Abstract
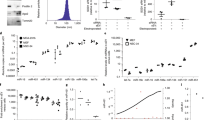
RNA interference (RNAi) is a natural cellular regulatory process that inhibits gene expression by transcriptional, post-transcriptional and translational mechanisms. Synthetic approaches that emulate this process (small interfering RNA (siRNA), short hairpin RNA (shRNA)) have been shown to be similarly effective in this regard. We developed a novel ‘bifunctional’ RNAi strategy, which further optimizes target gene knockdown outcome. A bifunctional construct (bi-sh-STMN1) was generated against Stathmin1, a critical tubulin modulator that is overexpressed in human cancers. The bifunctional construct is postulated to concurrently repress the translation of the target mRNA (cleavage-independent, mRNA sequestration and degradation) and degrade (through RNase H-like cleavage) post-transcriptional mRNA through cleavage-dependent activities. Bi-sh-STMN1 showed enhanced potency and durability in parallel comparisons with conventional shRNA and siRNAs targeting the same sequence. Enhanced STMN1 protein knockdown by bi-sh-STMN1 was accompanied by target site cleavage at the mRNA level showed by the rapid amplification of complementary DNA ends (RACE) assay. Bi-sh-STMN1 also showed knockdown kinetics at the mRNA level consistent with its multieffector silencing mechanisms. The bifunctional shRNA is a highly effective and advantageous approach mediating RNAi at concentrations significantly lower than conventional shRNA or siRNA. These results support further evaluations.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Irvine DV, Zaratiegui M, Tolia NH, Goto DB, Chitwood DH, Vaughn MW et al. Argonaute slicing is required for heterochromatic silencing and spreading. Science 2006; 313: 1134–1137.
Bailis JM, Forsburg SL . RNAi hushes heterochromatin. Genome Biol 2002; 3: Reviews1035.
Chu CY, Rana TM . Small RNAs: regulators and guardians of the genome. J Cell Physiol 2007; 213: 412–419.
Ghildiyal M, Zamore PD . Small silencing RNAs: an expanding universe. Nat Rev 2009; 10: 94–108.
Grimm D, Kay MA . Therapeutic application of RNAi: is mRNA targeting finally ready for prime time? J Clin Invest 2007; 117: 3633–3641.
Gewirtz AM . On future's doorstep: RNA interference and the pharmacopeia of tomorrow. J Clin Invest 2007; 117: 3612–3614.
Silva J, Chang K, Hannon GJ, Rivas FV . RNA-interference-based functional genomics in mammalian cells: reverse genetics coming of age. Oncogene 2004; 23: 8401–8409.
Wang S, Shi Z, Liu W, Jules J, Feng X . Development and validation of vectors containing multiple siRNA expression cassettes for maximizing the efficiency of gene silencing. BMC Biotechnol 2006; 6: 50.
Liu YP, Haasnoot J, ter Brake O, Berkhout B, Konstantinova P . Inhibition of HIV-1 by multiple siRNAs expressed from a single microRNA polycistron. Nucleic Acids Res 2008; 36: 2811–2824.
Li L, Lin X, Khvorova A, Fesik SW, Shen Y . Defining the optimal parameters for hairpin-based knockdown constructs. RNA (New York, NY) 2007; 13: 1765–1774.
Pillai RS . MicroRNA function: multiple mechanisms for a tiny RNA? RNA (New York, NY) 2005; 11: 1753–1761.
Zeng Y, Cai X, Cullen BR . Use of RNA polymerase II to transcribe artificial microRNAs. Methods Enzymol 2005; 392: 371–380.
Nemunaitis J, Senzer N, Khalil I, Shen Y, Kumar P, Tong A et al. Proof concept for clinical justification of network mapping for personalized cancer therapeutics. Cancer Gene Ther 2007; 14: 686–695.
Rana S, Maples PB, Senzer N, Nemunaitis J . Stathmin 1: a novel therapeutic target for anticancer activity. Expert Rev Anticancer Ther 2008; 8: 1461–1470.
Iancu C, Mistry SJ, Arkin S, Wallenstein S, Atweh GF . Effects of stathmin inhibition on the mitotic spindle. J Cell Sci 2001; 114 (Part 5): 909–916.
Alli E, Yang JM, Hait WN . Silencing of stathmin induces tumor-suppressor function in breast cancer cell lines harboring mutant p53. Oncogene 2006; 26: 1003–1012.
Zhang HZ, Wang Y, Gao P, Lin F, Liu L, Yu B et al. Silencing stathmin gene expression by survivin promoter-driven siRNA vector to reverse malignant phenotype of tumor cells. Cancer Biol Ther 2006; 5: 1457–1461.
Mistry SJ, Atweh GF . Therapeutic interactions between stathmin inhibition and chemotherapeutic agents in prostate cancer. Mol Cancer Ther 2006; 5: 3248–3257.
Ngo TT, Peng T, Liang XJ, Akeju O, Pastorino S, Zhang W et al. The 1p-encoded protein stathmin and resistance of malignant gliomas to nitrosoureas. J Natl Cancer Inst 2007; 99: 639–652.
Wang R, Dong K, Lin F, Wang X, Gao P, Wei SH et al. Inhibiting proliferation and enhancing chemosensitivity to taxanes in osteosarcoma cells by RNA interference-mediated downregulation of stathmin expression. Mol Med (Cambridge, MA) 2007; 13: 567–575.
