Abstract
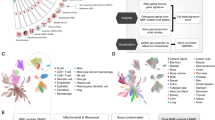
Increasing evidence suggests that changes in the cellular microenvironment contribute to tumorigenesis, but the molecular basis of these alterations is not well understood. Although epigenetic modifications of the neoplastic cells in tumors have been firmly implicated in tumorigenesis1, it is not known whether epigenetic modifications occur in the non-neoplastic stromal cells. To address this question in an unbiased and genome-wide manner, we developed a new method, methylation-specific digital karyotyping, and applied it to epithelial and myoepithelial cells, stromal fibroblasts from normal breast tissue, and in situ and invasive breast carcinomas. Our analyses showed that distinct epigenetic alterations occur in all three cell types during breast tumorigenesis in a tumor stage– and cell type–specific manner, suggesting that epigenetic changes have a role in the maintenance of the abnormal cellular microenvironment in breast cancer.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Fazzari, M.J. & Greally, J.M. Epigenomics: beyond CpG islands. Nat. Rev. Genet. 5, 446–455 (2004).
Allinen, M. et al. Molecular characterization of the tumor microenvironment in breast cancer. Cancer Cell 6, 17–32 (2004).
Bissell, M.J. & Radisky, D. Putting tumours in context. Nat. Rev. Cancer 1, 46–54 (2001).
Bissell, M.J., Radisky, D.C., Rizki, A., Weaver, V.M. & Petersen, O.W. The organizing principle: microenvironmental influences in the normal and malignant breast. Differentiation 70, 537–546 (2002).
Tlsty, T.D. & Hein, P.W. Know thy neighbor: stromal cells can contribute oncogenic signals. Curr. Opin. Genet. Dev. 11, 54–59 (2001).
Jones, P.A. & Takai, D. The role of DNA methylation in mammalian epigenetics. Science 293, 1068–1070 (2001).
Feinberg, A.P. & Tycko, B. The history of cancer epigenetics. Nat. Rev. Cancer 4, 143–153 (2004).
Adany, R., Heimer, R., Caterson, B., Sorrell, J.M. & Iozzo, R.V. Altered expression of chondroitin sulfate proteoglycan in the stroma of human colon carcinoma. Hypomethylation of PG-40 gene correlates with increased PG-40 content and mRNA levels. J. Biol. Chem. 265, 11389–11396 (1990).
Adany, R. & Iozzo, R.V. Altered methylation of versican proteoglycan gene in human colon carcinoma. Biochem. Biophys. Res. Commun. 171, 1402–1413 (1990).
Adany, R. & Iozzo, R.V. Hypomethylation of the decorin proteoglycan gene in human colon cancer. Biochem. J. 276, 301–306 (1991).
Huang, T.H., Perry, M.R. & Laux, D.E. Methylation profiling of CpG islands in human breast cancer cells. Hum. Mol. Genet. 8, 459–470 (1999).
Shi, H. et al. Expressed CpG island sequence tag microarray for dual screening of DNA hypermethylation and gene silencing in cancer cells. Cancer Res. 62, 3214–3220 (2002).
Plass, C. Cancer epigenomics. Hum. Mol. Genet. 11, 2479–2488 (2002).
Wang, T.L. et al. Digital karyotyping. Proc. Natl. Acad. Sci. USA 99, 16156–16161 (2002).
Dai, Z. et al. An AscI boundary library for the studies of genetic and epigenetic alterations in CpG islands. Genome Res. 12, 1591–1598 (2002).
Rhee, I. et al. DNMT1 and DNMT3b cooperate to silence genes in human cancer cells. Nature 416, 552–556 (2002).
Paz, M.F. et al. Genetic unmasking of epigenetically silenced tumor suppressor genes in colon cancer cells deficient in DNA methyltransferases. Hum. Mol. Genet. 12, 2209–2219 (2003).
Shigematsu, H. et al. Aberrant methylation of HIN-1 (high in normal-1) is a frequent event in many human malignancies. Int. J. Cancer 113, 600–604 (2005).
Krop, I. et al. Frequent HIN-1 promoter methylation and lack of expression in multiple human tumor types. Mol. Cancer Res. 2, 489–494 (2004).
Krop, I.E. et al. HIN-1, a putative cytokine highly expressed in normal but not cancerous mammary epithelial cells. Proc. Natl. Acad. Sci. USA 98, 9796–9801 (2001).
Feinberg, A.P. & Vogelstein, B. Hypomethylation distinguishes genes of some human cancers from their normal counterparts. Nature 301, 89–92 (1983).
