Abstract
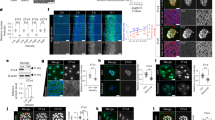
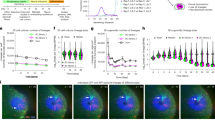
Postnatal bone marrow houses mesenchymal progenitor cells that are osteoblast precursors. These cells have established therapeutic potential, but they are difficult to maintain and expand in vitro, presumably because little is known about the mechanisms controlling their fate decisions. To investigate the potential role of Notch signaling in osteoblastogenesis, we used conditional alleles to genetically remove components of the Notch signaling system during skeletal development. We found that disruption of Notch signaling in the limb skeletogenic mesenchyme markedly increased trabecular bone mass in adolescent mice. Notably, mesenchymal progenitors were undetectable in the bone marrow of mice with high bone mass. As a result, these mice developed severe osteopenia as they aged. Moreover, Notch signaling seemed to inhibit osteoblast differentiation through Hes or Hey proteins, which diminished Runx2 transcriptional activity via physical interaction. These results support a model wherein Notch signaling in bone marrow normally acts to maintain a pool of mesenchymal progenitors by suppressing osteoblast differentiation. Thus, mesenchymal progenitors may be expanded in vitro by activating the Notch pathway, whereas bone formation in vivo may be enhanced by transiently suppressing this pathway.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Artavanis-Tsakonas, S., Rand, M.D. & Lake, R.J. Notch signaling: cell fate control and signal integration in development. Science 284, 770–776 (1999).
Chiba, S. Notch signaling in stem cell systems. Stem Cells 24, 2437–2447 (2006).
Schroeter, E.H., Kisslinger, J.A. & Kopan, R. Notch-1 signalling requires ligand-induced proteolytic release of intracellular domain. Nature 393, 382–386 (1998).
Honjo, T. The shortest path from the surface to the nucleus: RBP-Jκ/Su(H) transcription factor. Genes Cells 1, 1–9 (1996).
Kopan, R. & Goate, A. A common enzyme connects Notch signaling and Alzheimer's disease. Genes Dev. 14, 2799–2806 (2000).
Donoviel, D.B. et al. Mice lacking both presenilin genes exhibit early embryonic patterning defects. Genes Dev. 13, 2801–2810 (1999).
Herreman, A. et al. Presenilin 2 deficiency causes a mild pulmonary phenotype and no changes in amyloid precursor protein processing but enhances the embryonic lethal phenotype of presenilin 1 deficiency. Proc. Natl. Acad. Sci. USA 96, 11872–11877 (1999).
Krebs, L.T. et al. Notch signaling is essential for vascular morphogenesis in mice. Genes Dev. 14, 1343–1352 (2000).
Hamada, Y. et al. Mutation in ankyrin repeats of the mouse Notch2 gene induces early embryonic lethality. Development 126, 3415–3424 (1999).
Swiatek, P.J., Lindsell, C.E., del Amo, F.F., Weinmaster, G. & Gridley, T. Notch1 is essential for postimplantation development in mice. Genes Dev. 8, 707–719 (1994).
Domenga, V. et al. Notch3 is required for arterial identity and maturation of vascular smooth muscle cells. Genes Dev. 18, 2730–2735 (2004).
Hrabe de Angelis, M., McIntyre, J. II & Gossler, A. Maintenance of somite borders in mice requires the Delta homologue DII1. Nature 386, 717–721 (1997).
Duarte, A. et al. Dosage-sensitive requirement for mouse Dll4 in artery development. Genes Dev. 18, 2474–2478 (2004).
Gale, N.W. et al. Haploinsufficiency of delta-like 4 ligand results in embryonic lethality due to major defects in arterial and vascular development. Proc. Natl. Acad. Sci. USA 101, 15949–15954 (2004).
Xue, Y. et al. Embryonic lethality and vascular defects in mice lacking the Notch ligand Jagged1. Hum. Mol. Genet. 8, 723–730 (1999).
