Abstract
Neurofibromatosis type 1 (NF1) is caused by dominant loss-of-function mutations of the tumor suppressor NF1 containing 57 constitutive coding exons. A huge number of different pathogenic NF1 alterations has been reported. The aim of the present study was to evaluate the usefulness of a multiplex ligation-dependent probe amplification (MLPA) approach in NF1 patients to detect single and multi-exon NF1 gene copy number variations. A genotype–phenotype correlation was then performed in NF1 patients carrying these types of genetic alterations. Among 565 NF1 index cases from the French NF1 cohort, single and multi-exon deletions/duplications screening identified NF1 partial deletions/duplications in 22 patients (~4%) using MLPA analysis. Eight single exon deletions, 11 multiple exons deletions, 1 complex rearrangement and 2 duplications were identified. All results were confirmed using a custom array-CGH. MLPA and custom array-CGH allowed the identification of rearrangements that were missed by cDNA/DNA sequencing or microsatellite analysis. We then performed a targeted next-generation sequencing of NF1 that allowed confirmation of all 22 rearrangements. No clear genotype–phenotype correlations were found for the most clinically significant disease features of NF1 in patients with single and multi-exons NF1 gene copy number changes.
Similar content being viewed by others
Main
Neurofibromatosis type 1 (NF1, OMIM# 162200) is an autosomal tumor predisposition disorder with an estimated birth incidence of 1 in 3500.1 NF1 is caused by dominant loss-of-function mutations of the tumor suppressor NF1 (Neurofibromin 1; OMIM# 613113), located at 17q11.2 and containing 57 constitutive coding exons and three alternative spliced exons over ~280 kb. Almost half of all NF1 cases are caused by de novo sporadic mutations. A huge number of different pathogenic NF1 alterations have been reported.2, 3, 4, 5, 6, 7 Five to ten percents of NF1 patients present large 17q11.2 deletions associated with a more severe phenotype, the so-called ‘NF1 microdeletion syndrome’.8, 9, 10 For patients with intragenic NF1 mutations (> 90% of all NF1 cases), no clear-cut allele–phenotype correlations have been established so far11, 12, 13, 14 with the exception of a 3-bp inframe deletion associated with the absence of cutaneous neurofibromas.15 To date, few reports have estimated the contribution of single and multi-exon NF1 gene copy number changes to the NF1 mutation spectrum.6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18
The aim of the present study was (i) to evaluate the usefulness of a multiplex ligation-dependent probe amplification (MLPA) approach in NF1 patients and (ii) to perform a genotype–phenotype correlation in NF1 patients carrying these type of genetic alterations from the large NF French cohort. We then assessed whether all identified NF1 gene copy number changes could be detected by using a recently developed targeted next-generation sequencing (NGS) pipeline for NF1 screening.
The French NF1 database was previously described.14 Briefly, it represented a collection of 565 families, consisting of 1697 individuals among whom 1083 fulfilled National Institutes of Health diagnostic criteria for NF1.19 Single and multi-exon deletions/duplications screening was performed by MLPA analysis using the SALSA MLPA kits P081/P082 NF1, as recommended in the manufacturer’s protocol (MRC Holland, Amsterdam, the Netherlands). The two probe mixes included in this MLPA kit contain probes for all constitutive NF1 exons, except in exons 7, 9, 22, 23, 54 and 56. One probe is also present in the NF1 promoter and in intron 1. MLPA analysis identified NF1 partial deletions/duplications in ~4% (n=22/565) of all index cases, confirming previous literature data.6, 16, 17, 18 The 22 NF1 partial deletions/duplications are detailed in Table 1 and Figure 1. Eight single exon deletions (including four restricted deletions of the NF1 promoter and exon 1), 11 multiple exons deletions (including one mosaic exon 1–35 deletion in patient NF00587; Supplementary Figure S1), 1 complex rearrangement and 2 duplications were found. We analyzed the two NF1 intragenic duplications at the cDNA level in patients NF00735 and NF00966. The two duplications were found to be in a direct tandem orientation for both exons 28–29 and exons 49–57 duplications in patients NF00735 and NF00966, respectively (Supplementary Figures S3 and S4).
Location of the 22 NF1 single or multi-exons deletions in the NF1 gene (usual nomenclature). Exons are named according to NCBI nomenclature (exons numbered 1-58) and represented by rectangles (not proportional to their size). Vertical arrows indicate the location of microsatellites used to search large deletion (introns 1, 36 and 47). Horizontal lines F1 to F8 represent the RT-PCR fragments used for the sequencing of the NF1 gene transcript. Duplications and deletions are shown above and below the NF1 gene, respectively. NF1 patient references are indicated below the corresponding duplicated or deleted segments. Affected exons are indicated above the corresponding duplicated or deleted segments.
