Abstract
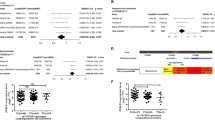
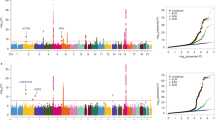
To identify genetic susceptibility loci for nasopharyngeal carcinoma (NPC), a genome-wide association study was performed using 464,328 autosomal SNPs in 1,583 NPC affected individuals (cases) and 1,894 controls of southern Chinese descent. The top 49 SNPs from the genome-wide association study were genotyped in 3,507 cases and 3,063 controls of southern Chinese descent from Guangdong and Guangxi. The seven supportive SNPs were further confirmed by transmission disequilibrium test analysis in 279 trios from Guangdong. We identified three new susceptibility loci, TNFRSF19 on 13q12 (rs9510787, Pcombined = 1.53 × 10−9, odds ratio (OR) = 1.20), MDS1-EVI1 on 3q26 (rs6774494, Pcombined = 1.34 × 10−8, OR = 0.84) and the CDKN2A-CDKN2B gene cluster on 9p21 (rs1412829, Pcombined = 4.84 × 10−7, OR = 0.78). Furthermore, we confirmed the role of HLA by revealing independent associations at rs2860580 (Pcombined = 4.88 × 10−67, OR = 0.58), rs2894207 (Pcombined = 3.42 × 10−33, OR = 0.61) and rs28421666 (Pcombined = 2.49 × 10−18, OR = 0.67). Our findings provide new insights into the pathogenesis of NPC by highlighting the involvement of pathways related to TNFRSF19 and MDS1-EVI1 in addition to HLA molecules.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Wei, W.I. & Sham, J.S. Nasopharyngeal carcinoma. Lancet 365, 2041–2054 (2005).
McDermott, A.L., Dutt, S.N. & Watkinson, J.C. The aetiology of nasopharyngeal carcinoma. Clin. Otolaryngol. Allied Sci. 26, 82–92 (2001).
Yu, M.C. & Yuan, J.M. Epidemiology of nasopharyngeal carcinoma. Semin. Cancer Biol. 12, 421–429 (2002).
Zeng, Y.X. & Jia, W.H. Familial nasopharyngeal carcinoma. Semin. Cancer Biol. 12, 443–450 (2002).
Li, D.J. et al. The dominance of China 1 in the spectrum of Epstein-Barr virus strains from Cantonese patients with nasopharyngeal carcinoma. J. Med. Virol. 81, 1253–1260 (2009).
Young, L.S. & Rickinson, A.B. Epstein-Barr virus: 40 years on. Nat. Rev. Cancer 4, 757–768 (2004).
Lu, S.J. et al. Linkage of a nasopharyngeal carcinoma susceptibility locus to the HLA region. Nature 346, 470–471 (1990).
Lu, C.C. et al. Nasopharyngeal carcinoma-susceptibility locus is localized to a 132 kb segment containing HLA-A using high-resolution microsatellite mapping. Int. J. Cancer 115, 742–746 (2005).
Feng, B.J. et al. Genome-wide scan for familial nasopharyngeal carcinoma reveals evidence of linkage to chromosome 4. Nat. Genet. 31, 395–399 (2002).
Xiong, W. et al. A susceptibility locus at chromosome 3p21 linked to familial nasopharyngeal carcinoma. Cancer Res. 64, 1972–1974 (2004).
Hu, L.F. et al. A genome-wide scan suggests a susceptibility locus on 5p 13 for nasopharyngeal carcinoma. Eur. J. Hum. Genet. 16, 343–349 (2008).
Tse, K.P. et al. Genome-wide association study reveals multiple nasopharyngeal carcinoma-associated loci within the HLA region at chromosome 6p21.3. Am. J. Hum. Genet. 85, 194–203 (2009).
Ng, C.C. et al. A genome-wide association study identifies ITGA9 conferring risk of nasopharyngeal carcinoma. J. Hum. Genet. 54, 392–397 (2009).
Chen, J. et al. Genetic structure of the Han Chinese population revealed by genome-wide SNP variation. Am. J. Hum. Genet. 85, 775–785 (2009).
Hildesheim, A. et al. Association of HLA class I and II alleles and extended haplotypes with nasopharyngeal carcinoma in Taiwan. J. Natl. Cancer Inst. 94, 1780–1789 (2002).
