Abstract
Inosine triphosphate pyrophosphatase (ITPase), the enzyme that hydrolyzes ITP and other deaminated purine nucleoside triphosphates to the corresponding purine nucleoside monophosphate and pyrophosphate, is encoded by the Itpa gene. In this study, we established Itpa knockout (KO) mice and used them to show that ITPase is required for the normal organization of sarcomeres in the heart. Itpa−/− mice died about 2 weeks after birth with features of growth retardation and cardiac myofiber disarray, similar to the phenotype of the cardiac α-actin KO mouse. Inosine nucleotides were found to accumulate in both the nucleotide pool and RNA of Itpa−/− mice. These data suggest that the role of ITPase in mice is to exclude ITP from the ATP pool, and the main target substrate of this enzyme is rITP. Our data also suggest that cardiomyopathy, which is mainly caused by mutations in sarcomeric protein-encoding genes, is also caused by a defect in maintaining the quality of the ATP pool, which is an essential requirement for sarcomere function.
Similar content being viewed by others
Main
Nucleotide and polynucleotide molecules are spontaneously damaged under the physiological conditions.1, 2, 3 These modifications often cause a nucleotide to pair with an alternate partner nucleotide in DNA.4, 5 For example, 8-oxo-dGTP, an oxidized product of dGTP, can pair with adenine as well as cytosine during replication, resulting in a transversion mutation.4 8-Oxoguanine in DNA appears through two independent pathways, either by an incorporation of the oxidized precursor, 8-oxo-dGTP during DNA synthesis, or by the direct oxidation of a guanine base in DNA.6, 7 To prevent the former pathway from being used, mammals possess the oxidized purine nucleoside triphosphatase encoded by the Mth1 gene, that degrades 8-oxo-dGTP into 8-oxo-dGMP and pyrophosphate,8, 9 and to protect against the latter pathway, 8-oxoguanine-DNA glycosylase, encoded by the Ogg1 gene, removes the oxidized base from the DNA.10, 11, 12, 13, 14, 15 To evaluate the roles of these enzymes in mouse, we produced Ogg1 and Mth1 single-knockout mice, as well as Ogg1 plus Mth1 double-knockout mice, and showed their tumor susceptibility and the accumulation of 8-oxoguanine in their DNA.6
Similar to oxidation, deamination is also a common chemical modification that nucleotides may undergo under physiological conditions.1, 2 In the case of deaminated purine bases, such as hypoxanthine, it is considered that inosine triphosphate pyrophosphatase (ITPase), endonuclease V and hypoxanthine-DNA glycosylase have a role in eliminating the deaminated bases from DNA.16, 17, 18, 19 Saparbaev and Laval19 reported that alkylpurine-DNA glycosylases engage in hypoxanthine-DNA glycosylase activity both in Escherichia coli and mammalian cells. The alkylpurine-DNA glycosylase-deficient (Mpg knockout) mouse has features of impaired base excision repair of alkylation-induced DNA damage, and increased sensitivity to methyl methanesulfonate and streptozotocin-induced diabetes.20, 21 However, there is no report regarding ITPase and endonuclease V knockout mouse as yet.
The enzyme activity of ITPase (EC 3.6.1.19) was first described in 1969.22 ITPase, which is encoded by the Itpa gene in mammals, hydrolyzes deaminated purine nucleoside triphosphates, such as dITP and ITP, to the corresponding purine nucleoside monophosphate and pyrophosphate.16, 23 The enzyme was identified as a novel NTP pyrophosphatase by a protein structure-based approach in 1999.24 We have reported the genomic organization and expression of the mouse Itpa gene in 2005.23 In this study to elucidate the biological significance of ITPase in mice, we produced an Itpa knockout. These Itpa−/− mice died about 2 weeks after birth with features of growth retardation and cardiac myofiber disarray.
Results
Biological features of Itpa−/− mice
To disrupt the Itpa gene, exons 2–4 that encode V23-I88 of mouse ITPase were replaced with the polII-neo cassette (Figure 1a). Two independently targeted ES clones were obtained and used to establish the knockout (KO) mouse strains. The Itpa gene-disrupted ES cells were injected into C57BL/6J blastocysts to produce Itpa KO mice. Disruption of the Itpa gene was confirmed by Southern blotting (ES clone, Figure 1b) as well as by genomic PCR (KO mice, Figure 1c). When we analyzed the expression of the Itpa gene in Itpa KO mice, we detected a shorter length of cDNA in the RT-PCR product of Itpa KO mouse RNAs using primer sets positioned in exons 1 and 8 (Figure 2a). The shorter length of cDNA was detected in both KO mouse strains. The unexpected cDNA contained a sequence corresponding to exons 1, 5, 6, 7 and 8, with one G insertion between exons 1 and 5 (Figure 2b). This abnormally spliced RNA encodes a polypeptide composed of 29 amino acids, that is, the first 22 amino acids encoded by exon 1 of the Itpa gene and an additional 7 misframed amino acids derived from the exon 5 sequence. The aberrant translation product lacks the major portion of ITPase (176 a.a./198 a.a.), including the residues thought to interact with ITP (Figure 2c).16 The null expression of the ITPase protein in the Itpa−/− mouse was confirmed by western blotting using a rabbit anti-ITPase antiserum. We did not detect any signal corresponding to the expected size of ITPase (MW, 21.9 kDa) in Itpa−/− mouse samples (Figure 3a). A 3.1 kDa peptide, the expected translation product of abnormally spliced RNA, was not detected in our system, perhaps because of an instability of the peptide or the detection limit of the antiserum (Supplementary Figure). Finally, no ITPase activity was detected in the erythrocyte extract of the Itpa−/− mouse (Figure 3b), and we concluded that the gene disruption had succeeded.