Among the 22 deletions/duplications, three were identified by microsatellites pre-screening before any sequencing investigation. Among the 19 remaining deletions/duplications, only seven were detected by NF1 cDNA study. Undetected rearrangements included at least one of the two primers used for the cDNA PCR, causing an absence of amplification of the mutated alleles. All results were confirmed using a high-resolution NF1 custom array-CGH targeting the 17q11.2 region, allowing characterization of the duplication/deletion boundaries and suggesting recurrent breakpoints (Supplementary Figure S1). The theoretical and practical aspects of this custom array-CGH are described in detail elsewhere.20 Array-CGH showed that the promoter-exon 1 deletion profile was the same for patients NF00173 and NF00416, suggesting a recurrent molecular mechanism or a familial unknown link. According to array CGH analyses, no additional gene was included in the deletions exceeding the NF1 gene locus in 5′ (NF00173, NF00416, NF00485, NF00587, NF01745) or 3′ (NF00162, NF00911, NF01693) directions.
NF1 sequencing at both cDNA and DNA levels enabled the detection of a broader mutation spectrum than any single level approach.14 However, MLPA and/or custom array-CGH allowed the identification of rearrangements that were missed by cDNA/DNA sequencing or microsatellite analysis (Table 1). In case of negative results, these last approaches should therefore be completed by the screening for single or multi-NF1 exons copy number variation using MLPA or array-CGH. These results led us to reconsider our strategy for molecular study of the NF1 gene (Supplementary Figure S2). In the first step, whole NF1 deletions are now screened using microsatellites analysis. Sequencing at cDNA and then DNA levels is performed when no whole deletion is found. MLPA is performed in case of negative screening at cDNA and DNA approaches.
We recently developed a targeted NGS of the NF1 gene using a multiplex PCR approach on a PGM sequencer (Ampliseq, Life Technologies).21 All the 22 rearrangements were confirmed using this targeted NGS approach used as a quantitative method by comparing the number of reads for each amplicon of each sample, as previously described (Supplementary Figure S5).21 A mosaic NF1 deletion was confirmed for one (NF00587) of the six patients’ samples with de novo rearrangements. However, it cannot be excluded that mosaic may be restricted to other tissues types than peripheral blood for the other five samples from sporadic index cases. Targeted-NGS allowed simultaneous NF1 sequencing and provided a quantitative aspect allowing the detection of exons copy number alteration. This coupled analysis provides information of both the sequence and copy number alterations and avoids the sequential implementation of different techniques for the identification of these two different types of alterations. For laboratories using NGS in routine molecular diagnostic, MLPA may thus be used as a second line confirmation test for samples with suspected deletion or duplication.
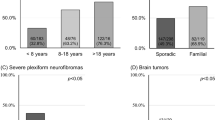
As the great majority of NF1 alterations are private mutations, genotype–phenotype correlations in NF1 can be more readily examined by categorizing NF1 mutations on the basis of the type of molecular defect. Genotype–phenotype correlations were statistically evaluated by comparing the 22 NF1 patients harboring NF1 exon deletions/duplications with a reference group composed of 439 unrelated patients with intragenic NF1 mutations from the NF-France database.14 All patients were clinically evaluated using the same standardized core questionnaire. Twelve major clinical features of NF1 were considered for the genotype–phenotype association study. Five were quantitative traits: number of café-au-lait (CAL) spots of small and large size, and number of plexiform, cutaneous and subcutaneous neurofibromas. The seven other clinical features were: skin-fold freckling, blue-red macules, Lisch nodules, facial dysmorphism, scoliosis, optic gliomas and learning disabilities. The 22 NF1 patients harboring single or multi-exon deletions and duplications in the NF1 gene consisted of 13 men (59%) and 9 women (41%) with a mean age at examination of 28.9±16.8 years (Supplementary Table S1). Around 27% (6/22) of patients had de novo mutations, adjudged based on the absence of a clinically affected parent. The number