Abstract
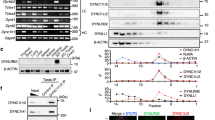
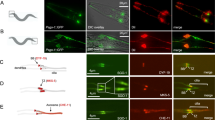
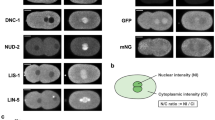
The spindle apparatus dictates the plane of cell cleavage, which is critical in the choice between symmetric or asymmetric division. Spindle positioning is controlled by an evolutionarily conserved pathway, which involves LIN-5/GPR-1/2/Gα in Caenorhabditis elegans, Mud/Pins/Gα in Drosophila and NuMA/LGN/Gα in humans1. GPR-1/2 and Gα localize LIN-5 to the cell cortex, which engages dynein and controls the cleavage plane during early mitotic divisions in C. elegans2,3,4,5,6. Here we identify ASPM-1 (abnormal spindle-like, microcephaly-associated) as a novel LIN-5 binding partner. ASPM-1, together with calmodulin (CMD-1), promotes meiotic spindle organization and the accumulation of LIN-5 at meiotic and mitotic spindle poles. Spindle rotation during maternal meiosis is independent of GPR-1/2 and Gα, yet requires LIN-5, ASPM-1, CMD-1 and dynein. Our data support the existence of two distinct LIN-5 complexes that determine localized dynein function: LIN-5/GPR-1/2/Gα at the cortex, and LIN-5/ASPM-1/CMD-1 at spindle poles. These functional interactions may be conserved in mammals, with implications for primary microcephaly.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Galli, M. & van den Heuvel, S. Determination of the cleavage plane in early C. elegans embryos. Annu. Rev. Genet. 42, 389–411 (2008).
Colombo, K. et al. Translation of polarity cues into asymmetric spindle positioning in Caenorhabditis elegans embryos. Science 300, 1957–1961 (2003).
Couwenbergs, C. et al. Heterotrimeric G protein signaling functions with dynein to promote spindle positioning in C. elegans. J. Cell Biol. 179, 15–22 (2007).
Gotta, M., Dong, Y., Peterson, Y. K., Lanier, S. M. & Ahringer, J. Asymmetrically distributed C. elegans homologs of AGS3/PINS control spindle position in the early embryo. Curr. Biol. 13, 1029–1037 (2003).
Nguyen-Ngoc, T., Afshar, K. & Gönczy, P. Coupling of cortical dynein and Gα proteins mediates spindle positioning in Caenorhabditis elegans. Nature Cell Biol. 9, 1294–1302 (2007).
Srinivasan, D. G., Fisk, R. M., Xu, H. & van den Heuvel, S. A complex of LIN-5 and GPR proteins regulates G protein signaling and spindle function in C. elegans. Genes Dev. 17, 1225–1239 (2003).
Albertson, D. G. & Thomson, J. N. Segregation of holocentric chromosomes at meiosis in the nematode, Caenorhabditis elegans. Chromosome Res. 1, 15–26 (1993).
Afshar, K. et al. RIC-8 is required for GPR-1/2-dependent Gα function during asymmetric division of C. elegans embryos. Cell 119, 219–230 (2004).
Lorson, M. A., Horvitz, H. R. & van den Heuvel, S. LIN-5 is a novel component of the spindle apparatus required for chromosome segregation and cleavage plane specification in Caenorhabditis elegans. J. Cell Biol. 148, 73–86 (2000).
Bond, J. et al. ASPM is a major determinant of cerebral cortical size. Nature Genet. 32, 316–320 (2002).
Fish, J. L., Kosodo, Y., Enard, W., Pääbo, S. & Huttner, W. B. Aspm specifically maintains symmetric proliferative divisions of neuroepithelial cells. Proc. Natl Acad. Sci. USA 103, 10438–10443 (2006).
Fisk Green, R., Lorson, M., Walhout, A. J., Vidal, M. & van den Heuvel, S. Identification of critical domains and putative partners for the Caenorhabditis elegans spindle component LIN-5. Mol. Genet. Genomics 271, 532–544 (2004).