Skibola, C.F. et al. Genetic variants at 6p21.33 are associated with susceptibility to follicular lymphoma. Nat. Genet. 41, 873–875 (2009).
Rickinson, A.B. & Moss, D.J. Human cytotoxic T lymphocyte responses to Epstein-Barr virus infection. Annu. Rev. Immunol. 15, 405–431 (1997).
Hu, S., Tamada, K., Ni, J., Vincenz, C. & Chen, L. Characterization of TNFRSF19, a novel member of the tumor necrosis factor receptor superfamily. Genomics 62, 103–107 (1999).
Eby, M.T., Jasmin, A., Kumar, A., Sharma, K. & Chaudhary, P.M. TAJ, a novel member of the tumor necrosis factor receptor family, activates the c-Jun N-terminal kinase pathway and mediates caspase-independent cell death. J. Biol. Chem. 275, 15336–15342 (2000).
Morikawa, Y., Hisaoka, T., Kitamura, T. & Senba, E. TROY, a novel member of the tumor necrosis factor receptor superfamily in the central nervous system. Ann. NY Acad. Sci. 1126, A1–A10 (2008).
Eliopoulos, A.G. & Young, L.S. LMP1 structure and signal transduction. Semin. Cancer Biol. 11, 435–444 (2001).
Métais, J.Y. & Dunbar, C.E. The MDS1–EVI1 gene complex as a retrovirus integration site: impact on behavior of hematopoietic cells and implications for gene therapy. Mol. Ther. 16, 439–449 (2008).
Kurokawa, M. et al. The evi-1 oncoprotein inhibits c-Jun N-terminal kinase and prevents stress-induced cell death. EMBO J. 19, 2958–2968 (2000).
Nitta, E. et al. Oligomerization of Evi-1 regulated by the PR domain contributes to recruitment of corepressor CtBP. Oncogene 24, 6165–6173 (2005).
Xu, J., Menezes, J., Prasad, U. & Ahmad, A. Elevated serum levels of transforming growth factor beta1 in Epstein-Barr virus-associated nasopharyngeal carcinoma patients. Int. J. Cancer 84, 396–399 (1999).
Chou, J. et al. Nasopharyngeal carcinoma–review of the molecular mechanisms of tumorigenesis. Head Neck 30, 946–963 (2008).
Sharpless, N.E. et al. Loss of p16Ink4a with retention of p19Arf predisposes mice to tumorigenesis. Nature 413, 86–91 (2001).
Krimpenfort, P. et al. p15Ink4b is a critical tumour suppressor in the absence of p16Ink4a. Nature 448, 943–946 (2007).
Lo, K.W. & Huang, D.P. Genetic and epigenetic changes in nasopharyngeal carcinoma. Semin. Cancer Biol. 12, 451–462 (2002).
Wrensch, M. et al. Variants in the CDKN2B and RTEL1 regions are associated with high-grade glioma susceptibility. Nat. Genet. 41, 905–908 (2009).
Chen, M.C. et al. The incidence and risk of second primary cancers in patients with nasopharyngeal carcinoma: a population-based study in Taiwan over a 25-year period (1979–2003). Ann. Oncol. 19, 1180–1186 (2008).
Scélo, G. et al. Second primary cancers in patients with nasopharyngeal carcinoma: a pooled analysis of 13 cancer registries. Cancer Causes Control 18, 269–278 (2007).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Price, A.L. et al. Principal components analysis corrects for stratification in genome-wide association studies. Nat. Genet. 38, 904–909 (2006).
Barrett, J.C., Fry, B., Maller, J. & Daly, M.J. Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics 21, 263–265 (2005).
Xu, S. et al. Genomic dissection of population substructure of Han Chinese and its implication in association studies. Am. J. Hum. Genet. 85, 762–774 (2009).
Kazeem, G.R. & Farrall, M. Integrating case-control and TDT studies. Ann. Hum. Genet. 69, 329–335 (2005).
Agresti, A. Categorical Data Analysis (John Wiley and Sons, New York, 1990).
Stephens, M., Smith, N.J. & Donnelly, P. A new statistical method for haplotype reconstruction from population data. Am. J. Hum. Genet. 68, 978–989 (2001).
de Bakker, P.I. et al. A high-resolution HLA and SNP haplotype map for disease association studies in the extended human MHC. Nat. Genet. 38, 1166–1172 (2006).