Jiang, R. et al. Defects in limb, craniofacial, and thymic development in Jagged2 mutant mice. Genes Dev. 12, 1046–1057 (1998).
Dunwoodie, S.L. et al. Axial skeletal defects caused by mutation in the spondylocostal dysplasia/pudgy gene Dll3 are associated with disruption of the segmentation clock within the presomitic mesoderm. Development 129, 1795–1806 (2002).
Shen, J. et al. Skeletal and CNS defects in presenilin-1–deficient mice. Cell 89, 629–639 (1997).
Wong, P.C. et al. Presenilin 1 is required for Notch1 and DII1 expression in the paraxial mesoderm. Nature 387, 288–292 (1997).
Pan, Y., Liu, Z., Shen, J. & Kopan, R. Notch1 and 2 cooperate in limb ectoderm to receive an early Jagged2 signal regulating interdigital apoptosis. Dev. Biol. 286, 472–482 (2005).
Logan, M. et al. Expression of CrerRecombinase in the developing mouse limb bud driven by a Prxl enhancer. Genesis 33, 77–80 (2002).
Miao, D. et al. Osteoblast-derived PTHrP is a potent endogenous bone anabolic agent that modifies the therapeutic efficacy of administered PTH 1–34. J. Clin. Invest. 115, 2402–2411 (2005).
Owen, M. & Friedenstein, A.J. Stromal stem cells: marrow-derived osteogenic precursors. Ciba Found. Symp. 136, 42–60 (1988).
Kuznetsov, S.A. et al. The interplay of osteogenesis and hematopoiesis: expression of a constitutively active PTH/PTHrP receptor in osteogenic cells perturbs the establishment of hematopoiesis in bone and of skeletal stem cells in the bone marrow. J. Cell Biol. 167, 1113–1122 (2004).
Nichols, A.M. et al. Notch pathway is dispensable for adipocyte specification. Genesis 40, 40–44 (2004).
Karsenty, G. & Wagner, E.F. Reaching a genetic and molecular understanding of skeletal development. Dev. Cell 2, 389–406 (2002).
Bai, S. et al. Notch1 regulates osteoclastogenesis directly in osteoclast precursors and indirectly via osteoblast lineage cells. J. Biol. Chem. published online, doi:10.1074/jbc.M707000200 (22 December 2007).
Iso, T., Kedes, L. & Hamamori, Y. HES and HERP families: multiple effectors of the Notch signaling pathway. J. Cell. Physiol. 194, 237–255 (2003).
Duncan, A.W. et al. Integration of Notch and Wnt signaling in hematopoietic stem cell maintenance. Nat. Immunol. 6, 314–322 (2005).
Vooijs, M. et al. Mapping the consequence of Notch1 proteolysis in vivo with NIP-CRE. Development 134, 535–544 (2007).
Soriano, P. Generalized lacZ expression with the ROSA26 Cre reporter strain. Nat. Genet. 21, 70–71 (1999).
Deregowski, V., Gazzerro, E., Priest, L., Rydziel, S. & Canalis, E. Notch 1 overexpression inhibits osteoblastogenesis by suppressing Wnt/β-catenin but not bone morphogenetic protein signaling. J. Biol. Chem. 281, 6203–6210 (2006).
Sciaudone, M., Gazzerro, E., Priest, L., Delany, A.M. & Canalis, E. Notch 1 impairs osteoblastic cell differentiation. Endocrinology 144, 5631–5639 (2003).
Shindo, K. et al. Osteogenic differentiation of the mesenchymal progenitor cells, Kusa is suppressed by Notch signaling. Exp. Cell Res. 290, 370–380 (2003).
Nobta, M. et al. Critical regulation of bone morphogenetic protein-induced osteoblastic differentiation by Delta1/Jagged1-activated Notch1 signaling. J. Biol. Chem. 280, 15842–15848 (2005).
Tezuka, K. et al. Stimulation of osteoblastic cell differentiation by Notch. J. Bone Miner. Res. 17, 231–239 (2002).
