Abstract
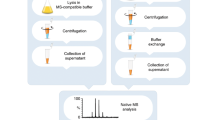
The analysis of highly hydrophobic proteins is still an analytical challenge. Using a recombinant gamma-aminobutyric acid A (GABAA)-receptor subunit as a model protein, we developed a gel-based proteomic approach for high MS/MS-peptide sequence coverage identification. Protein samples were separated by multi-dimensional gel electrophoresis and the three protein spots representing the GABAA-receptor subunit α-1 from the last electrophoretic step were used for in-gel digestion with trypsin, chymotrypsin and subtilisin, followed by subsequent mass-spectrometric identification by nano-ESI-LC-MS/MS Qstar XL (quadrupole time-of-flight (qQTOF)) and linear ion trap (LIT) LTQ XL identification. This protocol allows the unambiguous identification of the GABAA-receptor α-1 subunit protein with 100% sequence coverage, thus covering all four hydrophobic transmembrane domains. This protocol differs from other methods in the selection of enzymes, digestion conditions and use of the two mass spectrometry principles. The protocol takes ∼10 d to complete and may represent a step forward in the complex analysis of other membrane or hydrophobic proteins.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Kashino, Y. Separation methods in the analysis of protein membrane complexes. J. Chromatogr. B. Analyt. Technol. Biomed. Life Sci. 797, 191–216 (2003).
Speers, A.E. & Wu, C.C. Proteomics of integral membrane proteins–theory and application. Chem. Rev. 107, 3687–3714 (2007).
Rabilloud, T., Chevallet, M., Luche, S. & Lelong, C. Fully denaturing two-dimensional electrophoresis of membrane proteins: a critical update. Proteomics 8, 3965–3973 (2008).
Zahedi, R.P., Moebius, J. & Sickmann, A. Two-dimensional BAC/SDS-PAGE for membrane proteomics. Subcell. Biochem. 43, 13–20 (2007).
Braun, R.J., Kinkl, N., Beer, M. & Ueffing, M. Two-dimensional electrophoresis of membrane proteins. Anal. Bioanal. Chem. 389, 1033–1045 (2007).
Barnard, E.A. et al. International Union of Pharmacology. XV. Subtypes of gamma-aminobutyric acidA receptors: classification on the basis of subunit structure and receptor function. Pharmacol. Rev. 50, 291–313 (1998).
Sieghart, W. & Sperk, G. Subunit composition, distribution and function of GABA(A) receptor subtypes. Curr. Top. Med. Chem. 2, 795–816 (2002).
Sarter, M., Schneider, H.H. & Stephens, D.N. Treatment strategies for senile dementia: antagonist beta-carbolines. Trends Neurosci. 11, 13–17 (1988).
Paulsen, O. & Moser, E.I. A model of hippocampal memory encoding and retrieval: GABAergic control of synaptic plasticity. Trends Neurosci. 21, 273–278 (1998).
Macdonald, R.L. & Olsen, R.W. GABAA receptor channels. Annu. Rev. Neurosci. 17, 569–602 (1994).
Sieghart, W. & Ernst, M. Heterogeneity of GABA-A receptors: revived interest in the development of subtype-selective drugs. Curr. Med. Chem-Centr. Nervous Syst. 5, 217–242 (2005).
Akentieva, N.P. et al. Society for Neurosciences, Meeting Atlanta, Abstract 527.521 (2006).
Schindler, J., Lewandrowski, U., Sickmann, A., Friauf, E. & Nothwang, H.G. Proteomic analysis of brain plasma membranes isolated by affinity two-phase partitioning. Mol. Cell Proteomics 5, 390–400 (2006).
Kang, S.U., Fuchs, K., Sieghart, W. & Lubec, G. Gel-based mass spectrometric analysis of recombinant GABA(A) receptor subunits representing strongly hydrophobic transmembrane proteins. J. Proteome Res. 7, 3498–3506 (2008).
Baer, A.S. et al. Myelin-mediated inhibition of oligodendrocyte precursor differentiation can be overcome by pharmacological modulation of Fyn-RhoA and protein kinase C signalling. Brain 132, 465–481 (2009).
Chen, W.Q., Kang, S.U. & Lubec, G. Protein profiling by the combination of two independent mass spectrometry techniques. Nat. Protoc. 1, 1446–1452 (2006).
