Abstract
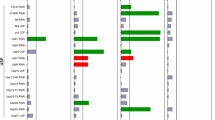
Huntington disease is a fatal neurodegenerative disorder caused by expansion of a polyglutamine tract in the protein huntingtin (Htt)1, which leads to its aggregation in nuclear and cytoplasmic inclusion bodies2. We recently identified 52 loss-of-function mutations in yeast genes that enhance the toxicity of a mutant Htt fragment3. Here we report the results from a genome-wide loss-of-function suppressor screen in which we identified 28 gene deletions that suppress toxicity of a mutant Htt fragment. The suppressors are known or predicted to have roles in vesicle transport, vacuolar degradation, transcription and prion-like aggregation. Among the most potent suppressors was Bna4 (kynurenine 3-monooxygenase), an enzyme in the kynurenine pathway of tryptophan degradation that has been linked directly to the pathophysiology of Huntington disease in humans by a mechanism that may involve reactive oxygen species4. This finding is suggestive of a conserved mechanism of polyglutamine toxicity from yeast to humans and identifies new candidate therapeutic targets for the treatment of Huntington disease.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
The Huntington's Disease Collaborative Research Group. A novel gene containing a trinucleotide repeat that is expanded and unstable on Huntington's disease chromosomes. Cell 72, 971–983 (1993).
Scherzinger, E. et al. Huntingtin-encoded polyglutamine expansions form amyloid-like protein aggregates in vitro and in vivo. Cell 90, 549–558 (1997).
Willingham, S., Outeiro, T.F., DeVit, M.J., Lindquist, S.L. & Muchowski, P.J. Yeast genes that enhance the toxicity of a mutant huntingtin fragment or alpha-synuclein. Science 302, 1769–1772 (2003).
Schwarcz, R. The kynurenine pathway of tryptophan degradation as a drug target. Curr. Opin. Pharmacol. 4, 12–17 (2004).
Meriin, A.B. et al. Huntington toxicity in yeast model depends on polyglutamine aggregation mediated by a prion-like protein Rnq1. J. Cell Biol. 157, 997–1004 (2002).
Muchowski, P.J., Ning, K., D'Souza-Schorey, C. & Fields, S. Requirement of an intact microtubule cytoskeleton for aggregation and inclusion body formation by a mutant huntingtin fragment. Proc. Natl. Acad. Sci. USA 99, 727–732 (2002).
Arrasate, M., Mitra, S., Schweitzer, E.S., Segal, M.R. & Finkbeiner, S. Inclusion body formation reduces levels of mutant huntingtin and the risk of neuronal death. Nature 431, 805–810 (2004).
Ravikumar, B., Duden, R. & Rubinsztein, D.C. Aggregate-prone proteins with polyglutamine and polyalanine expansions are degraded by autophagy. Hum. Mol. Genet. 11, 1107–1117 (2002).
Qin, Z.H. et al. Autophagy regulates the processing of amino terminal huntingtin fragments. Hum. Mol. Genet. 12, 3231–3244 (2003).
Ravikumar, B. et al. Inhibition of mTOR induces autophagy and reduces toxicity of polyglutamine expansions in fly and mouse models of Huntington disease. Nat. Genet. 36, 585–595 (2004).
Sugars, K.L. & Rubinsztein, D.C. Transcriptional abnormalities in Huntington disease. Trends Genet. 19, 233–238 (2003).
Steffan, J.S. et al. Histone deacetylase inhibitors arrest polyglutamine-dependent neurodegeneration in Drosophila. Nature 413, 739–743 (2001).
Hockly, E. et al. Suberoylanilide hydroxamic acid, a histone deacetylase inhibitor, ameliorates motor deficits in a mouse model of Huntington's disease. Proc. Natl. Acad. Sci. USA 100, 2041–2046 (2003).
Ferrante, R.J. et al. Histone deacetylase inhibition by sodium butyrate chemotherapy ameliorates the neurodegenerative phenotype in Huntington's disease mice. J. Neurosci. 23, 9418–9427 (2003).
Mallory, M.J. & Strich, R. Ume1p represses meiotic gene transcription in Saccharomyces cerevisiae through interaction with the histone deacetylase Rpd3p. J. Biol. Chem. 278, 44727–44734 (2003).
Michelitsch, M.D. & Weissman, J.S. A census of glutamine/asparagine-rich regions: implications for their conserved function and the prediction of novel prions. Proc. Natl. Acad. Sci. USA 97, 11910–11915 (2000).
Schwarcz, R., Whetsell, W.O. Jr. & Mangano, R.M. Quinolinic acid: an endogenous metabolite that produces axon-sparing lesions in rat brain. Science 219, 316–318 (1983).
Guidetti, P., Luthi-Carter, R.E., Augood, S.J. & Schwarcz, R. Neostriatal and cortical quinolinate levels are increased in early grade Huntington's disease. Neurobiol. Dis. 17, 455–461 (2004).
