Cell 121, 1097–1108 (2005).
Bernstein, E. et al. Dicer is essential for mouse development. Nature Genet. 35, 215–217 (2003).
Johnston, R. J. & Hobert, O. A microRNA controlling left/right neuronal asymmetry in Caenorhabditis elegans. Nature 426, 845–849 (2003).
Johnston, R. J. Jr et al. MicroRNAs acting in a double-negative feedback loop to control a neuronal cell fate decision. Proc. Natl Acad. Sci. USA 102 12449–12454 (2005).
Chang, S. et al. MicroRNAs act sequentially and asymmetrically to control chemosensory laterality in the nematode. Nature 430, 785–789 (2004). The above three studies from the Hobert lab are among the clearest demonstrations of miRNA circuitry and the complex relationship between miRNAs and transcription factors.
Wheeler, G. et al. Identification of new central nervous system specific mouse microRNAs. FEBS Lett. 580, 2195–2200 (2006).
Monticelli, S. et al. MicroRNA profiling of the murine hematopoietic system. Genome Biol. 6, R71 (2005).
Krichevsky, A. M. et al. Specific microRNAs modulate embryonic stem cell-derived neurogenesis. Stem Cells 24, 857–864 (2006).
Smirnova, L. et al. Regulation of miRNA expression during neural cell specification. Eur. J. Neurosci. 21, 1469–1477 (2006).
Lai, E. C., Burks, C. & Posakony, J. W. The K box, a conserved 3´ UTR sequence motif, negatively regulates accumulation of enhancer of split complex transcripts. Development 125, 4077–4088 (1998).
Naguibneva, I. et al. The microRNA miR-181 targets the homeobox protein Hox-A11 during mammalian myoblast differentiation. Nature Cell Biol. 8, 278–284 (2006).
Desai, A. R. & McConnell, S. K. Progressive restriction in fate potential by neural progenitors during cerebral cortical development. Development 127, 2863–2872 (2000).
Blackshaw, S. et al. Comprehensive analysis of photoreceptor gene expression and the identification of candidate retinal disease genes. Cell 107, 579–589 (2001).
Lin, S. Y. et al. The C. elegans hunchback homolog, hbl-1, controls temporal patterning and is a probable microRNA target. Dev. Cell 4, 639–650 (2003).
Irish, V., Lehmann, R. & Akam, M. The Drosophila posterior-group gene nanos functions by repressing hunchback activity. Nature 338, 646–648 (1989).
Pearson, B. J. & Doe, C. Q. Specification of temporal identity in the developing nervous system. Annu. Rev. Cell Dev. Biol. 20, 619–647 (2004).
Cleary, M. D. & Doe, C. Q. Regulation of neuroblast competence: multiple temporal identity factors specify distinct neuronal fates within a single early competence window. Genes Dev. 20, 429–434 (2006).
Hengst, U. et al. Functional and selective RNA interference in developing axons and growth cones. J. Neurosci. 26, 5727–5732 (2006).
Campbell, D. S. & Holt, C. E. Chemotropic responses of retinal growth cones mediated by rapid local protein synthesis and degradation. Neuron 32, 1013–1026 (2001).
Wu, K. Y. et al. Local translation of RhoA regulates growth cone collapse. Nature 436, 1020–1024 (2005).
Deglincerti, A., Hengst, U. & Jaffrey, S. R. Regulation of local protein translation in axons and growth cones by microRNAs. Soc. Neurosci. Abstr. 617. 9/B5 (2006).
Buss, R. R. & Oppenheim, R. W. Role of programmed cell death in normal neuronal development and function. Anat. Sci. Int. 79, 191–197 (2004).
Chan, J. A., Krichevsky, A. M. & Kosik, K. S. MicroRNA-21 is an antiapoptotic factor in human glioblastoma cells. Cancer Res. 65, 6029–6033 (2005).
Harfe, B. D. et al. The RNaseIII enzyme Dicer is required for morphogenesis but not patterning of the vertebrate limb. Proc. Natl Acad. Sci. USA 102, 10898–10903 (2005).
Martin, K. C. & Kosik, K. S. Synaptic tagging — who's it? Nature Rev. Neurosci. 3, 813–820 (2002).