Garg, V. et al. Mutations in NOTCH1 cause aortic valve disease. Nature 437, 270–274 (2005).
Zamurovic, N., Cappellen, D., Rohner, D. & Susa, M. Coordinated activation of notch, Wnt, and transforming growth factor-β signaling pathways in bone morphogenic protein 2–induced osteogenesis. Notch target gene Hey1 inhibits mineralization and Runx2 transcriptional activity. J. Biol. Chem. 279, 37704–37715 (2004).
Conlon, R.A., Reaume, A.G. & Rossant, J. Notch1 is required for the coordinate segmentation of somites. Development 121, 1533–1545 (1995).
Pan, Y. et al. Gamma-secretase functions through Notch signaling to maintain skin appendages but is not required for their patterning or initial morphogenesis. Dev. Cell 7, 731–743 (2004).
McCright, B., Lozier, J. & Gridley, T. Generation of new Notch2 mutant alleles. Genesis 44, 29–33 (2006).
Yu, H. et al. APP processing and synaptic plasticity in presenilin-1 conditional knockout mice. Neuron 31, 713–726 (2001).
Hilton, M.J., Tu, X., Cook, J., Hu, H. & Long, F. Ihh controls cartilage development by antagonizing Gli3, but requires additional effectors to regulate osteoblast and vascular development. Development 132, 4339–4351 (2005).
Tu, X. et al. Noncanonical Wnt signaling through G protein–linked PKCδ activation promotes bone formation. Dev. Cell 12, 113–127 (2007).
Huppert, S.S. et al. Embryonic lethality in mice homozygous for a processing-deficient allele of Notch1. Nature 405, 966–970 (2000).
Ong, C.T. et al. Target selectivity of vertebrate notch proteins. Collaboration between discrete domains and CSL-binding site architecture determines activation probability. J. Biol. Chem. 281, 5106–5119 (2006).
Acknowledgements
This work was supported in part by US National Institutes of Health grants DK065789 (F.L.), HD044056 (R.K.), AR046852 (F.P.R.), AR046523 (S.L.T.), DK11794 (H.M.K.) and 5T32AR07033 (M.J.H.). We thank R. Civitelli (Washington University) for pCMV-Runx2 and p60SE2-Luc, R. Kageyama (Kyoto University) for pSV2-CMV-Hes1, P. Robey and S. Kuznetsov for advice on bone marrow CFU-F assays and J. Shen (Harvard Medical School) and Thomas Gridley (Jackson Laboratory) for mouse strains. We also thank D. Towler and M. Silva for their help with the response to reviewers' comments.
Author information
Authors and Affiliations
Contributions
M.J.H., animal studies and figure preparation; X.T., cell culture work and animal studies; X.W. coimmunoprecipitation experiments; S.B., H.Z., F.P.R. and S.L.T., TRAP staining, serum CTX1 assays and discussion; T.K. and H.M.K., Col1-Cre mice; R.K., Notch reagents and manuscript editing; F.L., project directing, figure preparation and manuscript writing.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figs. 1–5 and Supplementary Table 1 (PDF 937 kb)
Rights and permissions
About this article
Cite this article
Hilton, M., Tu, X., Wu, X. et al. Notch signaling maintains bone marrow mesenchymal progenitors by suppressing osteoblast differentiation. Nat Med 14, 306–314 (2008). https://doi.org/10.1038/nm1716
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm1716
This article is cited by
-
Cellular senescence in skeletal disease: mechanisms and treatment
Cellular & Molecular Biology Letters (2023)
-
Mesenchymal stem cells and dental implant osseointegration during aging: from mechanisms to therapy
Stem Cell Research & Therapy (2023)
-
Spatial transcriptomic interrogation of the murine bone marrow signaling landscape
Bone Research (2023)
-
Loss of Notch signaling in skeletal stem cells enhances bone formation with aging
Bone Research (2023)
-
Integrated single-cell and bulk RNA sequencing analysis identified pyroptosis-related signature for diagnosis and prognosis in osteoarthritis
Scientific Reports (2023)