Abstract
Decades of research went into understanding immune cell development and function without awareness that consideration of a key element, microRNA (miRNA), was lacking. The discovery of miRNAs as regulators of developmental events in model organisms suggested to many investigators that miRNA might be involved in the immune system. In the past few years, widespread examination of this possibility has produced notable results. Results have shown that miRNAs affect mammalian immune cell differentiation, the outcome of immune responses to infection and the development of diseases of immunological origin. Some miRNAs repress expression of target proteins with well established functions in hematopoiesis. Here we bring together much of this work, which has so far only scratched the surface of this very fertile field of investigation, and show how the results illuminate many historic questions about hematopoiesis and immune function.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Baulcombe, D. RNA silencing in plants. Nature 431, 356–363 (2004).
Fire, A. et al. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 391, 806–811 (1998).
Lee, R.C., Feinbaum, R.L. & Ambros, V. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 75, 843–854 (1993).
Cai, X., Hagedorn, C.H. & Cullen, B.R. Human microRNAs are processed from capped, polyadenylated transcripts that can also function as mRNAs. RNA 10, 1957–1966 (2004).
Lee, Y. et al. MicroRNA genes are transcribed by RNA polymerase II. EMBO J. 23, 4051–4060 (2004).
Han, J. et al. Molecular basis for the recognition of primary microRNAs by the Drosha-DGCR8 complex. Cell 125, 887–901 (2006).
Yi, R., Qin, Y., Macara, I.G. & Cullen, B.R. Exportin-5 mediates the nuclear export of pre-microRNAs and short hairpin RNAs. Genes Dev. 17, 3011–3016 (2003).
Chendrimada, T.P. et al. MicroRNA silencing through RISC recruitment of eIF6. Nature 447, 823–828 (2007).
Chendrimada, T.P. et al. TRBP recruits the Dicer complex to Ago2 for microRNA processing and gene silencing. Nature 436, 740–744 (2005).
O'Carroll, D. et al. A Slicer-independent role for Argonaute 2 in hematopoiesis and the microRNA pathway. Genes Dev. 21, 1999–2004 (2007).
Cobb, B.S. et al. T cell lineage choice and differentiation in the absence of the RNase III enzyme Dicer. J. Exp. Med. 201, 1367–1373 (2005).
Muljo, S.A. et al. Aberrant T cell differentiation in the absence of Dicer. J. Exp. Med. 202, 261–269 (2005).
Koralov, S.B. et al. Dicer ablation affects antibody diversity and cell survival in the B lymphocyte lineage. Cell 132, 860–874 (2008). This paper describes an early B cell–specific ablation of Dicer expression that leads to a block in B cell development at the transition from pro–B cell to pre–B cell and identifies dysregulation of expression of the miR-17–92 cluster and its target gene Bim as the culprits of this defect.
Bartel, D.P. & Chen, C.Z. Micromanagers of gene expression: the potentially widespread influence of metazoan microRNAs. Nat. Rev. Genet. 5, 396–400 (2004).
Landgraf, P. et al. A mammalian microRNA expression atlas based on small RNA library sequencing. Cell 129, 1401–1414 (2007).
Georgantas, R.W., III et al. CD34+ hematopoietic stem-progenitor cell microRNA expression and function: a circuit diagram of differentiation control. Proc. Natl. Acad. Sci. USA 104, 2750–2755 (2007).
Garzon, R. et al. MicroRNA fingerprints during human megakaryocytopoiesis. Proc. Natl. Acad. Sci. USA 103, 5078–5083 (2006).
Zhan, M., Miller, C.P., Papayannopoulou, T., Stamatoyannopoulos, G. & Song, C.Z. MicroRNA expression dynamics during murine and human erythroid differentiation. Exp. Hematol. 35, 1015–1025 (2007).
Cobb, B.S. et al. A role for Dicer in immune regulation. J. Exp. Med. 203, 2519–2527 (2006).
Neilson, J.R., Zheng, G.X., Burge, C.B. & Sharp, P.A. Dynamic regulation of miRNA expression in ordered stages of cellular development. Genes Dev. 21, 578–589 (2007).
O'Connell, R.M., Taganov, K.D., Boldin, M.P., Cheng, G. & Baltimore, D. MicroRNA-155 is induced during the macrophage inflammatory response. Proc. Natl. Acad. Sci. USA 104, 1604–1609 (2007).