Azuma-Mukai A, Oguri H, Mituyama T, Qian ZR, Asai K, Siomi H et al. Characterization of endogenous human argonautes and their miRNA partners in RNA silencing. Proc Natl Acad Sci USA 2008; 105: 7964–7969.
Lee Y, Kim M, Han J, Yeom KH, Lee S, Baek SH et al. MicroRNA genes are transcribed by RNA polymerase II. EMBO J 2004; 23: 4051–4060.
Lagos-Quintana M, Rauhut R, Meyer J, Borkhardt A, Tuschl T . New microRNAs from mouse and human. RNA (New York, NY) 2003; 9: 175–179.
Mendell JT . miRiad roles for the miR-17-92 cluster in development and disease. Cell 2008; 133: 217–222.
Wu L, Fan J, Belasco JG . Importance of translation and nonnucleolytic ago proteins for on-target RNA interference. Curr Biol 2008; 18: 1327–1332.
Matranga C, Tomari Y, Shin C, Bartel DP, Zamore PD . Passenger-strand cleavage facilitates assembly of siRNA into Ago2-containing RNAi enzyme complexes. Cell 2005; 123: 607–620.
Leuschner PJ, Ameres SL, Kueng S, Martinez J . Cleavage of the siRNA passenger strand during RISC assembly in human cells. EMBO Rep 2006; 7: 314–320.
Pizzorno MC, O'Hare P, Sha L, LaFemina RL, Hayward GS . Trans-activation and autoregulation of gene expression by the immediate-early region 2 gene products of human cytomegalovirus. J Virol 1988; 62: 1167–1179.
Soutschek J, Akinc A, Bramlage B, Charisse K, Constien R, Donoghue M et al. Therapeutic silencing of an endogenous gene by systemic administration of modified siRNAs. Nature 2004; 432: 173–178.
Roush SF, Slack FJ . Micromanagement: a role for microRNAs in mRNA stability. ACS Chem Biol 2006; 1: 132–134.
Wu L, Fan J, Belasco JG . MicroRNAs direct rapid deadenylation of mRNA. Proc Natl Acad Sci USA 2006; 103: 4034–4039.
Zeng Y, Cullen BR . Sequence requirements for micro RNA processing and function in human cells. RNA (New York, NY) 2003; 9: 112–123.
Silva JM, Li MZ, Chang K, Ge W, Golding MC, Rickles RJ et al. Second-generation shRNA libraries covering the mouse and human genomes. Nat Genet 2005; 37: 1281–1288.
Chen C, Ridzon DA, Broomer AJ, Zhou Z, Lee DH, Nguyen JT et al. Real-time quantification of microRNAs by stem-loop RT-PCR. Nucleic Acids Res 2005; 33: e179.
McManus MT, Petersen CP, Haines BB, Chen J, Sharp PA . Gene silencing using micro-RNA designed hairpins. RNA (New York, NY) 2002; 8: 842–850.
Landthaler M, Gaidatzis D, Rothballer A, Chen PY, Soll SJ, Dinic L et al. Molecular characterization of human Argonaute-containing ribonucleoprotein complexes and their bound target mRNAs. RNA (New York, NY) 2008; 14: 2580–2596.
Robb GB, Brown KM, Khurana J, Rana TM . Specific and potent RNAi in the nucleus of human cells. Nat Struct Mol Biol 2005; 12: 133–137.
Langlois MA, Boniface C, Wang G, Alluin J, Salvaterra PM, Puymirat J et al. Cytoplasmic and nuclear retained DMPK mRNAs are targets for RNA interference in myotonic dystrophy cells. J Biol Chem 2005; 280: 16949–16954.
Jarve A, Muller J, Kim IH, Rohr K, MacLean C, Fricker G et al. Surveillance of siRNA integrity by FRET imaging. Nucleic Acids Res 2007; 35: e124.
Chiu YL, Rana TM . siRNA function in RNAi: a chemical modification analysis. RNA (New York, NY) 2003; 9: 1034–1048.
Rossi JJ . Expression strategies for short hairpin RNA interference triggers. Human Gene Ther 2008; 19: 313–317.
Acknowledgements
We gratefully acknowledge the generous support of the Baylor Health Care System Foundation—Lane Newsom Drennan Fund, the Jasper L and Jack Denton Wilson Foundation, the WW Caruth, Jr Foundation Fund of Communities Foundation of Texas, Merkle Family Foundation and the Mary Crowley Cancer Research Centers. We would also like to acknowledge Brenda Marr and Susan Mill for their competent and knowledgeable assistance in the preparation of this paper.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Rao, D., Maples, P., Senzer, N. et al. Enhanced target gene knockdown by a bifunctional shRNA: a novel approach of RNA interference. Cancer Gene Ther 17, 780–791 (2010). https://doi.org/10.1038/cgt.2010.35
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cgt.2010.35
Keywords
This article is cited by
-
Secondary structure RNA elements control the cleavage activity of DICER
Nature Communications (2022)
-
Preclinical Justification of pbi-shRNA EWS/FLI1 Lipoplex (LPX) Treatment for Ewing's Sarcoma
Molecular Therapy (2016)
-
Design of Effective Primary MicroRNA Mimics With Different Basal Stem Conformations
Molecular Therapy - Nucleic Acids (2016)
-
Phase 1 Trial of Bi-shRNA STMN1 BIV in Refractory Cancer
Molecular Therapy (2015)
-
Vertically integrated translational studies of PDX1 as a therapeutic target for pancreatic cancer via a novel bifunctional RNAi platform
Cancer Gene Therapy (2014)