or prevalence of each of 12 major clinical features of NF1 were contrasted with those observed in a reference group composed of NF1 patients carrying either a truncating, missense or in-frame splice NF1 mutation (N=439). A comparison with the more homogeneous subgroup of patients harboring truncating mutations was also performed (Supplementary Table S1). To our knowledge, only a few previous studies have investigated the influence of the type of constitutional NF1 mutation on the disease phenotypic variability.11, 12, 13 The association of the type of NF1 mutation with each of the 12 clinical features individually was investigated through multiple regression analysis. All regression models included age at examination (as a continuous variable), gender and mutation inheritance (de novo versus inherited) as possible explanatory variables to control for potential confounding. Although patients with exon deletions/duplications exhibited a higher prevalence of scoliosis (55 versus 38%) and learning disabilities (59 versus 46%) when compared with either the general NF1 population or the subgroup of patients carrying truncating mutations (Supplementary Table S2), our analysis did not reveal any significant association with any of the 12 clinical features considered at either a Bonferroni-corrected or nominal significance threshold (P<0.05). These results are consistent with our previous findings showing a limited contribution of the allelic heterogeneity of the constitutional NF1 mutation to the variable expressivity of the disease,13 with the exception of those patients with the NF1 microdeletion syndrome.8, 9 The knowledge of the NF1 mutation type is not expected to provide any valuable prognostic information to patients and clinicians in the future, suggesting that genetic modifiers contribute to the variable expression of NF1.22, 23
References
Carey, J. C ., Baty, B. J ., Johnson, J. P ., Morrison, T ., Skolnick, M . & Kivlin, J . The genetic aspects of neurofibromatosis. Ann. NY Acad. Sci. 486, 45–56 (1986).
Fahsold, R ., Hoffmeyer, S ., Mischung, C ., Gille, C ., Ehlers, C ., Kücükceylan, N . et al. Minor lesion mutational spectrum of the entire NF1 gene does not explain its high mutability but points to a functional domain upstream of the GAP-related domain. Am. J. Hum. Genet. 66, 790–818 (2000).
Messiaen, L. M ., Callens, T ., Mortier, G ., Beysen, D ., Vandenbroucke, I ., Van Roy, N . et al. Exhaustive mutation analysis of the NF1 gene allows identification of 95% of mutations and reveals a high frequency of unusual splicing defects. Hum. Mutat. 15, 541–555 (2000).
Ars, E ., Kruyer, H ., Morell, M ., Pros, E ., Serra, E ., Ravella, A . et al. Recurrent mutations in the NF1 gene are common among neurofibromatosis type 1 patients. J. Med. Genet. 40, e82 (2003).
Mattocks, C ., Baralle, D ., Tarpey, P ., ffrench-Constant, C ., Bobrow, M . & Whittaker, J . Automated comparative sequence analysis identifies mutations in 89% of NF1 patients and confirms a mutation cluster in exons 11-17 distinct from the GAP related domain. J. Med. Genet. 41, e48 (2004).
Wimmer, K ., Yao, S ., Claes, K ., Kehrer-Sawatzki, H ., Tinschert, S ., De Raedt, T . et al. Spectrum of single- and multiexon NF1 copy number changes in a cohort of 1,100 unselected NF1 patients. Genes Chromosomes Cancer 45, 265–276 (2006).
Pros, E ., Gómez, C ., Martín, T ., Fábregas, P ., Serra, E . & Lázaro, C . Nature and mRNA effect of 282 different NF1 point mutations: focus on splicing alterations. Hum. Mutat. 29, E173–E193 (2008).
Mautner, V. F ., Kluwe, L ., Friedrich, R. E ., Roehl, A. C ., Bammert, S ., Högel, J . et al. Clinical characterisation of 29 neurofibromatosis type-1 patients with molecularly ascertained 1.4 Mb type-1 NF1 deletions. J. Med. Genet. 47, 623–630 (2008).
Pasmant, E ., Sabbagh, A ., Spurlock, G ., Laurendeau, I ., Grillo, E . & Hamel, M. J ., members of the NF France Network NF1 microdeletions in neurofibromatosis type 1: from genotype to phenotype. Hum. Mutat. 31, E1506–E1518 (2010).
De Raedt, T ., Brems, H ., Wolkenstein, P ., Vidaud, D ., Pilotti, S ., Perrone, F . et al. Elevated risk for MPNST in NF1 microdeletion patients. Am. J. Hum. Genet. 72, 1288–1292 (2003).
Easton, D. F ., Ponder, M. A ., Huson, S. M . & Ponder, B. A . An analysis of variation in expression of neurofibromatosis (NF) type 1 (NF1): evidence for modifying genes. Am. J. Hum. Genet. 53, 305–313 (1993).