Generation of the Itpa−/− mouse. (a) Schematic representation of a partial restriction map of the mouse Itpa locus, targeting vector and targeted allele. PCR primers to detect the wild type and knockout alleles are indicated by open and closed triangles, respectively. ‘p’ indicates the probe fragment used in Southern blot analysis. As there are several Itpa pseudogenes in the mouse genome,23 we cloned the exact Itpa gene located on chromosome 2 and used it to construct a targeting vector. (b) Southern blot analysis of Itpa+/− ES cells. The 5′-flanking probe (p) was used to distinguish the EcoRI-digested targeted allele. An approximate 6 kb targeted allele (KO) was found only in the lane of Itpa+/− ES cells. (c) Targeted allele confirmed by the genomic PCR of mouse tail DNA. Amplified DNAs derived from wild type (WT) and knockout (KO) alleles are indicated by open and closed triangles, respectively
RT-PCR analysis of Itpa−/− mouse RNA. (a) An aberrant RNA transcribed from the targeted allele. Positions of the primers used in the RT-PCR are shown in Figure 2b. The length of cDNA expected from the wild-type allele is shown with an open triangle, and the unexpected cDNA from the targeted allele, with a closed triangle. (1) and (2) indicate the different KO mouse strains. cDNAs prepared from the liver RNA were shown. The shorter molecule was also observed in the heart cDNA of the Itpa−/− mouse (data not shown). (b) An aberrant transcript encodes a short peptide composed of 29 amino acids. Amino acid sequences predicted from the nucleotide sequence of RT-PCR products and the predicted splicing occurring in wild and Itpa−/− mice are presented. Positions of the primers used in the RT-PCR are shown by open triangles. Peptide coding regions are shown in magenta. (c) Comparison of the predicted amino-acid sequence between wild-type ITPase and the short peptide encoded by the aberrant mRNA transcribed from the targeted allele. In the amino-acid sequence of the short peptide from Itpa−/− mouse cDNA, frame-shifted amino acids are indicated in the blue area
(a) Western blot analysis of Itpa−/− mouse tissues. ITPase is indicated by a closed triangle. (1) and (2) indicate the different KO mouse strains. Full imaging of the blot is shown in the Supplementary Figure with a long exposed image. (b) No ITPase activity was detected in the erythrocyte extract of the Itpa−/− mouse. The amount of IMP released from ITP by the erythrocyte extract was determined by HPLC as described in the Materials and Methods section. Itpa+/+, open circle; Itpa+/−, gray circle; Itpa−/−, closed circle
The Itpa+/− mouse was indistinguishable from its Itpa+/+ littermates with respect to size, viability, fertility and behavior. About 10% of offspring obtained by the mating of Itpa+/− mice showed the Itpa−/− genotype (Figure 4a). The birth ratio indicates that more than half of the Itpa−/− mice died before birth. Although there was a significant amount of milk in the stomach of Itpa−/− newborn mice, these mice showed growth retardation and died about 14 days after birth (Figure 4b-d). Prior to death, the Itpa−/− mouse showed ataxia, abnormal breathing (Supplemental Movie 1) and heart abnormalities. These phenotypes were common in the two knockout mouse lines, independently established from different Itpa+/− ES cells. On pathological analysis, the Itpa−/− mouse showed such features as immature hair follicles, hyperkeratosis of the forestomach, decreased extramedullary hematopoiesis in liver and spleen, and germ cell hypoplasia in testis (data not shown), in addition to hypoplasia of the heart (Figure 5a-c). The heart of the Itpa−/− mouse was smaller and immature, especially the ventricle regions. Analysis of a 10 days old Itpa−/− mouse heart showed that both ventricular chamber walls were thinner than those of the Itpa+/+ mouse heart (Figure 5d-f). The striated staining of desmin corresponding to the Z-disc was absent in the Itpa−/− heart and disorganization of myocardial fiber was observed (Figure 5g-i). By ultrastructural analysis of Itpa−/− mouse cardiomyocytes, it was evident that the number of sarcomeres was decreased and that the sarcomere structure was broken and disorganized. The shape of mitochondria was changed, although the inner and outer membrane structure seemed normal (Figure 5j-l). We detected neither apoptotic cardiomyocytes nor symptoms of myocardial infarction in Itpa−/− mouse heart (data not shown). These results indicate that an ITPase deficiency causes disruption of the sarcomere structure in cardiomyocytes. Using echocardiography, we compared the cardiac performance of the Itpa−/− mouse to that of the wild type. As shown in Supplementary Movie 2, movement of the Itpa−/− mouse heart was unsynchronized and jerky.
