Abstract
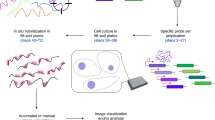
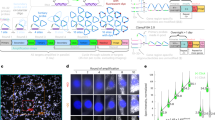
In situ hybridization methods enable the mapping of mRNA expression within intact biological samples1,2. With current approaches, it is challenging to simultaneously map multiple target mRNAs within whole-mount vertebrate embryos3,4,5,6, representing a significant limitation in attempting to study interacting regulatory elements in systems most relevant to human development and disease. Here, we report a multiplexed fluorescent in situ hybridization method based on orthogonal amplification with hybridization chain reactions (HCR)7. With this approach, RNA probes complementary to mRNA targets trigger chain reactions in which fluorophore-labeled RNA hairpins self-assemble into tethered fluorescent amplification polymers. The programmability and sequence specificity of these amplification cascades enable multiple HCR amplifiers to operate orthogonally at the same time in the same sample. Robust performance is achieved when imaging five target mRNAs simultaneously in fixed whole-mount and sectioned zebrafish embryos. HCR amplifiers exhibit deep sample penetration, high signal-to-background ratios and sharp signal localization.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Qian, X., Jin, L. & Lloyd, R.V. In situ hybridization: basic approaches and recent development. J. Histotechnol. 27, 53–67 (2004).
Silverman, A. & Kool, E. Oligonucleotide probes for RNA-targeted fluorescence in situ hybridization. Adv. Clin. Chem. 43, 79–115 (2007).
Thisse, B. et al. Spatial and temporal expression of the zebrafish genome by large-scale in situ hybridization screening. in Zebrafish: Genetics, Genomics and Informatics, vol. 77, edn. 2 (eds. Detrich, H.W., Zon, L.I. & Westerfield, M.) 505–519 (Elsevier, 2004).
Denkers, N., Garcia-Villalba, P., Rodesch, C.K., Nielson, K.R. & Mauch, T.J. FISHing for chick genes: Triple-label whole-mount fluorescence in situ hybridization detects simultaneous and overlapping gene expression in avian embryos. Dev. Dyn. 229, 651–657 (2004).
Barroso-Chinea, P. et al. Detection of two different mRNAs in a single section by dual in situ hybridization: A comparison between colorimetric and fluorescent detection. J. Neurosci. Methods 162, 119–128 (2007).
Acloque, H., Wilkinson, D.G. & Nieto, M.A. In situ hybridization analysis of chick embryos in whole-mount and tissue sections. in Avian Embryology, 2nd Edition vol. 87 (ed. Bronner-Fraser, M.) 169–185 (Elsevier, 2008).
Dirks, R.M. & Pierce, N.A. Triggered amplification by hybridization chain reaction. Proc. Natl. Acad. Sci. USA 101, 15275–15278 (2004).
Gall, J.G. & Pardue, M.L. Formation and detection of RNA-DNA hybrid molecules in cytological preparations. Proc. Natl. Acad. Sci. USA 63, 378–383 (1969).
Lawrence, J.B., Singer, R.H. & Marselle, L.M. Highly localized tracks of specific transcripts within interphase nuclei visualized by in situ hybridization. Cell 57, 493–502 (1989).
Tautz, D. & Pfeifle, C. A non-radioactive in situ hybridization method for the localization of specific RNAs in Drosophila embryos reveals translational control of the segmentation gene hunchback. Chromosoma 98, 81–85 (1989).
Kislauskis, E.H., Li, Z., Singer, R.H. & Taneja, K.L. Isoform-specific 3′-untranslated sequences sort α-cardiac and β-cytoplasmic actin messenger RNAs to different cytoplasmic compartments. J. Cell. Biol. 123, 165–172 (1993).
O'Neill, J.W. & Bier, E. Double-label in situ hybridization using biotin digoxigenin-tagged RNA probes. Biotechniques 17, 870–875 (1994).
Femino, A.M., Fay, F.S., Fogarty, K. & Singer, R.H. Visualization of single RNA transcripts in situ. Science 280, 585–590 (1998).
Zaidi, A.U., Enomoto, H., Milbrandt, J. & Roth, K.A. Dual fluorescent in situ hybridization and immunohistochemical detection with tyramide signal amplification. J. Histochem. Cytochem. 48, 1369–1375 (2000).
Player, A.N., Shen, L.-P., Kenny, D., Antao, V.P. & Kolberg, J.A. Single-copy gene detection using branched DNA (bDNA) in situ hybridization. J. Histochem. Cytochem. 49, 603–612 (2001).
Levsky, J.M., Shenoy, S.M., Pezo, R.C. & Singer, R.H. Single-cell gene expression profiling. Science 297, 836–840 (2002).
Kosman, D. et al. Multiplex detection of RNA expression in Drosophila embryos. Science 305, 846 (2004).
Lambros, M.B.K., Natrajan, R. & Reis-Filho, J.S. Chromogenic and fluorescent in situ hybridization in breast cancer. Hum. Pathol. 38, 1105–1122 (2007).
Raj, A., van den Bogaard, P., Rifkin, S.A., van Oudenaarden, A. & Tyagi, S. Imaging individual mRNA molecules using multiple singly labeled probes. Nat. Methods 5, 877–879 (2008).
