Abstract
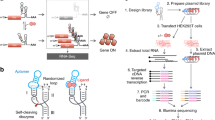
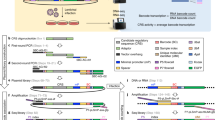
Engineered zinc-finger transcription factors (ZF-TF) are powerful tools to modulate the expression of specific genes. Complex libraries of ZF-TF can be delivered into cells to scan the genome for genes responsible for a particular phenotype or to select the most effective ZF-TF to regulate an individual gene. In both cases, the construction of highly representative and unbiased libraries is critical. In this protocol, we describe a user-friendly ZF technology suitable for the creation of complex libraries and the construction of customized ZF-TFs. The new technology described here simplifies the building of ZF libraries, avoids PCR-introduced bias and ensures equal representation of every module. We also describe the construction of a customized ZF-TF that can be transferred to a number of expression vectors. This protocol can be completed in 9–11 d.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Elrod-Erickson, M., Rould, M.A., Nekludova, L. & Pabo, C.O. Zif268 protein-DNA complex refined at 1.6 A: a model system for understanding zinc finger-DNA interactions. Structure 4, 1171–1180 (1996).
Pavletich, N.P. & Pabo, C.O. Zinc finger-DNA recognition: crystal structure of a Zif268-DNA complex at 2.1 A. Science 252, 809–817 (1991).
Segal, D.J., Dreier, B., Beerli, R.R. & Barbas III, C.F. Toward controlling gene expression at will: selection and design of zinc finger domains recognizing each of the 5′-GNN-3′ DNA target sequences. Proc. Natl. Acad. Sci. USA 96, 2758–2763 (1999).
Dreier, B., Beerli, R.R., Segal, D.J., Flippin, J.D. & Barbas III, C.F. Development of zinc finger domains for recognition of the 5′-ANN-3′ family of DNA sequences and their use in the construction of artificial transcription factors. J. Biol. Chem. 276, 29466–29478 (2001).
Dreier, B. et al. Development of zinc finger domains for recognition of the 5′-CNN-3′ family DNA sequences and their use in the construction of artificial transcription factors. J. Biol. Chem. 280, 35588–35597 (2005).
Jamieson, A.C. et al. Controlling gene expression in Drosophila using engineered zinc finger protein transcription factors. Biochem. Biophys. Res. Commun. 348, 873–879 (2006).
Snowden, A.W. et al. Repression of vascular endothelial growth factor A in glioblastoma cells using engineered zinc finger transcription factors. Cancer Res. 63, 8968–8976 (2003).
Tan, S. et al. Zinc-finger protein-targeted gene regulation: genomewide single-gene specificity. Proc. Natl. Acad. Sci. USA 100, 11997–12002 (2003).
Bartsevich, V.V., Miller, J.C., Case, C.C. & Pabo, C.O. Engineered zinc finger proteins for controlling stem cell fate. Stem Cells 21, 632–637 (2003).
Beerli, R.R., Segal, D.J., Dreier, B. & Barbas III, C.F. Toward controlling gene expression at will: specific regulation of the erbB-2/HER-2 promoter by using polydactyl zinc finger proteins constructed from modular building blocks. Proc. Natl. Acad. Sci. USA 95, 14628–14633 (1998).
Beerli, R.R., Dreier, B. & Barbas III, C.F. Positive and negative regulation of endogenous genes by designed transcription factors. Proc. Natl. Acad. Sci. USA 97, 1495–1500 (2000).
Falke, D., Fisher, M., Ye, D. & Juliano, R.L. Design of artificial transcription factors to selectively regulate the pro-apoptotic bax gene. Nucleic Acids Res. 31, e10 (2003).
Falke, D., Fisher, M.H. & Juliano, R.L. Selective transcription of p53 target genes by zinc finger-p53 DNA binding domain chimeras. Biochim. Biophys. Acta. 1681, 15–27 (2004).
Xu, D., Ye, D., Fisher, M. & Juliano, R.L. Selective inhibition of P-glycoprotein expression in multidrug-resistant tumor cells by a designed transcriptional regulator. J. Pharmacol. Exp. Ther. 302, 963–971 (2002).
Rebar, E.J. et al. Induction of angiogenesis in a mouse model using engineered transcription factors. Nat. Med. 8, 1427–1432 (2002).
Liu, P.Q. et al. Regulation of an endogenous locus using a panel of designed zinc finger proteins targeted to accessible chromatin regions. Activation of vascular endothelial growth factor A. J. Biol. Chem. 276, 11323–11334 (2001).