Frey, U. & Morris, R. G. Synaptic tagging and long-term potentiation. Nature 385, 533–536 (1997).
Nguyen, P. V. & Kandel, E. R. Brief theta-burst stimulation induces a transcription-dependent late phase of LTP requiring cAMP in area CA1 of the mouse hippocampus. Learn. Mem. 4, 230–243 (1997).
Montarolo, P. G. et al. A critical period for macromolecular synthesis in long-term heterosynaptic facilitation in Aplysia. Science 234, 1249–1254 (1986).
Davis, H. P. & Squire, L. R. Protein synthesis and memory: a review. Psychol. Bull. 96, 518–559 (1984).
Aakalu, G. et al. Dynamic visualization of local protein synthesis in hippocampal neurons. Neuron 30, 489–502 (2001). Set up the rationale for performing experiments that sought a role for miRNAs in local dendritic translation.
Bagni, C. et al. Chemical stimulation of synaptosomes modulates αCa2+/calmodulin-dependent protein kinase II mRNA association to polysomes. J. Neurosci. 20, RC76 (2000).
Weiler, I. J. et al. Fragile X mental retardation protein is translated near synapses in response to neurotransmitter activation. Proc. Natl Acad. Sci. USA 94, 5395–5400 (1997).
Kang, H. & Schuman, E. M. A requirement for local protein synthesis in neurotrophin-induced hippocampal synaptic plasticity. Science 273, 1402–1406 (1996).
Huber, K. M., Roder, J. C. & Bear, M. F. Chemical induction of mGluR5- and protein synthesis--dependent long-term depression in hippocampal area CA1. J. Neurophysiol. 86, 321–325 (2001).
Huber, K. M., Kayser, M. S. & Bear, M. F. Role for rapid dendritic protein synthesis in hippocampal mGluR-dependent long-term depression. Science 288, 1254–1257 (2000).
Schratt, G. M. et al. A brain-specific microRNA regulates dendritic spine development. Nature 439, 283–289 (2006). The strongest suggestion so far that miRNAs are important regulators of plasticity.
Olsen, P. H. & Ambros, V. The lin-4 regulatory RNA controls developmental timing in Caenorhabditis elegans by blocking LIN-14 protein synthesis after the initiation of translation. Dev. Biol. 216, 671–680 (1999).
Krichevsky, A. M. & Kosik, K. S. Neuronal RNA granules: a link between RNA localization and stimulation-dependent translation. Neuron 32, 683–696 (2001).
Pinkstaff, J. K. et al. Internal initiation of translation of five dendritically localized neuronal mRNAs. Proc. Natl Acad. Sci. USA 98, 2770–2775 (2001).
Ashraf, S. I. et al. Synaptic protein synthesis associated with memory is regulated by the RISC pathway in Drosophila. Cell 124, 191–205 (2006). A paper that will grow in importance not only for its insights regarding miRNA regulation as a function of synaptic activity, but the potential of the approach to reveal a molecular-level image of brain activity.
Steward, O. & Schuman, E. M. Protein synthesis at synaptic sites on dendrites. Annu. Rev. Neurosci. 24, 299–325 (2001).
Zhong, J., Zhang, T. & Bloch, L. M. Dendritic mRNAs encode diversified functionalities in hippocampal pyramidal neurons. BMC Neurosci. 7, 17 (2006).
Glanzer, J. G. & Eberwine, J. H. Mechanisms of translational control in dendrites. Neurobiol. Aging 24, 1105–1111 (2003).
Bockers, T. M. et al. Differential expression and dendritic transcript localization of Shank family members: identification of a dendritic targeting element in the 3′ untranslated region of Shank1 mRNA. Mol. Cell. Neurosci. 26, 182–190 (2004).
Blichenberg, A. et al. Identification of a cis-acting dendritic targeting element in MAP2 mRNAs. J. Neurosci. 19, 8818–8829 (1999).
Mayford, M. et al. The 3′-untranslated region of CaMKIIa is a cis-acting signal for the localization and translation of mRNA in dendrites. Proc. Natl Acad. Sci. USA 93, 13250–13255 (1996).