Amann, R. & Fuchs, B.M. Single-cell identification in microbial communities by improved fluorescence in situ hybridization techniques. Nat. Rev. Microbiol. 6, 339–348 (2008).
Larsson, C., Grundberg, I., Soderberg, O. & Nilsson, M. In situ detection and genotyping of individual mRNA molecules. Nat. Methods 7, 395–397 (2010).
Harland, R.M. In situ hybridization: an improved whole-mount method for Xenopus embryos. Methods Cell Biol. 36, 685–695 (1991).
Speel, E.J.M., Hopman, A.H.N. & Komminoth, P. Amplification methods to increase the sensitivity of in situ hybridization: Play CARD(S). J. Histochem. Cytochem. 47, 281–288 (1999).
Feldkamp, U. & Niemeyer, C.M. Rational design of DNA nanoarchitectures. Angew Chem. Int. Ed. 45, 1856–1876 (2006).
Venkataraman, S., Dirks, R.M., Rothemund, P.W.K., Winfree, E. & Pierce, N.A. An autonomous polymerization motor powered by DNA hybridization. Nat. Nanotechnol. 2, 490–494 (2007).
Choi, H.M.T. Programmable In Situ Amplification for Multiplexed Bioimaging. PhD thesis, California Institute of Technology (2009).
Simmel, F.C. & Dittmer, W.U. DNA nanodevices. Small 1, 284–299 (2005).
Bath, J. & Turberfield, A.J. DNA nanomachines. Nat. Nanotechnol. 2, 275–284 (2007).
Feldkamp, U. & Niemeyer, C.M. Rational engineering of dynamic DNA systems. Angew. Chem. Int. Ed. 47, 3871–3873 (2008).
Yurke, B., Turberfield, A.J., Mills, J.A.P., Simmel, F.C. & Neumann, J.L. A DNA-fuelled molecular machine made of DNA. Nature 406, 605–608 (2000).
Dirks, R.M., Lin, M., Winfree, E. & Pierce, N.A. Paradigms for computational nucleic acid design. Nucleic Acids Res. 32, 1392–1403 (2004).
Zadeh, J.N., Wolfe, B.R. & Pierce, N.A. Nucleic acid sequence design via efficient ensemble defect optimization. J. Compu. Chem. published online, doi:10.1002/jcc.2163 (17 August 2010).
Zadeh, J.N. et al. NUPACK: analysis and design of nucleic acid systems. J. Compu. Chem. published online, doi:10.1002/jcc.21596 (19 July 2010).
Dirks, R.M., Bois, J.S., Schaeffer, J.M., Winfree, E. & Pierce, N.A. Thermodynamic analysis of interacting nucleic acid strands. SIAM Rev. 49, 65–88 (2007).
Acknowledgements
We thank V.A. Beck, J.S. Bois, S. Venkataraman, J.R. Vieregg and P. Yin for discussions. We thank C. Johnson and A.J. Ewald for performing preliminary studies. We thank J.N. Zadeh for the use of unpublished software. We thank the Caltech Biological Imaging Center and A. Collazo of the House Ear Institute for the use of multispectral confocal microscopes. This work was funded by the US National Institutes of Health (R01 EB006192 and P50 HG004071), the National Science Foundation (CCF-0448835 and CCF-0832824) and the Beckman Institute at Caltech.
Author information
Authors and Affiliations
Contributions
S.E.F. and N.A.P. conceived the application of HCR to multiplexed bioimaging; H.M.T.C., J.Y.C., J.E.P. and N.A.P. engineered HCR hairpins for use in stringent hybridization buffers; H.M.T.C. and N.A.P. designed the experiments; H.M.T.C. performed the experiments; L.A.T. selected targets, provided technical guidance and performed the control experiments using traditional in situ hybridization; H.M.T.C., L.A.T., S.E.F. and N.A.P. analyzed the data; H.M.T.C. and N.A.P. wrote the manuscript; and all authors edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare competing financial interests in the form of US patents and pending US and EU patents.
Supplementary information
Supplementary Text and Figures
Supplementary Notes (PDF 23142 kb)
Supplementary Movie 1
Image stack for Figure 3b (AVI 8342 kb)
Supplementary Movie 2
Image stack for Figure 3d (AVI 4130 kb)
Rights and permissions
About this article
Cite this article
Choi, H., Chang, J., Trinh, L. et al. Programmable in situ amplification for multiplexed imaging of mRNA expression. Nat Biotechnol 28, 1208–1212 (2010). https://doi.org/10.1038/nbt.1692
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.1692
This article is cited by
-
A rapid and sensitive, multiplex, whole mount RNA fluorescence in situ hybridization and immunohistochemistry protocol
Plant Methods (2023)
-
A one-pot isothermal Cas12-based assay for the sensitive detection of microRNAs
Nature Biomedical Engineering (2023)
-
Quantitative whole-tissue 3D imaging reveals bacteria in close association with mouse jejunum mucosa
npj Biofilms and Microbiomes (2023)
-
Spatially resolved in vivo imaging of inflammation-associated mRNA via enzymatic fluorescence amplification in a molecular beacon
Nature Biomedical Engineering (2022)
-
Nucleic acid-based fluorescent sensor systems: a review
Polymer Journal (2022)