Li, S. et al. A map of the interactome network of the metazoan C. elegans. Science 303, 540–543 (2004).
Park, D. H. & Rose, L. S. Dynamic localization of LIN-5 and GPR-1/2 to cortical force generation domains during spindle positioning. Dev. Biol. 315, 42–54 (2008).
Clark-Maguire, S. & Mains, P. E. Localization of the mei-1 gene product of Caenorhaditis elegans, a meiotic-specific spindle component. J. Cell Biol. 126, 199–209 (1994).
Yang, H. Y., Mains, P. E. & McNally, F. J. Kinesin-1 mediates translocation of the meiotic spindle to the oocyte cortex through KCA-1, a novel cargo adapter. J. Cell Biol. 169, 447–457 (2005).
Yang, H. Y., McNally, K. & McNally, F. J. MEI-1/katanin is required for translocation of the meiosis I spindle to the oocyte cortex in C. elegans. Dev. Biol. 260, 245–259 (2003).
O'Rourke, S. M., Dorfman, M. D., Carter, J. C. & Bowerman, B. Dynein modifiers in C. elegans: light chains suppress conditional heavy chain mutants. PLoS Genet. 3, e128 (2007).
Hamill, D. R., Severson, A. F., Carter, J. C. & Bowerman, B. Centrosome maturation and mitotic spindle assembly in C. elegans require SPD-5, a protein with multiple coiled-coil domains. Dev. Cell 3, 673–684 (2002).
Gassmann, R. et al. A new mechanism controlling kinetochore–microtubule interactions revealed by comparison of two dynein-targeting components: SPDL-1 and the Rod/Zwilch/Zw10 complex. Genes Dev. 22, 2385–2399 (2008).
do Carmo Avides, M. & Glover, D. M. Abnormal spindle protein, Asp, and the integrity of mitotic centrosomal microtubule organizing centers. Science 283, 1733–1735 (1999).
Wakefield, J. G., Bonaccorsi, S. & Gatti, M. The Drosophila protein asp is involved in microtubule organization during spindle formation and cytokinesis. J. Cell Biol. 153, 637–648 (2001).
Gonzalez, C. et al. Mutations at the asp locus of Drosophila lead to multiple free centrosomes in syncytial embryos, but restrict centrosome duplication in larval neuroblasts. J. Cell Sci. 96, 605–616 (1990).
Heald, R. et al. Self-organization of microtubules into bipolar spindles around artificial chromosomes in Xenopus egg extracts. Nature 382, 420–425 (1996).
Merdes, A., Heald, R., Samejima, K., Earnshaw, W. C. & Cleveland, D. W. Formation of spindle poles by dynein/dynactin-dependent transport of NuMA. J. Cell Biol. 149, 851–862 (2000).
Bowman, S. K., Neumuller, R. A., Novatchkova, M., Du, Q. & Knoblich, J. A. The Drosophila NuMA Homolog Mud regulates spindle orientation in asymmetric cell division. Dev. Cell 10, 731–742 (2006).
Du, Q. & Macara, I. G. Mammalian Pins is a conformational switch that links NuMA to heterotrimeric G proteins. Cell 119, 503–516 (2004).
Izumi, Y., Ohta, N., Hisata, K., Raabe, T. & Matsuzaki, F. Drosophila Pins-binding protein Mud regulates spindle-polarity coupling and centrosome organization. Nature Cell Biol. 8, 586–593 (2006).
Siller, K. H., Cabernard, C. & Doe, C. Q. The NuMA-related Mud protein binds Pins and regulates spindle orientation in Drosophila neuroblasts. Nature Cell Biol. 8, 594–600 (2006).
Mains, P. E., Kemphues, K. J., Sprunger, S. A., Sulston, I. A. & Wood, W. B. Mutations affecting the meiotic and mitotic divisions of the early Caenorhabditis elegans embryo. Genetics 126, 593–605 (1990).
Konno, D. et al. Neuroepithelial progenitors undergo LGN-dependent planar divisions to maintain self-renewability during mammalian neurogenesis. Nature Cell Biol. 10, 93–101 (2008).