Feng, B.J. et al. Multiple loci within the major histocompatibility complex confer risk of psoriasis. PLoS Genet. 5, e1000606 (2009).
Stephens, M. & Scheet, P. Accounting for decay of linkage disequilibrium in haplotype inference and missing-data imputation. Am. J. Hum. Genet. 76, 449–462 (2005).
Mensah, F.K. et al. Haplotype uncertainty in association studies. Genet. Epidemiol. 31, 348–357 (2007).
Marchini, J., Howie, B., Myers, S., McVean, G. & Donnelly, P. A new multipoint method for genome-wide association studies by imputation of genotypes. Nat. Genet. 39, 906–913 (2007).
Acknowledgements
We would like to thank all the subjects and healthy volunteers who participated in this work. We also want to thank the staffs of Bank of Tumor Resource at Sun Yat-sen University Cancer Center (SYSUCC) for help in sample storage. We also want to thank W.-Y. Meah, X. Chen, H.-B. Toh, K.-K. Heng, C.-H. Wong and D.E.K. Tan, who performed the genotyping, and R.T.H. Ong, J. Chen, K.-S. Sim and E. Tantoso for their assistance in data analysis. This work was funded by the High-Tech Research and Development Program of China (863 Plan, 2006AA02A404), the Natural Science Foundation of China (30621063, 30872929 and u0732005), the National Basic Research Program of China (973 Plan; 2004CB518604 and 2006CB910104), Beijing Science & Technology nova program (2006A54), the National Science and Technology Support Program of China (2006BAI02A11) and the Agency for Science, Technology and Research of Singapore.
Author information
Authors and Affiliations
Contributions
Y.-X.Z. was the overall study principal investigator who conceived the study and obtained financial support. Y.-X.Z., J.-X.B. and J.L. designed and oversaw the study. W.-H.J., B.-J.F., Q.-S.F. and L.-Z.C. initiated or participated in the recruitment of Guangdong Chinese samples and preparation of biological samples. H.Z., G.Z. and F.H. were responsible for the recruitment and sample preparation of Guangxi Chinese samples. J.-X.B. conducted sample inclusion and data management. B.-J.F. helped to impute the HLA alleles. J.-X.B. and Y.L. undertook the statistical analyses under guidance from J.L. and with help from H.-Q.L. J.-X.B. interpreted the results, drafted and synthesized the manuscript. E.T.L., T.K. and E.S.T. reviewed the manuscript and J.L. helped to revise it.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Note, Supplementary Tables 1–17 and Supplementary Figures 1–5 (PDF 2569 kb)
Supplementary Table 1
Summary of previous studies on the associations of HLA genes and nasopharyngeal carcinoma. (XLS 32 kb)
Supplementary Table 2
Summary of previous studies on the associations of non-MHC genes and nasopharyngeal carcinoma, based on candidate-gene approach. (XLS 34 kb)
Supplementary Table 3
Our GWAS results of the SNPs that were reported by the previously studies. (XLS 28 kb)
Supplementary Table 5
Association results of the 42 SNPs subjected to the validation analysis. (XLS 30 kb)
Supplementary Table 15
Susceptibility loci of different malignancies identified by GWAS up to March 9, 2010. (XLS 158 kb)
Rights and permissions
About this article
Cite this article
Bei, JX., Li, Y., Jia, WH. et al. A genome-wide association study of nasopharyngeal carcinoma identifies three new susceptibility loci. Nat Genet 42, 599–603 (2010). https://doi.org/10.1038/ng.601
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.601
This article is cited by
-
Nasopharyngeal carcinoma: nationwide trends in subtype-specific incidence and survival over 3 decades in a non-endemic area
Journal of Cancer Research and Clinical Oncology (2024)
-
The impact of HLA polymorphism on herpesvirus infection and disease
Immunogenetics (2023)
-
CACA guidelines for holistic integrative management of nasopharyngeal carcinoma
Holistic Integrative Oncology (2023)
-
A novel causal model for nasopharyngeal carcinoma
Cancer Causes & Control (2022)
-
A polygenic risk score for nasopharyngeal carcinoma shows potential for risk stratification and personalized screening
Nature Communications (2022)