Guidetti, P. & Schwarcz, R. 3-Hydroxykynurenine potentiates quinolinate but not NMDA toxicity in the rat striatum. Eur. J. Neurosci. 11, 3857–3863 (1999).
Guidetti, P., Reddy, P.H., Tagle, D.A. & Schwarcz, R. Early kynurenergic impairment in Huntington's disease and in a transgenic animal model. Neurosci. Lett. 283, 233–235 (2000).
Goda, K., Hamane, Y., Kishimoto, R. & Ogishi, Y. Radical scavenging properties of tryptophan metabolites. Estimation of their radical reactivity. Adv. Exp. Med. Biol. 467, 397–402 (1999).
Rover, S., Cesura, A.M., Huguenin, P., Kettler, R. & Szente, A. Synthesis and biochemical evaluation of N-(4-phenylthiazol-2-yl)benzenesulfonamides as high-affinity inhibitors of kynurenine 3-hydroxylase. J. Med. Chem. 40, 4378–4385 (1997).
Moroni, F., Cozzi, A., Peruginelli, F., Carpenedo, R. & Pellegrini-Giampietro, D.E. Neuroprotective effects of kynurenine-3-hydroxylase inhibitors in models of brain ischemia. Adv. Exp. Med. Biol. 467, 199–206 (1999).
Richter, A. & Hamann, M. The kynurenine 3-hydroxylase inhibitor Ro 61-8048 improves dystonia in a genetic model of paroxysmal dyskinesia. Eur. J. Pharmacol. 478, 47–52 (2003).
Schwarcz, R. & Pellicciari, R. Manipulation of brain kynurenines: glial targets, neuronal effects, and clinical opportunities. J. Pharmacol. Exp. Ther. 303, 1–10 (2002).
Wyttenbach, A. et al. Heat shock protein 27 prevents cellular polyglutamine toxicity and suppresses the increase of reactive oxygen species caused by huntingtin. Hum. Mol. Genet. 11, 1137–1151 (2002).
Perez-Severiano, F. et al. Increased formation of reactive oxygen species, but no changes in glutathione peroxidase activity, in striata of mice transgenic for the Huntington's disease mutation. Neurochem. Res. 29, 729–733 (2004).
Sapp, E. et al. Early and progressive accumulation of reactive microglia in the Huntington disease brain. J. Neuropathol. Exp. Neurol. 60, 161–172 (2001).
Ryu, J.K., Kim, S.U. & McLarnon, J.G. Blockade of quinolinic acid-induced neurotoxicity by pyruvate is associated with inhibition of glial activation in a model of Huntington's disease. Exp. Neurol. 187, 150–159 (2004).
Zhang, X. et al. A potent small molecule inhibits polyglutamine aggregation in Huntington's disease neurons and suppresses neurodegeneration in vivo. Proc. Natl. Acad. Sci. USA 102, 892–897 (2005).
Acknowledgements
We thank M. Sherman for the pYES2-Htt25Q and pYES2-Htt103Q plasmids, S. Lindquist for the RNQ antibody and the pdr1Δpdr3Δ drug testing strain, W. Frostl for Ro 61-8048, R. Schwarcz for discussions about our data and advice regarding this project and K. Neireiter for his illustration. P.J.M. is supported by the US National Institute of Neurological Disease and Stroke, by a US National Institutes of Health construction award, by the Alzheimer's Disease Research Center at the University of Washington and by the Hereditary Disease Foundation under the auspices of the Cure Huntington's Disease Initiative. F.G. is supported by a postdoctoral fellowship from the HighQ foundation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Expression of Htt103Q and Rnq1 in Suppressor Strains. (PDF 986 kb)
Rights and permissions
About this article
Cite this article
Giorgini, F., Guidetti, P., Nguyen, Q. et al. A genomic screen in yeast implicates kynurenine 3-monooxygenase as a therapeutic target for Huntington disease. Nat Genet 37, 526–531 (2005). https://doi.org/10.1038/ng1542
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng1542
This article is cited by
-
Genome-wide screening in pluripotent cells identifies Mtf1 as a suppressor of mutant huntingtin toxicity
Nature Communications (2023)
-
Vitamin B6, B12 and folate modulate deregulated pathways and protein aggregation in yeast model of Huntington disease
3 Biotech (2023)
-
Modifier pathways in polyglutamine (PolyQ) diseases: from genetic screens to drug targets
Cellular and Molecular Life Sciences (2022)
-
Ablation of kynurenine 3-monooxygenase rescues plasma inflammatory cytokine levels in the R6/2 mouse model of Huntington’s disease
Scientific Reports (2021)
-
Full-length in meso structure and mechanism of rat kynurenine 3-monooxygenase inhibition
Communications Biology (2021)