Morin, X., Jaouen, F. & Durbec, P. Control of planar divisions by the G-protein regulator LGN maintains progenitors in the chick neuroepithelium. Nature Neurosci. 10, 1440–1448 (2007).
Sanada, K. & Tsai, L. H. G protein βγ subunits and AGS3 control spindle orientation and asymmetric cell fate of cerebral cortical progenitors. Cell 122, 119–131 (2005).
Boxem, M. et al. A protein domain-based interactome network for C. elegans early embryogenesis. Cell 134, 534–545 (2008).
Grill, S. W., Howard, J., Schäffer, E., Stelzer, E. H. & Hyman, A. A. The distribution of active force generators controls mitotic spindle position. Science 301, 518–521 (2003).
Acknowledgements
We are grateful to Steve Gygi and Ross Tomaino (Taplin Biological Mass Spectrometry, Harvard Medical School) for analysis of peptides by mass spectrometry (HMS), to Sean O'Rourke and Bruce Bowerman for the GFP–DHC-1 strain EU1561, to Inge The for assistance with ASPM-1 antibody production, Kristiaan de Vries for scoring embryonic lethality, and the Caenorhabditis Genetics Center (National Institutes of Health, National Center for Research Resources) for providing strains. We thank Adri Thomas and Marjolein Wildwater for critically reading the manuscript. This work was supported in part by National Institutes of Health grant GM57990 to S.v.d.H, Swiss National Science Foundation grant 3100A0-102087 to P.G., and Marie Curie International Reintegration grant MIRG-CT-2007-046458 to M.B.
Author information
Authors and Affiliations
Contributions
M.v.d.V. was primarily responsible for the experiments, data analysis and manuscript writing. C.B. performed IP-western experiments and fusion protein purification. A.P. contributed in the initial phase of the project. T.N-N. and P.G. performed spindle severing assays. M.B. performed Y2H experiments with support from M.V. S.v.d.H. contributed to experimental design, supervision and manuscript writing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Information (PDF 1852 kb)
Supplementary Information
Supplementary Movie 1 (MOV 1176 kb)
Supplementary Information
Supplementary Movie 2 (MOV 1418 kb)
Supplementary Information
Supplementary Movie 3 (MOV 1521 kb)
Supplementary Information
Supplementary Movie 4 (MOV 412 kb)
Supplementary Information
Supplementary Movie 5 (MOV 765 kb)
Supplementary Information
Supplementary Movie 6 (MOV 1322 kb)
Supplementary Information
Supplementary Movie 7 (MOV 2376 kb)
Supplementary Information
Supplementary Movie 8 (MOV 2162 kb)
Supplementary Information
Supplementary Movie 9 (MOV 640 kb)
Supplementary Information
Supplementary Movie 10 (MOV 2114 kb)
Supplementary Information
Supplementary Movie 11 (MOV 144 kb)
Supplementary Information
Supplementary Movie 12 (MOV 184 kb)
Rights and permissions
About this article
Cite this article
van der Voet, M., Berends, C., Perreault, A. et al. NuMA-related LIN-5, ASPM-1, calmodulin and dynein promote meiotic spindle rotation independently of cortical LIN-5/GPR/Gα. Nat Cell Biol 11, 269–277 (2009). https://doi.org/10.1038/ncb1834
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb1834
This article is cited by
-
Genetic basis of brain size evolution in cetaceans: insights from adaptive evolution of seven primary microcephaly (MCPH) genes
BMC Evolutionary Biology (2017)
-
Microtubule minus-end regulation at spindle poles by an ASPM–katanin complex
Nature Cell Biology (2017)
-
Chromosome segregation occurs by microtubule pushing in oocytes
Nature Communications (2017)
-
Distinct molecular cues ensure a robust microtubule-dependent nuclear positioning in the Drosophila oocyte
Nature Communications (2017)
-
A tissue-specific protein purification approach in Caenorhabditis elegans identifies novel interaction partners of DLG-1/Discs large
BMC Biology (2016)