Bocker, W. et al. Common adult stem cells in the human breast give rise to glandular and myoepithelial cell lineages: a new cell biological concept. Lab. Invest. 82, 737–746 (2002).
Kremenskoy, M. et al. Genome-wide analysis of DNA methylation status of CpG islands in embryoid bodies, teratomas, and fetuses. Biochem. Biophys. Res. Commun. 311, 884–890 (2003).
Caron, H. et al. The human transcriptome map: clustering of highly expressed genes in chromosomal domains. Science 291, 1289–1292 (2001).
Ushijima, T. Detection and interpretation of altered methylation patterns in cancer cells. Nat. Rev. Cancer 5, 223–231 (2005).
Bell, A.C. & Felsenfeld, G. Methylation of a CTCF-dependent boundary controls imprinted expression of the IGF2 gene. Nature 405, 482–485 (2000).
Saha, S. et al. Using the transcriptome to annotate the genome. Nat. Biotechnol. 20, 508–512 (2002).
Cai, L. et al. Clustering analysis of SAGE data using a Poisson approach. Genome Biol. 5, R51 (2004).
Herman, J.G., Graff, J.R., Myohanen, S., Nelkin, B.D. & Baylin, S.B. Methylation-specific PCR: a novel PCR assay for methylation status of CpG islands. Proc. Natl. Acad. Sci. USA 93, 9821–9826 (1996).
Acknowledgements
We thank A. Richardson, G. Lodeiro and R. Gomes for their help with the acquisition of human tissue samples; R. Gelman for help with statistical analysis; B. Vogelstein and K. Kinzler for providing genomic DNA from HCT116 wild-type and DKO cells and their continuous support and encouragement; and B. Vogelstein, I. Krop and other members of the laboratory of K.P. for critical reading of the manuscript and their constructive criticism throughout the execution of this project. This work was supported by the National Cancer Institute Specialized Program in Research Excellence in Breast Cancer at Dana-Farber/Harvard Cancer Center, by Department of Defense Breast Cancer Center of Excellence grants awarded to K.P. and by a Department of Defense Postdoctoral Fellowship grant awarded to J.Y.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
M.H. and K.P. submitted a patent application on the MSDK method and the findings described in this manuscript, in accordance with the policies of the Dana Farber Cancer Institute.
Supplementary information
Supplementary Fig. 1
SNP array analysis of genomic DNA samples used for MSDK analysis. (PDF 139 kb)
Supplementary Fig. 2
Genome-wide overview of the location of CpG islands, predicted AscI sites overlayed with methylation (MDSK), and gene expression (SAGE) patterns. (PDF 196 kb)
Supplementary Table 1
Statistical analysis of MSDK libraries generated from HCT116 WT and DKO cells. (PDF 143 kb)
Supplementary Table 2
List of breast tissue samples used for methylation assays. (PDF 81 kb)
Supplementary Table 3
Statistical analysis of MSDK libraries generated from normal (N-EPI-7) and tumor (I-EPI-7) breast epithelial cells. (PDF 128 kb)
Supplementary Table 4
Statistical analysis of MSDK libraries generated from normal (N-STR-I7) and tumor (I-STR-7) breast stromal cells. (PDF 142 kb)
Supplementary Table 5
Statistical analysis of MSDK libraries generated from normal (N-MYOEP-4) and DCIS tumor (D-MYOEP-6) breast myoepithelial cells. (PDF 134 kb)
Supplementary Table 6
Statistical analysis of MSDK libraries generated from normal breast myoepithelial (N-MYOEP-4) and epithelial (N-EPI-I7) cells. (PDF 136 kb)
Rights and permissions
About this article
Cite this article
Hu, M., Yao, J., Cai, L. et al. Distinct epigenetic changes in the stromal cells of breast cancers. Nat Genet 37, 899–905 (2005). https://doi.org/10.1038/ng1596
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng1596
This article is cited by
-
The expression of stromal biomarkers in small papillary thyroid carcinomas
World Journal of Surgical Oncology (2022)
-
The importance of being CAFs (in cancer resistance to targeted therapies)
Journal of Experimental & Clinical Cancer Research (2022)
-
Neuronal hyperexcitability drives central and peripheral nervous system tumor progression in models of neurofibromatosis-1
Nature Communications (2022)
-
Inference of epigenetic subnetworks by Bayesian regression with the incorporation of prior information
Scientific Reports (2022)
-
Signaling pathways in cancer-associated fibroblasts and targeted therapy for cancer
Signal Transduction and Targeted Therapy (2021)