Gommans, W.M. et al. Engineering zinc finger protein transcription factors to downregulate the epithelial glycoprotein-2 promoter as a novel anti-cancer treatment. Mol. Carcinog. 46, 391–401 (2007).
Nomura, W. & Barbas III, C.F. In vivo site-specific DNA methylation with a designed sequence-enabled DNA methylase. J. Am. Chem. Soc. 129, 8676–8677 (2007).
Smith, A.E. & Ford, K.G. Specific targeting of cytosine methylation to DNA sequences in vivo . Nucleic Acids Res. 35, 740–754 (2007).
Minczuk, M., Papworth, M.A., Kolasinska, P., Murphy, M.P. & Klug, A. Sequence-specific modification of mitochondrial DNA using a chimeric zinc finger methylase. Proc. Natl. Acad. Sci. USA 103, 19689–19694 (2006).
Snowden, A.W., Gregory, P.D., Case, C.C. & Pabo, C.O. Gene-specific targeting of H3K9 methylation is sufficient for initiating repression in vivo . Curr. Biol. 12, 2159–2166 (2002).
Gordley, R.M., Smith, J.D., Graslund, T. & Barbas III, C.F. Evolution of programmable zinc finger-recombinases with activity in human cells. J. Mol. Biol. 367, 802–813 (2007).
Akopian, A., He, J., Boocock, M.R. & Stark, W.M. Chimeric recombinases with designed DNA sequence recognition. Proc. Natl. Acad. Sci. USA 100, 8688–8691 (2003).
Yant, S.R., Huang, Y., Akache, B. & Kay, M.A. Site-directed transposon integration in human cells. Nucleic Acids Res. 35, e50 (2007).
Tan, W., Dong, Z., Wilkinson, T.A., Barbas III, C.F. & Chow, S.A. Human immunodeficiency virus type 1 incorporated with fusion proteins consisting of integrase and the designed polydactyl zinc finger protein E2C can bias integration of viral DNA into a predetermined chromosomal region in human cells. J. Virol. 80, 1939–1948 (2006).
Kim, Y.G., Cha, J. & Chandrasegaran, S. Hybrid restriction enzymes: zinc finger fusions to Fok I cleavage domain. Proc. Natl. Acad. Sci. USA 93, 1156–1160 (1996).
Bibikova, M. et al. Stimulation of homologous recombination through targeted cleavage by chimeric nucleases. Mol. Cell Biol. 21, 289–297 (2001).
Bibikova, M., Golic, M., Golic, K.G. & Carroll, D. Targeted chromosomal cleavage and mutagenesis in Drosophila using zinc-finger nucleases. Genetics 161, 1169–1175 (2002).
Bibikova, M., Beumer, K., Trautman, J.K. & Carroll, D. Enhancing gene targeting with designed zinc finger nucleases. Science 300, 764 (2003).
Porteus, M.H. & Baltimore, D. Chimeric nucleases stimulate gene targeting in human cells. Science 300, 763 (2003).
Porteus, M.H. Mammalian gene targeting with designed zinc finger nucleases. Mol. Ther. 13, 438–446 (2006).
Lloyd, A., Plaisier, C.L., Carroll, D. & Drews, G.N. Targeted mutagenesis using zinc-finger nucleases in Arabidopsis. Proc. Natl. Acad. Sci. USA 102, 2232–2237 (2005).
Wright, D.A. et al. High-frequency homologous recombination in plants mediated by zinc-finger nucleases. Plant J. 44, 693–705 (2005).
Urnov, F.D. et al. Highly efficient endogenous human gene correction using designed zinc-finger nucleases. Nature 435, 646–651 (2005).
Alwin, S. et al. Custom zinc-finger nucleases for use in human cells. Mol. Ther. 12, 610–617 (2005).
Moehle, E.A. et al. Targeted gene addition into a specified location in the human genome using designed zinc finger nucleases. Proc. Natl. Acad. Sci. USA 104, 3055–3060 (2007).
Beerli, R.R., Schopfer, U., Dreier, B. & Barbas III, C.F. Chemically regulated zinc finger transcription factors. J. Biol. Chem. 275, 32617–32627 (2000).
Ghosh, I., Stains, C.I., Ooi, A.T. & Segal, D.J. Direct detection of double-stranded DNA: Molecular methods and applications for DNA diagnostics. Mol. Biosyst. 2, 551–560 (2006).