Taganov, K.D., Boldin, M.P., Chang, K.J. & Baltimore, D. NF-κB-dependent induction of microRNA miR-146, an inhibitor targeted to signaling proteins of innate immune responses. Proc. Natl. Acad. Sci. USA 103, 12481–12486 (2006). This paper identifies three endotoxin-responsive miRNAs (miR-146, miR-155 and miR-132) in monocytic cells and proposes involvement of the miR-146 family in the negative regulation of Toll-like receptor signaling.
Wu, H. et al. miRNA Profiling of Naive, Effector and memory CD8 T Cells. PLoS ONE 2, e1020 (2007).
Calin, G.A. & Croce, C.M. MicroRNA signatures in human cancers. Nat. Rev. Cancer 6, 857–866 (2006).
Tam, W., Ben-Yehuda, D. & Hayward, W.S. BIC, a novel gene activated by proviral insertions in avian leukosis virus-induced lymphomas, is likely to function through its noncoding RNA. Mol. Cell. Biol. 17, 1490–1502 (1997).
Haasch, D. et al. T cell activation induces a noncoding RNA transcript sensitive to inhibition by immunosuppressant drugs and encoded by the proto-oncogene, BIC. Cell. Immunol. 217, 78–86 (2002).
Thai, T.H. et al. Regulation of the germinal center response by microRNA-155. Science 316, 604–608 (2007). This report describes analysis of the physiological functions of miR-155 in the immune system through the targeted deletion of this miRNA in mice.
Eis, P.S. et al. Accumulation of miR-155 and BIC RNA in human B cell lymphomas. Proc. Natl. Acad. Sci. USA 102, 3627–3632 (2005).
Fulci, V. et al. Quantitative technologies establish a novel microRNA profile of chronic lymphocytic leukemia. Blood 109, 4944–4951 (2007).
Kluiver, J. et al. BIC and miR-155 are highly expressed in Hodgkin, primary mediastinal and diffuse large B cell lymphomas. J. Pathol. 207, 243–249 (2005).
O'Connell, R.M. et al. Sustained expression of microRNA-155 in hematopoietic stem cells causes a myeloproliferative disorder. J. Exp. Med. 205, 585–594 (2008).
Tam, W. & Dahlberg, J.E. miR-155/BIC as an oncogenic microRNA. Genes Chromosom. Cancer 45, 211–212 (2006).
van den Berg, A. et al. High expression of B-cell receptor inducible gene BIC in all subtypes of Hodgkin lymphoma. Genes Chromosom. Cancer 37, 20–28 (2003).
Costinean, S. et al. Pre-B cell proliferation and lymphoblastic leukemia/high-grade lymphoma in E(mu)-miR155 transgenic mice. Proc. Natl. Acad. Sci. USA 103, 7024–7029 (2006).
Rodriguez, A. et al. Requirement of bic/microRNA-155 for normal immune function. Science 316, 608–611 (2007). This paper analyzes the physiological functions of miR-155 in the immune system through the targeted deletion of this miRNA in mice.
Vigorito, E. et al. microRNA-155 regulates the generation of immunoglobulin class-switched plasma cells. Immunity 27, 847–859 (2007).
Dorsett, Y. et al. MicroRNA-155 suppresses activation-induced cytidine deaminase-mediated Myc-Igh translocation. Immunity 28, 630–638 (2008).
Teng, G. et al. MicroRNA-155 is a negative regulator of activation-induced cytidine deaminase. Immunity 28, 621–629 (2008).
Zheng, Y. et al. Genome-wide analysis of Foxp3 target genes in developing and mature regulatory T cells. Nature 445, 936–940 (2007).
Brown, B.D. et al. Endogenous microRNA can be broadly exploited to regulate transgene expression according to tissue, lineage and differentiation state. Nat. Biotechnol. 25, 1457–1467 (2007).
Kluiver, J. et al. Regulation of pri-microRNA BIC transcription and processing in Burkitt lymphoma. Oncogene 26, 3769–3776 (2007).
Yin, Q. et al. microRNA-155 is an Epstein-Barr Virus induced gene that modulates Epstein Barr virus regulated gene expression pathways. J. Virol. 82, 5295–5306 (2008).
Garzon, R. et al. Distinctive microRNA signature of acute myeloid leukemia bearing cytoplasmic mutated nucleophosmin. Proc. Natl. Acad. Sci. USA 105, 3945–3950 (2008).
Gottwein, E. et al. A viral microRNA functions as an orthologue of cellular miR-155. Nature 450, 1096–1099 (2007).
Skalsky, R.L. et al. Kaposi's sarcoma-associated herpesvirus encodes an ortholog of miR-155. J. Virol. 81, 12836–12845 (2007).
