Abstract
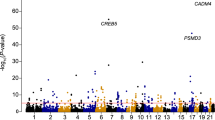
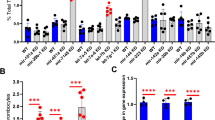
Platelets are the second most abundant cell type in blood and are essential for maintaining haemostasis. Their count and volume are tightly controlled within narrow physiological ranges, but there is only limited understanding of the molecular processes controlling both traits. Here we carried out a high-powered meta-analysis of genome-wide association studies (GWAS) in up to 66,867 individuals of European ancestry, followed by extensive biological and functional assessment. We identified 68 genomic loci reliably associated with platelet count and volume mapping to established and putative novel regulators of megakaryopoiesis and platelet formation. These genes show megakaryocyte-specific gene expression patterns and extensive network connectivity. Using gene silencing in Danio rerio and Drosophila melanogaster, we identified 11 of the genes as novel regulators of blood cell formation. Taken together, our findings advance understanding of novel gene functions controlling fate-determining events during megakaryopoiesis and platelet formation, providing a new example of successful translation of GWAS to function.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Frazer, K. A. et al. A second generation human haplotype map of over 3.1 million SNPs. Nature 449, 851–861 (2007)
Devlin, B. & Roeder, K. Genomic control for association studies. Biometrics 55, 997–1004 (1999)
Lo, K. S. et al. Genetic association analysis highlights new loci that modulate hematological trait variation in Caucasians and African Americans. Hum. Genet. 129, 307–317 (2011)
Meisinger, C. et al. A genome-wide association study identifies three loci associated with mean platelet volume. Am. J. Hum. Genet. 84, 66–71 (2009)
Soranzo, N. et al. A novel variant on chromosome 7q22.3 associated with mean platelet volume, counts, and function. Blood 113, 3831–3837 (2009)
Soranzo, N. et al. A genome-wide meta-analysis identifies 22 loci associated with eight hematological parameters in the HaemGen consortium. Nature Genet. 41, 1182–1190 (2009)
Bath, P. M. & Butterworth, R. J. Platelet size: measurement, physiology and vascular disease. Blood Coagul. Fibrinolysis 7, 157–161 (1996)
Kamatani, Y. et al. Genome-wide association study of hematological and biochemical traits in a Japanese population. Nature Genet. 42, 210–215 (2010)
Manolio, T. A. et al. Finding the missing heritability of complex diseases. Nature 461, 747–753 (2009)
Park, J.-H. et al. Estimation of effect size distribution from genome-wide association studies and implications for future discoveries. Nature Genet. 42, 570–575 (2010)
Lango Allen, H. et al. Hundreds of variants clustered in genomic loci and biological pathways affect human height. Nature 467, 832–838 (2010)
Teslovich, T. et al. Biological, clinical and population relevance of 95 loci for blood lipids. Nature 466, 707–713 (2010)
Paul, D. S. et al. Maps of open chromatin guide the functional follow-up of genome-wide association signals: application to hematological traits. PLoS Genet. 7, e1002139 (2011)
Watkins, N. A. et al. A HaemAtlas: characterizing gene expression in differentiated human blood cells. Blood 113, 1–9 (2009)
Lu, J. et al. MicroRNA-mediated control of cell fate in megakaryocyte-erythrocyte progenitors. Dev. Cell 14, 843–853 (2008)
Crozatier, M. & Meister, M. Drosophila haematopoiesis. Cell. Microbiol. 9, 1117–1126 (2007)
Fossett, N. et al. The Friend of GATA proteins U-shaped, FOG-1, and FOG-2 function as negative regulators of blood, heart, and eye development in Drosophila. Proc. Natl Acad. Sci. USA 98, 7342–7347 (2001)
Goldfarb, A. N. Transcriptional control of megakaryocyte development. Oncogene 26, 6795–6802 (2007)
Gregory, G. D. et al. FOG1 requires NuRD to promote hematopoiesis and maintain lineage fidelity within the megakaryocytic-erythroid compartment. Blood 115, 2156–2166 (2010)
Wendling, F. et al. cMpl ligand is a humoral regulator of megakaryocytopoiesis. Nature 369, 571–574 (1994)
Serbanovic-Canic, J. et al. Silencing of RhoA nucleotide exchange factor, ARHGEF3 reveals its unexpected role in iron uptake. Blood 118, 4967–4976 (2011)
Chu, S. G. et al. Mean platelet volume as a predictor of cardiovascular risk: a systematic review and meta-analysis. J. Thromb. Haemost. 8, 148–156 (2010)
Klovaite, J., Benn, M., Yazdanyar, S. & Nordestgaard, B. G. High platelet volume and increased risk of myocardial infarction: 39,531 participants from the general population. J. Thromb. Haemost. 9, 49–56 (2010)