Ooi, A.T., Stains, C.I., Ghosh, I. & Segal, D.J. Sequence-enabled reassembly of beta-lactamase (SEER-LAC): a sensitive method for the detection of double-stranded DNA. Biochemistry 45, 3620–3625 (2006).
Stains, C.I., Furman, J.L., Segal, D.J. & Ghosh, I. Site-specific detection of DNA methylation utilizing mCpG-SEER. J. Am. Chem. Soc. 128, 9761–9765 (2006).
Liu, Q., Segal, D.J., Ghiara, J.B. & Barbas III, C.F. Design of polydactyl zinc-finger proteins for unique addressing within complex genomes. Proc. Natl. Acad. Sci. USA 94, 5525–5530 (1997).
Guan, X. et al. Heritable endogenous gene regulation in plants with designed polydactyl zinc finger transcription factors. Proc. Natl. Acad. Sci. USA 99, 13296–13301 (2002).
Segal, D.J., Crotty, J.W., Bhakta, M.S., Barbas III, C.F. & Horton, N.C. Structure of Aart, a designed six-finger zinc finger peptide, bound to DNA. J. Mol. Biol. 363, 405–421 (2006).
Blancafort, P., Magnenat, L. & Barbas III, C.F. Scanning the human genome with combinatorial transcription factor libraries. Nat. Biotechnol. 21, 269–274 (2003).
Magnenat, L., Blancafort, P. & Barbas III, C.F. In vivo selection of combinatorial libraries and designed affinity maturation of polydactyl zinc finger transcription factors for ICAM-1 provides new insights into gene regulation. J. Mol. Biol. 341, 635–649 (2004).
Lee, D.K., Kim, Y.H., Kim, J.S. & Seol, W. Induction and characterization of taxol-resistance phenotypes with a transiently expressed artificial transcriptional activator library. Nucleic Acids Res. 32, e116 (2004).
Zhao, X.H., Zhu, X.D., Liu, J., Rao, X.J. & Huang, P.T. Construction of a SV40 promoter specific artificial transcription factor. Sheng Wu Gong Cheng Xue Bao 19, 608–612 (2003).
Blancafort, P. et al. Genetic reprogramming of tumor cells by zinc finger transcription factors. Proc. Natl. Acad. Sci. USA 102, 11716–11721 (2005).
Lund, C.V., Blancafort, P., Popkov, M. & Barbas III, C.F. Promoter-targeted phage display selections with preassembled synthetic zinc finger libraries for endogenous gene regulation. J. Mol. Biol. 340, 599–613 (2004).
Lindhout, B.I., Pinas, J.E., Hooykaas, P.J. & van der Zaal, B.J. Employing libraries of zinc finger artificial transcription factors to screen for homologous recombination mutants in Arabidopsis. Plant J. 48, 475–483 (2006).
Park, K.S. et al. Phenotypic alteration of eukaryotic cells using randomized libraries of artificial transcription factors. Nat. Biotechnol. 21, 1208–1214 (2003).
Park, K.S., Jang, Y.S., Lee, H. & Kim, J.S. Phenotypic alteration and target gene identification using combinatorial libraries of zinc finger proteins in prokaryotic cells. J. Bacteriol. 187, 5496–5499 (2005).
Blancafort, P. et al. Modulation of drug resistance by artificial transcription factors. Mol. Cancer Ther. 7, 688–697 (2008).
Tschulena, U., Peterson, K.R., Gonzalez, B., Fedosyuk, H. & Barbas III, C.F. Positive selection of DNA-protein interactions in mammalian cells through phenotypic coupling with retrovirus production. Nat. Struct. Mol. Biol. 16, 1195–1199 (2009).
Meng, X., Thibodeau-Beganny, S., Jiang, T., Joung, J.K. & Wolfe, S.A. Profiling the DNA-binding specificities of engineered Cys2His2 zinc finger domains using a rapid cell-based method. Nucleic Acids Res. 35, e81 (2007).
Durai, S., Bosley, A., Abulencia, A.B., Chandrasegaran, S. & Ostermeier, M. A bacterial one-hybrid selection system for interrogating zinc finger-DNA interactions. Comb. Chem. High Throughput Screen 9, 301–311 (2006).
Bae, K.H. & Kim, J.S. One-step selection of artificial transcription factors using an in vivo screening system. Mol. Cells 21, 376–380 (2006).