Jiang, J., Lee, E.J. & Schmittgen, T.D. Increased expression of microRNA-155 in Epstein-Barr virus transformed lymphoblastoid cell lines. Genes Chromosom. Cancer 45, 103–106 (2006).
Mrazek, J., Kreutmayer, S.B., Grasser, F.A., Polacek, N. & Huttenhofer, A. Subtractive hybridization identifies novel differentially expressed ncRNA species in EBV-infected human B cells. Nucleic Acids Res. 35, e73 (2007).
Cameron, J.E. et al. Epstein-Barr virus latent membrane protein 1 induces cellular MicroRNA miR-146a, a modulator of lymphocyte signaling pathways. J. Virol. 82, 1946–1958 (2008).
Motsch, N., Pfuhl, T., Mrazek, J., Barth, S. & Grasser, F.A. Epstein-Barr virus-encoded latent membrane protein 1 (LMP1) induces the expression of the cellular microRNA miR-146a. RNA Biol., 4, 131–137 (2007).
Monticelli, S. et al. MicroRNA profiling of the murine hematopoietic system. Genome Biol. 6, R71 (2005).
Griffiths-Jones, S. The microRNA registry. Nucleic Acids Res. 32, D109–D111 (2004).
Obernosterer, G., Leuschner, P.J., Alenius, M. & Martinez, J. Post-transcriptional regulation of microRNA expression. RNA 12, 1161–1167 (2006).
Thomson, J.M. et al. Extensive post-transcriptional regulation of microRNAs and its implications for cancer. Genes Dev. 20, 2202–2207 (2006).
Lim, C.A. et al. Genome-wide mapping of RELA(p65) binding identifies E2F1 as a transcriptional activator recruited by NF-κB upon TLR4 activation. Mol. Cell 27, 622–635 (2007).
Chang, T.C. et al. Widespread microRNA repression by Myc contributes to tumorigenesis. Nat. Genet. 40, 43–50 (2008).
Kawai, T. & Akira, S. Signaling to NF-κB by Toll-like receptors. Trends Mol. Med. 13, 460–469 (2007).
Lin, S.L., Chiang, A., Chang, D. & Ying, S.Y. Loss of mir-146a function in hormone-refractory prostate cancer. RNA 14, 417–424 (2008).
Xiao, C. et al. MiR-150 controls B cell differentiation by targeting the transcription factor c-Myb. Cell 131, 146–159 (2007). This paper describes the function of miR-150 in the control of B cell development through a combination of gain- and loss-of-function approaches.
Lu, J. et al. microRNA-mediated control of cell fate in megakaryocyte-erythrocyte progenitors. Dev. Cell 14, 843–853 (2008). It is known that miR-150 promotes megakaryocytic differentiation at the expense of erythroid differentiation, thereby specifying lineage 'choice' at the megakaryocyte-erythroid progenitor stage of hematopoietic development. This work expands the hematopoietic function of miR-150, previously confined to lymphoid development, while also confirming previous findings of c-Myb as an important target of miR-150.
Zhou, B., Wang, S., Mayr, C., Bartel, D.P. & Lodish, H.F. miR-150, a microRNA expressed in mature B and T cells, blocks early B cell development when expressed prematurely. Proc. Natl. Acad. Sci. USA 104, 7080–7085 (2007).
Montecino-Rodriguez, E. & Dorshkind, K. New perspectives in B-1 B cell development and function. Trends Immunol. 27, 428–433 (2006).
Emambokus, N. et al. Progression through key stages of haemopoiesis is dependent on distinct threshold levels of c-Myb. EMBO J. 22, 4478–4488 (2003).
Chen, C.Z., Li, L., Lodish, H.F. & Bartel, D.P. MicroRNAs modulate hematopoietic lineage differentiation. Science 303, 83–86 (2004). This early work demonstrates functions for miRNA in regulating lineage decisions during hematopoiesis, setting the paradigm in this field. It identifies miR-181, miR-142 and miR-223 as being important in hematopoietic development.
Li, Q.J. et al. miR-181a is an intrinsic modulator of T cell sensitivity and selection. Cell 129, 147–161 (2007).
Choong, M.L., Yang, H.H. & McNiece, I. MicroRNA expression profiling during human cord blood-derived CD34 cell erythropoiesis. Exp. Hematol. 35, 551–564 (2007).
Naguibneva, I. et al. The microRNA miR-181 targets the homeobox protein Hox-A11 during mammalian myoblast differentiation. Nat. Cell Biol. 8, 278–284 (2006).