Castle, B ., Baser, M. E ., Huson, S. M ., Cooper, D. N . & Upadhyaya, M . Evaluation of genotype-phenotype correlations in neurofibromatosis type 1. J. Med. Genet. 40, e109 (2003).
Sabbagh, A ., Pasmant, E ., Laurendeau, I ., Parfait, B ., Barbarot, S ., Guillot, B . et al. Unravelling the genetic basis of variable clinical expression in neurofibromatosis 1. Hum. Mol. Genet. 18, 2768–2778 (2009).
Sabbagh, A ., Pasmant, E ., Imbard, A ., Luscan, A ., Soares, M ., Blanché, H . et al. NF1 molecular characterization and neurofibromatosis type I genotype-phenotype correlation: the French experience. Hum. Mutat. 34, 1510–1518 (2013).
Upadhyaya, M ., Huson, S. M ., Davies, M ., Thomas, N ., Chuzhanova, N ., Giovannini, S . et al. An absence of cutaneous neurofibromas associated with a 3- bp inframe deletion in exon 17 of the NF1 gene (c.2970–2972 delAAT): evidence of a clinically significant NF1 genotype-phenotype correlation. Am. J. Hum. Genet. 80, 140–151 (2007).
De Luca, A ., Schirinzi, A ., Buccino, A ., Bottillo, I ., Sinibaldi, L ., Torrente, I . et al. Novel and recurrent mutations in the NF1 gene in Italian patients with neurofibromatosis type 1. Hum. Mutat. 23, 629 (2004).
De Luca, A ., Bottillo, I ., Dasdia, M. C ., Morella, A ., Lanari, V ., Bernardini, L . et al. Deletions of NF1 gene and exons detected by multiplex ligation-dependent probe amplification. J. Med. Genet. 44, 800–808 (2007).
Valero, M. C ., Martín, Y ., Hernández-Imaz, E ., Marina Hernández, A ., Meleán, G ., Valero, A. M . et al. A highly sensitive genetic protocol to detect NF1 mutations. J. Mol. Diagn. 13, 113–122 (2011).
National Institutes of Health Consensus Development Conference Statement Neurofibromatosis. Arch. Neurol. 45, 575–578 (1988).
Pasmant, E ., Sabbagh, A ., Masliah-Planchon, J ., Haddad, V ., Hamel, M. J ., Laurendeau, I . et al. Detection and characterization of NF1 microdeletions by custom high resolution array CGH. J. Mol. Diagn. 11, 524–529 (2009).
Pasmant, E ., Parfait, B ., Luscan, A ., Goussard, P ., Briand-Suleau, A ., Laurendeau, I . et al. Neurofibromatosis type 1 molecular diagnosis: what can NGS do for you when you have a large gene with loss of function mutations? Eur. J. Hum. Genet. (e-pub ahead of print 30 July 2014).
Pasmant, E ., Sabbagh, A ., Masliah-Planchon, J ., Ortonne, N ., Laurendeau, I ., Melin, L . et al. Role of noncoding RNA ANRIL in genesis of plexiform neurofibromas in neurofibromatosis type 1. J. Natl Cancer. Inst. 103, 1713–1722 (2011).
Pasmant, E ., Vidaud, M ., Vidaud, D . & Wolkenstein, P . Neurofibromatosis type 1: from genotype to phenotype. J. Med. Genet. 49, 483–489 (2012).
Acknowledgements
This work was supported in part by grants from Association Neurofibromatoses et Recklinghausen, Ligue Française Contre les Neurofibromatoses, the French Clinical Research program (PHRC 2002), INSERM (Nf1GeneModif project) and Ministère de l’Enseignement Supérieur et de la Recherche.
Author information
Authors and Affiliations
Consortia
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on Journal of Human Genetics website
Supplementary information
Rights and permissions
About this article
Cite this article
Imbard, A., Pasmant, E., Sabbagh, A. et al. NF1 single and multi-exons copy number variations in neurofibromatosis type 1. J Hum Genet 60, 221–224 (2015). https://doi.org/10.1038/jhg.2015.6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2015.6
This article is cited by
-
Genetic diagnosis of neurofibromatosis type 1: targeted next- generation sequencing with Multiple Ligation-Dependent Probe Amplification analysis
Journal of Biomedical Science (2018)
-
Alternative outcomes of pathogenic complex somatic structural variations in the genomes of NF1 and NF2 patients
neurogenetics (2017)