Kindler, S. et al. Molecular characterization of dendritically localized transcripts encoding MAP2. Brain Res. Mol. Brain Res. 36, 63–69 (1996).
Mori, Y. et al. Two cis-acting elements in the 3′ untranslated region of α-CaMKII regulate its dendritic targeting. Nature Neurosci. 3, 1079–1084 (2000).
Lugli, G. et al. Dicer and eIF2c are enriched at postsynaptic densities in adult mouse brain and are modified by neuronal activity in a calpain-dependent manner. J. Neurochem. 94, 896–905 (2005).
Steward, O. & Worley, P. F. Selective targeting of newly synthesized arc mRNA to active synapses requires NMDA receptor activation. Neuron 30, 227–240 (2001).
Steward, O. et al. Synaptic activation causes the mRNA for the IEG Arc to localize selectively near activated postsynaptic sites on dendrites. Neuron 21, 741–751 (1998).
Jin, P. et al. Biochemical and genetic interaction between the fragile X mental retardation protein and the microRNA pathway. Nature Neurosci. 7, 113–117 (1998).
Vo, N. et al. A cAMP-response element binding protein-induced microRNA regulates neuronal morphogenesis. Proc. Natl Acad. Sci. USA 102, 16426–16431 (2005).
Impey, S. et al. Stimulation of cAMP response element (CRE)-mediated transcription during contextual learning. Nature Neurosci. 1, 595–601 (1998).
Antar, L. N. & Bassell, G. J. Sunrise at the synapse: the FMRP mRNP shaping the synaptic interface. Neuron 37, 555–558 (2003).
Esquela-Kerscher, A. & Slack, F. J. Oncomirs — microRNAs with a role in cancer. Nature Rev. Cancer 6, 259–269 (2006). An excellent overview, among the several that have been written, on the emerging relationship between miRNAs and cancer.
Zhang, L. et al. microRNAs exhibit high frequency genomic alterations in human cancer. Proc. Natl Acad. Sci. USA 103, 9136–9141 (2006).
Clop, A. et al. A mutation creating a potential illegitimate microRNA target site in the myostatin gene affects muscularity in sheep. Nature Genet. 38, 813–818 (2006). This paper is the leading edge of what will emerge as a windfall of genetic data related to polymorphisms at miRNA target sites with functional consequences.
Abelson, J. F. et al. Sequence variants in SLITRK1 are associated with Tourette's syndrome. Science 310, 317–320 (2005). Presents a novel and provocative basis for a poorly understood syndrome.
Lim, L. P. et al. The microRNAs of Caenorhabditis elegans. Genes Dev. 17, 991–1008 (2003).
Conaco, C. et al. Reciprocal actions of REST and a microRNA promote neuronal identity. Proc. Natl Acad. Sci. USA 103, 2422–2427 (2006). Opens the way to a network approach to the acquisition and maintenance of cell identity.
Grimm, D. et al. Fatality in mice due to oversaturation of cellular microRNA/short hairpin RNA pathways. Nature 441, 537–541 (2006). An important cautionary note.
Acknowledgements
Thanks to the members of the Kosik lab for insightful discussions and particularly to S. Banerjee and I. Wang for additional comments on the text. Also thanks to M. McManus for comments on the sensor technique.
Author information
Authors and Affiliations
Ethics declarations
Competing interests
The author declares no competing financial interests.
Related links
Related links
DATABASES
OMIM
FURTHER INFORMATION
Glossary
- Patterning
-
Establishing, often by diffusion gradients, a distribution of gene expression over a spatial region that defines the region's emerging anatomical organization, and could prefigure tissue differentiation.
- Plasticity
-
Changes that occur in the organization of the brain, either at the macroscopic level with anatomical reorganization, or at the molecular and physiological level, when synapses change their firing thresholds with repetitive stimulation.
- Pre-miRNA
-
The 70–100 nucleotide hairpin RNA processed from the primary microRNA in the nucleus by the double-stranded RNA-specific ribonuclease Drosha.
- Pri-miRNA
-
The initial transcript encoding the microRNA.
- Guide strand
-
The microRNA strand that is incorporated into the RNA-induced silencing complex and forms base pairs with the mRNA strand as a mature microRNA.