Mendell, J.T. MicroRNAs: critical regulators of development, cellular physiology and malignancy. Cell Cycle 4, 1179–1184 (2005).
Ventura, A. et al. Targeted deletion reveals essential and overlapping functions of the miR-17 through 92 family of miRNA clusters. Cell 132, 875–886 (2008).
Xiao, C. et al. Lymphoproliferative disease and autoimmunity in mice with increased miR-17–92 expression in lymphocytes. Nat. Immunol. 9, 405–414 (2008).
Fontana, L. et al. MicroRNAs 17–5p-20a-106a control monocytopoiesis through AML1 targeting and M-CSF receptor upregulation. Nat. Cell Biol. 9, 775–787 (2007). Monocytopoiesis is dependent on downregulation of the miR-17–92 cluster, which allows expression of AML1 that in turn is a transcriptional repressor of this miRNA cluster. This paper demonstrates a mutually inhibitory miRNA–transcription factor circuit that regulates monocytic lineage differentiation.
Johnnidis, J.B. et al. Regulation of progenitor cell proliferation and granulocyte function by microRNA-223. Nature 451, 1125–1129 (2008). Mice lacking miR-223 have more hyperfunctional neutrophils and pathologic sequelae thereof, somewhat unexpectedly, given previous data showing that this miRNA promoted granulopoiesis. A new function for miRNA in linking granulocytic lineage development with functional activation is suggested by this study.
Masaki, S., Ohtsuka, R., Abe, Y., Muta, K. & Umemura, T. Expression patterns of microRNAs 155 and 451 during normal human erythropoiesis. Biochem. Biophys. Res. Commun. 364, 509–514 (2007).
Fazi, F. et al. Epigenetic silencing of the myelopoiesis regulator microRNA-223 by the AML1/ETO oncoprotein. Cancer Cell 12, 457–466 (2007).
Fazi, F. et al. A minicircuitry comprised of microRNA-223 and transcription factors NFI-A and C/EBPα regulates human granulopoiesis. Cell 123, 819–831 (2005).
Garzon, R. et al. MicroRNA gene expression during retinoic acid-induced differentiation of human acute promyelocytic leukemia. Oncogene 26, 4148–4157 (2007).
Fukao, T. et al. An evolutionarily conserved mechanism for microRNA-223 expression revealed by microRNA gene profiling. Cell 129, 617–631 (2007).
Felli, N. et al. MicroRNAs 221 and 222 inhibit normal erythropoiesis and erythroleukemic cell growth via kit receptor down-modulation. Proc. Natl. Acad. Sci. USA 102, 18081–18086 (2005).
Rosa, A. et al. The interplay between the master transcription factor PU.1 and miR-424 regulates human monocyte/macrophage differentiation. Proc. Natl. Acad. Sci. USA 104, 19849–19854 (2007).
Pheasant, M. & Mattick, J.S. Raising the estimate of functional human sequences. Genome Res. 17, 1245–1253 (2007).
Acknowledgements
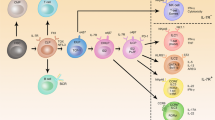
We apologize to those whose work we were unable to cite because of space limitations. We thank C. Koenig for developing the diagram presented in Figure 1. Supported by the US National Institutes of Health (D.B), the Irvington Institute Fellowship Program of the Cancer Research Institute (R.M.O.) and the American Society of Hematology (D.S.R.).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Baltimore, D., Boldin, M., O'Connell, R. et al. MicroRNAs: new regulators of immune cell development and function. Nat Immunol 9, 839–845 (2008). https://doi.org/10.1038/ni.f.209
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.f.209
This article is cited by
-
Early Leishmania infectivity depends on miR-372/373/520d family-mediated reprogramming of polyamines metabolism in THP-1-derived macrophages
Scientific Reports (2024)
-
miR-20b-5p exerts protective effects against experimental autoimmune encephalomyelitis in mice by inhibiting NLRP3 transcription and NLRP3/ASC/caspase-1 axis activation
Molecular & Cellular Toxicology (2023)
-
A noncoding secret to stay young—protecting Treg cells to keep the balance in the liver
Science China Life Sciences (2023)
-
MicroRNA profile of circulating CD4+ T cells in aged patients with atherosclerosis obliterans
BMC Cardiovascular Disorders (2022)
-
MicroRNA-155 expression with Brucella infection in vitro and in vivo and decreased serum levels of MicroRNA-155 in patients with brucellosis
Scientific Reports (2022)