Bierczynska-Krzysik, A., Kang, S.U., Silberrring, J. & Lubec, G. Mass spectrometrical identification of brain proteins including highly insoluble and transmembrane proteins. Neurochem. Int. 49, 245–255 (2006).
Oberacher, H. et al. On the inter-instrument and the inter-laboratory transferability of a tandem mass spectral reference library: 2. Optimization and characterization of the search algorithm. J. Mass Spectrom. 44, 485–493 (2009).
Barrera, N.P. et al. Atomic force microscopy reveals the stoichiometry and subunit arrangement of the alpha4beta3delta GABA(A) receptor. Mol. Pharmacol. 73, 960–967 (2008).
Knight, A.R., Stephenson, F.A., Tallman, J.F. & Ramabahdran, T.V. Monospecific antibodies as probes for the stoichiometry of recombinant GABA(A) receptors. Receptors Channels 7, 213–226 (2000).
Louiset, E., McKernan, R., Sieghart, W. & Vaudry, H. Subunit composition and pharmacological characterization of gamma-aminobutyric acid type A receptors in frog pituitary melanotrophs. Endocrinology 141, 1083–1092 (2000).
Rapallino, M.V. et al. Immunocytochemical study of alpha 1 and beta 2/3 subunits of GABAA receptors in freehand isolated vestibular Deiters' neurons. Receptors Channels 9, 77–81 (2003).
Lubec, G. & Afjehi-Sadat, L. Limitations and pitfalls in protein identification by mass spectrometry. Chem. Rev. 107, 3568–3584 (2007).
Wittig, I., Braun, H.P. & Schagger, H. Blue native PAGE. Nat. Protoc. 1, 418–428 (2006).
Reisinger, V. & Eichacker, L.A. Solubilization of membrane protein complexes for blue native PAGE. J. Proteomics 71, 277–283 (2008).
Lubec, G., Krapfenbauer, K. & Fountoulakis, M. Proteomics in brain research: potentials and limitations. Prog. Neurobiol. 69, 193–211 (2003).
Smith, P.K. et al. Measurement of protein using bicinchoninic acid. Anal. Biochem. 150, 76–85 (1985).
Schagger, H., Cramer, W.A. & von Jagow, G. Analysis of molecular masses and oligomeric states of protein complexes by blue native electrophoresis and isolation of membrane protein complexes by two-dimensional native electrophoresis. Anal. Biochem. 217, 220–230 (1994).
Acknowledgements
We acknowledge the contribution by the Verein zur Durchführung der wissenschaftlichen Forschung auf dem Gebiet der Neonatologie und Kinderintensivmedizin 'Unser Kind'.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Figure 1
Spectra of [311 T.AMDWFIAV.C 318], a part of TMD3 in GABAA receptor α-1 subunit, obtained by nano-ESI-LC-MS/MS Qstar XL. (PDF 73 kb)
Supplementary Figure 2
Positive ionization MS/MS spectrum of the peptide m/z 703.58 amu (PDF 16 kb)
Supplementary Table 1
List of identified GABAA receptor α1 subunit peptides (DOC 753 kb)
Rights and permissions
About this article
Cite this article
Kang, SU., Fuchs, K., Sieghart, W. et al. Gel-based mass spectrometric analysis of a strongly hydrophobic GABAA-receptor subunit containing four transmembrane domains. Nat Protoc 4, 1093–1102 (2009). https://doi.org/10.1038/nprot.2009.92
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2009.92
This article is cited by
-
Formation of GABAA receptor complexes containing α1 and α5 subunits is paralleling a multiple T-maze learning task in mice
Brain Structure and Function (2017)
-
Dorsal hippocampal brain receptor complexes linked to the protein synthesis-dependent late phase (LTP) in the rat
Brain Structure and Function (2015)
-
The use of native gels for the concomitant determination of protein sequences and modifications by mass spectrometry with subsequent conformational and functional analysis of native proteins following electro-elution
Amino Acids (2013)
-
Hippocampal levels of GluR1 and GluR2 complexes are modulated by training in the multiple T-Maze in C57BL/6J mice
Brain Structure and Function (2012)
-
Modafinil improves performance in the multiple T-Maze and modifies GluR1, GluR2, D2 and NR1 receptor complex levels in the C57BL/6J mouse
Amino Acids (2012)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.