- Passenger strand
-
The strand of the microRNA duplex which is removed during the assembly of the RNA-induced silencing complex and which is usually degraded.
- Seed region
-
Positions 2–8 of the 5′ end of the microRNA that are most likely to form an exact match with an mRNA target and are often conserved.
- PAZ motif
-
A conserved nucleic acid-binding structure that is found in members of the DCR and AGO protein families.
- PIWI motif
-
A conserved structure that is found in members of the AGO protein family. It is structurally similar to ribonuclease H domains and, in at least some cases, has endoribonuclease activity.
- Slicer
-
Target RNA cleavage activity in RNA interference, mediated by argonaute 2.
- P bodies
-
Cytoplasmic structures that contain untranslated mRNAs and can serve as sites of mRNA degradation and storage.
- Homeotic phenotype
-
Arises due to mutations in homeotic genes that do not eliminate elements of the pattern but rather cause these elements to develop with inappropriate identities.
- Maternal–zygotic mutants
-
In zebrafish, maternally derived products are synthesized during oogenesis and accumulate to regulate early embryonic development. Maternal–zygotic mutants allow for systematic analysis of the maternal effects of known zygotic lethal mutations.
- Neurulation
-
The formation of the embryonic neural plate and its transformation into the neural tube.
- Neurocoel
-
The cavity or system of cavities in the interior of the vertebrate CNS comprising the central canal of the spinal cord and the ventricles of the brain.
- Rhombomere
-
A segment or swelling in the vertebrate embryo that is part of the developing rhombencephalon, which will become the pons, cerebellum and medulla. The fate of a rhombomere is affected by differential expression of Hox genes.
- 2′O-methyl antisense oligoribonucleotides
-
These bind to target microRNAs to block their function by specifically inactivating the RNA interference activity associated with miRNA–protein complexes, and are stable and specific over a wide range of concentrations.
- Maternal to zygotic transition
-
In early embryos, initial protein synthesis occurs primarily through maternal mRNAs that are synthesized during oogenesis in the mother and stored in the oocyte. Eventually the embryo undergoes a transition and transcribes its own genes.
- Growth cones
-
A dynamic, actin-supported extension of a developing axon seeking its synaptic target.
- Endochondrial ossification
-
Involves the conversion of a type of cartilage called hyaline cartilage to bone. Hyaline cartilage is formed into a likeness of the future bone by chondroblast cells.
- Polysome
-
A functional unit of protein synthesis that consists of several ribosomes that are attached along the length of a single molecule of mRNA. It is the shortened form of the term polyribosome.
- Internal initiation
-
Translation initiation occurs by an internal ribosomal-binding mechanism in contrast to ribosomal binding at the 5′ end of the mRNA, facilitated by an interaction between the methylated cap structure at the end of the mRNA and the cap-binding protein complex eIF–4F.
- Glomeruli
-
Anatomical substructure of the Drosophila antennal lobe.
- Interactome
-
The network of protein interactions.
- Transcriptome
-
Historically the set of all mRNAs produced in a cell or a population of cells. However, more recently it has been suggested that the term must stipulate whether non-coding RNAs are included or not.
Rights and permissions
About this article
Cite this article
Kosik, K. The neuronal microRNA system. Nat Rev Neurosci 7, 911–920 (2006). https://doi.org/10.1038/nrn2037
Issue Date:
DOI: https://doi.org/10.1038/nrn2037
This article is cited by
-
Death Induced by Survival gene Elimination (DISE) correlates with neurotoxicity in Alzheimer’s disease and aging
Nature Communications (2024)
-
Brain-Derived Exosomal miRNA Profiles upon Experimental SAE Rats and Their Comparison with Peripheral Exosomes
Molecular Neurobiology (2024)
-
RNA methyltransferase NSun2 deficiency promotes neurodegeneration through epitranscriptomic regulation of tau phosphorylation
Acta Neuropathologica (2023)
-
Spinal cord injury: a study protocol for a systematic review and meta-analysis of microRNA alterations
Systematic Reviews (2022)
-
Dexmedetomidine attenuates oxygen-glucose deprivation/ reperfusion-induced inflammation through the miR-17-5p/ TLR4/ NF-κB axis
BMC Anesthesiology (2022)