Abstract
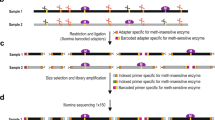
The base composition of PCR products derived from sodium bisulfite-modified templates is methylation dependent. Hence, methylated and unmethylated, PCR products show different melting profiles when subjected to thermal denaturation. The methylation-sensitive high-resolution melting (MS-HRM) protocol is based on the comparison of the melting profiles of PCR products from unknown samples with profiles specific for PCR products derived from methylated and unmethylated control DNAs. The protocol consists of PCR amplification of bisulfite-modified DNA with primers designed to proportionally amplify both methylated and unmethylated templates and subsequent high-resolution melting analysis of the PCR product. The MS-HRM protocol allows in-tube determination of the methylation status of the locus of interest following sodium bisulfite modification of template DNA in less than 3 h. Here, we provide a protocol for MS-HRM, which enables highly sensitive, labor- and cost-efficient single-locus methylation studies on the basis of DNA high-resolution melting technology.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Bird, A.P. & Wolffe, A.P. Methylation-induced repression—belts, braces, and chromatin. Cell 99, 451–454 (1999).
Jones, P.A. & Laird, P.W. Cancer epigenetics comes of age. Nat. Genet. 21, 163–167 (1999).
Esteller, M., Corn, P.G., Baylin, S.B. & Herman, J.G. A gene hypermethylation profile of human cancer. Cancer Res. 61, 3225–3229 (2001).
Costello, J.F. et al. Aberrant CpG-island methylation has non-random and tumour-type-specific patterns. Nat. Genet. 24, 132–138 (2000).
Palmisano, W.A. et al. Predicting lung cancer by detecting aberrant promoter methylation in sputum. Cancer Res. 60, 5954–5958 (2000).
Wojdacz, T.K., Dobrovic, A. & Algar, E.M. Rapid detection of methylation change at H19 in human imprinting disorders using methylation-sensitive high-resolution melting. Hum. Mutat. 29, 1255–1260 (2008).
Feinberg, A.P. Epigenetics at the epicenter of modern medicine. JAMA 299, 1345–1350 (2008).
Wojdacz, T.K. & Hansen, L.L. Techniques used in studies of age-related DNA methylation changes. Ann. N.Y. Acad. Sci. 1067, 479–487 (2006).
Dobrovic, A. Methods for Analyses of DNA Methylation. Molecular Diagnostics for the Clinical Laboratorian. (Humana Press, Totowa, NJ, 11, 2005).
Herman, J.G., Graff, J.R., Myohanen, S., Nelkin, B.D. & Baylin, S.B. Methylation-specific PCR: a novel PCR assay for methylation status of CpG islands. Proc. Natl. Acad. Sci. USA 93, 9821–9826 (1996).
Eads, C.A. et al. MethyLight: a high-throughput assay to measure DNA methylation. Nucleic Acids Res. 28, E32 (2000).
Frommer, M. et al. A genomic sequencing protocol that yields a positive display of 5-methylcytosine residues in individual DNA strands. Proc. Natl. Acad. Sci. USA 89, 1827–1831 (1992).
Clark, S.J., Harrison, J., Paul, C.L. & Frommer, M. High sensitivity mapping of methylated cytosines. Nucleic Acids Res. 22, 2990–2997 (1994).
Eads, C.A. & Laird, P.W. Combined bisulfite restriction analysis (COBRA). Methods Mol. Biol. 200, 71–85 (2002).
Bianco, T., Hussey, D. & Dobrovic, A. Methylation-sensitive, single-strand conformation analysis (MS-SSCA): a rapid method to screen for and analyze methylation. Hum. Mutat. 14, 289–293 (1999).
Worm, J., Aggerholm, A. & Guldberg, P. In-tube DNA methylation profiling by fluorescence melting curve analysis. Clin. Chem. 47, 1183–1189 (2001).
Wojdacz, T.K. & Dobrovic, A. Methylation-sensitive high resolution melting (MS-HRM): a new approach for sensitive and high-throughput assessment of methylation. Nucleic Acids Res. 35, e41 (2007).
Warnecke, P.M. et al. Detection and measurement of PCR bias in quantitative methylation analysis of bisulphite-treated DNA. Nucleic Acids Res. 25, 4422–4426 (1997).
Wojdacz, T.K. & Hansen, L.L. Reversal of PCR bias for improved sensitivity of the DNA methylation melting curve assay. Biotechniques 41, 274, 276, 278 (2006).
Shen, L., Guo, Y., Chen, X., Ahmed, S. & Issa, J.P. Optimizing annealing temperature overcomes bias in bisulfite PCR methylation analysis. Biotechniques 42, 48, 50, 52 passim (2007).
Wittwer, C.T., Reed, G.H., Gundry, C.N., Vandersteen, J.G. & Pryor, R.J. High-resolution genotyping by amplicon melting analysis using LCGreen. Clin. Chem. 49, 853–860 (2003).
Wojdacz, T.K., Hansen, L.L. & Dobrovic, A. A new approach to primer design for the control of PCR bias in methylation studies. BMC Res. Notes 1, 54 (2008).
Gudnason, H., Dufva, M., Bang, D.D. & Wolff, A. Comparison of multiple DNA dyes for real-time PCR: effects of dye concentration and sequence composition on DNA amplification and melting temperature. Nucleic Acids Res. 35, e127 (2007).
Acknowledgements
We thank the Lundbeck, Toyota and Harboe foundations (grants to T.K.W. and L.L.H.), the National Health and Medical Research Council of Australia and US Army Medical Research and Materiel Command (grants to A.D.) for the support of the research leading to this publication.
Author information
Authors and Affiliations
Contributions
A.D. and L.L.H. contributed equally to this work.
Corresponding author
Ethics declarations
Competing interests
The authors are co-inventors on patent applications on aspects of the MS-HRM methodology. The patents have been filed for by University of Aarhus and Peter MacCallum Cancer Centre.
Rights and permissions
About this article
Cite this article
Wojdacz, T., Dobrovic, A. & Hansen, L. Methylation-sensitive high-resolution melting. Nat Protoc 3, 1903–1908 (2008). https://doi.org/10.1038/nprot.2008.191
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2008.191
This article is cited by
-
Lower Levels of TET2 Gene Expression, with a Higher Level of TET2 Promoter Methylation in Patients with AML; Evidence for the Role of Aberrant Methylation in AML Pathogenesis
Indian Journal of Hematology and Blood Transfusion (2024)
-
Aberrant promoter hypermethylation of miR-335 and miR-145 is involved in breast cancer PD-L1 overexpression
Scientific Reports (2023)
-
Arsenic concentration in topsoil of central Chile is associated with aberrant methylation of P53 gene in human blood cells: a cross-sectional study
Environmental Science and Pollution Research (2022)
-
Global DNA methylation profile at LINE-1 repeats and promoter methylation of genes involved in DNA damage response and repair pathways in human peripheral blood mononuclear cells in response to γ-radiation
Molecular and Cellular Biochemistry (2022)
-
Laboratory methods to decipher epigenetic signatures: a comparative review
Cellular & Molecular Biology Letters (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.