Garner, C. et al. Two candidate genes for low platelet count identified in an Asian Indian kindred by genome-wide linkage analysis: glycoprotein IX and thrombopoietin. Eur. J. Hum. Genet. 14, 101–108</lpage> (2006)
Cline, M. S. et al. Integration of biological networks and gene expression data using Cytoscape. Nature Protocols 2, 2366–2382 (2007)
Acknowledgements
Reactome is supported by grants from the US National Institutes of Health (P41 HG003751), EU grant LSHG-CT-2005-518254 ‘ENFIN’, and the EBI Industry Programme. A.C. and D.L.S. are supported by the Wellcome Trust grants WT 082597/Z/07/Z and WT 077037/Z/05/Z and WT 077047/Z/05/Z, respectively. J.S.C. and K.V. are supported by the European Union NetSim training fellowship scheme (number 215820). A. Radhakrishnan, A.A., H.L.J., J.J., J.S. and W.H.O. are funded by the National Institute for Health Research, UK. Augusto Rendon is funded by programme grant RG/09/012/28096 from the British Heart Foundation. J.M.P. is supported by an Advanced ERC grant. IntAct is funded by the EC under SLING, grant agreement no. 226073 (Integrating Activity) within Research Infrastructures of the FP7, under PSIMEX, contract no. FP7-HEALTH-2007-223411. C.G. received support by the European Community’s Seventh Framework Programme (FP7/2007-2013), ENGAGE Consortium, grant agreement HEALTH-F4-2007- 201413. N.S. is supported by the Wellcome Trust (Grant Number 098051). Acknowledgements by individual participating studies are provided in Supplementary Information.
Author information
Authors and Affiliations
Contributions
Study design group: C.G., S. Sanna, A.A.H., A. Rendon, M.A.F., W.H.O., N.S.; manuscript writing group: C.G., A. Radhakrishnan, S. Sanna, A.A.H., A. Rendon, M.A.F., W.H.O., N.S.; data preparation, meta-analysis and secondary analysis group: A. Radhakrishnan, B.K., W.T., E.P., G.P., R.M., M.A.F., C.G., N.S.; bioinformatics analyses, pathway analyses and protein–protein interaction network group: S.M., J.-W.N.A., S.J., J.K., Y.M., L.B., A. Rendon, W.H.O.; transcript profiling methods and data group: K.V., A. Rendon, L.W., A.H.G., T.-P.Y., F. Cambien, J.E., C. Hengstenberg, N.J.S., H. Schunkert, P.D., W.H.O.; M. musculus models: R.R.-S.; D. rerio knockdown models: A.C., J.S.-C., D. Stemple, W.H.O.; D. melanogaster knockdown models: U.E., J. Penninger and A.A.H. All other author contributions and roles are listed in Supplementary Information.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
The file contains Supplementary Text, Supplementary References, Supplementary Acknowledgements and Author Contributions, Supplementary Tables 2-8 (see separate file for Supplementary Table 1), Supplementary Data and Supplementary Figures 1-6 - see the contents page for more details. This file was replaced online on 02 April 2012 as the Supplementary Tables and supplementary Figures were missing from the original file. (PDF 9853 kb)
Supplementary Table 1
The table shows a summary of population descriptives, trait measurements and details of genotyping and analysis metrics and methods. This file was replaced on line on 13 April 2012 as the file originally posted on line erroneously contained Supplementary Tables 2-8. (XLS 44 kb)
Supplementary Data
The file shows Protein-protein interaction network, containing gene expression levels and other annotation features. This file was removed from our website on 13 April 2012 as the original file posted was corrupted. To download this information please go to the following link - https://haemgen.haem.cam.ac.uk/files/. (ZIP 84 kb)
PowerPoint slides
Rights and permissions
About this article
Cite this article
Gieger, C., Radhakrishnan, A., Cvejic, A. et al. New gene functions in megakaryopoiesis and platelet formation. Nature 480, 201–208 (2011). https://doi.org/10.1038/nature10659
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10659
This article is cited by
-
Deciphering the omicron variant: integrated omics analysis reveals critical biomarkers and pathophysiological pathways
Journal of Translational Medicine (2024)
-
Jeanette Erdmann (1965–2023)
Nature Genetics (2023)
-
Determinants of Variable Total Platelet Count in Healthy Plateletpheresis Donor
Indian Journal of Hematology and Blood Transfusion (2023)
-
Genomic Variation Affecting MPV and PLT Count in Association with Development of Ischemic Stroke and Its Subtypes
Molecular Neurobiology (2023)
-
Physiological and pathophysiological functions of NLRP6: pro- and anti-inflammatory roles
Communications Biology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.