Bartsevich, V.V. & Juliano, R.L. Regulation of the MDR1 gene by transcriptional repressors selected using peptide combinatorial libraries. Mol. Pharmacol. 58, 1–10 (2000).
Ihara, H. et al. In vitro selection of zinc finger DNA-binding proteins through ribosome display. Biochem. Biophys. Res. Commun. 345, 1149–1154 (2006).
Sepp, A. & Choo, Y. Cell-free selection of zinc finger DNA-binding proteins using in vitro compartmentalization. J. Mol. Biol. 354, 212–219 (2005).
Carroll, D., Morton, J.J., Beumer, K.J. & Segal, D.J. Design, construction and in vitro testing of zinc finger nucleases. Nat. Protoc. 1, 1329–1341 (2006).
Wright, D.A. et al. Standardized reagents and protocols for engineering zinc finger nucleases by modular assembly. Nat. Protoc. 1, 1637–1652 (2006).
Maeder, M.L. et al. Rapid 'open-source' engineering of customized zinc-finger nucleases for highly efficient gene modification. Mol. Cell 31, 294–301 (2008).
Desjarlais, J.R. & Berg, J.M. Use of a zinc-finger consensus sequence framework and specificity rules to design specific DNA binding proteins. Proc. Natl. Acad. Sci. USA 90, 2256–2260 (1993).
Mandell, J.G. & Barbas III, C.F. Zinc finger tools: custom DNA-binding domains for transcription factors and nucleases. Nucleic Acids Res. 34, W516–W523 (2006).
Gordley, R.M., Gersbach, C.A. & Barbas III, C.F. Synthesis of programmable integrases. Proc. Natl. Acad. Sci. USA 106, 5053–5058 (2009).
Liu, X. et al. Transforming growth factor beta-induced phosphorylation of Smad3 is required for growth inhibition and transcriptional induction in epithelial cells. Proc. Natl. Acad. Sci. USA 94, 10669–10674 (1997).
Sidhu, S.S., Lowman, H.B., Cunningham, B.C. & Wells, J.A. Phage display for selection of novel binding peptides. Methods Enzymol. 328, 333–363 (2000).
Barbas III, C.F., Burton, D.R., Scott, J.K. & Silverman, G.J. Phage Display: A Laboratory Manual (Cold Spring Harbor Laboratory Press, 2001).
Acknowledgements
We thank S. Juraja and S. Alonso for critical reading of the manuscript and members of our group for helpful suggestions. L.J.S. is supported by The American Cancer Society Illinois Division—Linda M. Campbell Postdoctoral Fellowship. Funding was provided by grants from the US National Institutes of Health.
Author information
Authors and Affiliations
Contributions
B.G. and L.J.S. contributed equally to this work. C.F.B. III conceived of the SuperZif library construction concept and directed the research. Y.Y., B.G. and C.F.B. III designed the SuperZif vectors. R.P.F. designed and constructed the pSCV vector, modified the SuperZiFCNN vector and aided in library construction. B.G., L.J.S. and L.A. constructed and tested the libraries. The paper was written by L.J.S. with assistance from B.G., C.F.B. III and R.P.F.
Note: Supplementary information is available via the HTML version of this article.
Corresponding author
Supplementary information
Supplementary Information
Supplementary Sequence Archive (DOC 323 kb)
Supplementary Figure 1
Vector maps of SuperZiF vectors and pSCV. (PDF 262 kb)
Supplementary Figure 2
Vector maps of expression vectors in Table 1. (PDF 597 kb)
Rights and permissions
About this article
Cite this article
Gonzalez, B., Schwimmer, L., Fuller, R. et al. Modular system for the construction of zinc-finger libraries and proteins. Nat Protoc 5, 791–810 (2010). https://doi.org/10.1038/nprot.2010.34
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2010.34
This article is cited by
-
Induction of apoptosis in imatinib sensitive and resistant chronic myeloid leukemia cells by efficient disruption of bcr-abl oncogene with zinc finger nucleases
Journal of Experimental & Clinical Cancer Research (2018)
-
CRISPR/Cas9 genome editing technology significantly accelerated herpes simplex virus research
Cancer Gene Therapy (2018)
-
Genome-editing Technologies for Gene and Cell Therapy
Molecular Therapy (2016)
-
Histone methyltransferase Ash1L mediates activity-dependent repression of neurexin-1α
Scientific Reports (2016)
-
Efficient delivery of nuclease proteins for genome editing in human stem cells and primary cells
Nature Protocols (2015)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.