Itpa−/− mice died before weaning. (a) The birth ratio of the Itpa−/− mouse was not in accordance with the Mendel's laws. To determine the birth ratio of the Itpa−/− mouse, Itpa+/− mice (N1), which were the offspring of chimera male mice and C57BL/6J females, were used for mating. (b) Itpa−/− mice were smaller than Itpa+/+ and Itpa+/− mice. Ten days old male mice from the same litter were compared. (c) Itpa−/− mice died about 2 weeks after birth. The survival curve is shown as a Kaplan–Meier's plot. Itpa+/+, open circle; Itpa+/−, gray circle; Itpa−/−, closed circle. (d) Itpa−/− mice showed growth retardation. Itpa+/+, open circle; Itpa+/−, gray circle; Itpa−/−, closed circle. Error bar, S.E.M
The Itpa−/− mouse showed heart abnormalities. (a–c) Hypoplasia of Itpa−/− mouse heart (10 days old). (d–f) A horizontal section of heart stained by hematoxylin and eosin (11 days old). (g–i) Desmin staining. The Z-disc was not observed in the Itpa−/− mouse (11 days old). (j–l) Electron micrographic analysis. Microfibrillar disarray was evident in Itpa−/− mouse heart (10 days old)
Accumulation of inosine nucleotides in the Itpa−/− mouse
Under ITPase-deficient conditions, ITP was expected to appear in the nucleotide pool and the accumulated ITP to be incorporated into RNA. To analyze the contents of the nucleotide pool, we set up nucleotide separation conditions under which 15 kinds of nucleotides (ATP, CTP, GTP, UTP, ITP and their mono- and diphosphate forms) could be separated using an HPLC system equipped with a C-18 column (Figure 6a). As shown in Figure 6b, we confirmed that the ITP accumulation corresponded to about 10% of the ATP molecules in the nucleotide pool of the Itpa−/− mouse (10 days old) erythrocytes. No ITP was detected in Itpa+/+ and Itpa+/− mouse erythrocytes. In the case of organs, including the heart, however, we could not detect any ITP peak in the nucleotide pool of the Itpa−/− mouse (data not shown). One possibility is that the accumulated ITP is spent in its incorporation into RNA by transcription, except in erythrocytes where transcription does not occur. Alternatively, a rapid degradation of nucleotides may prevent detection of minor nucleotide species in tissues. To determine the inosine nucleotide that was incorporated into RNA, we digested total heart RNA with the nuclease P1, and separated the digest on a C-18 column. As shown in Figure 6c, we detected IMP in Itpa−/− mouse RNA, and the amount was equivalent to about 1% of the AMP. These results clearly showed that ITP was produced in mouse cells, and that ITPase degraded the deaminated nucleotides to maintain the quality of the ATP pool.
Accumulation of inosine nucleotide in the Itpa−/− mouse. The absorbance of each sample at 248.5 nm (the wavelength corresponding to the maximum absorbance of the inosine nucleotide at pH 6) was plotted. (a) HPLC conditions for the separation of nucleotides. A volume of 10 μl of a standard nucleotide mixture containing ATP, ADP, AMP, CTP, CDP, CMP, GTP, GDP, GMP, ITP, IDP, IMP, UTP, UDP and UMP (25 μ M each) were chromatographed on a SunFire 5.0 μmC18 column (4.6 × 250 mm) as described in the Materials and Methods section. After each run, the column was washed with 50% methanol. Under these conditions, we obtained a good separation pattern except for UMP and GDP. The AMP peak was not included on the chart due to its long retention time (13.663 min). (b) Accumulation of ITP in the nucleotide pool of Itpa−/− mouse erythrocytes. The peak area corresponding to the accumulated ITP is shown in green. No ITP was evident in Itpa+/+ and Itpa+/− mouse erythrocytes. (c) IMP detected in the total RNA of Itpa−/− mouse heart. To determine the amount of inosine nucleotide in RNA, we digested total RNA with the nuclease P1, and analyzed the digest on a C-18 column. The peak area of IMP observed in the Itpa−/− mouse is shown in green
Expression of the Actc1 gene in Itpa−/− mouse
As shown in Figure 5j, it was evident that the number of sarcomeres was decreased and that the sarcomere structure was broken and disorganized. This kind of abnormality is also a feature of human cardiomyopathy and its mouse models.29 Although a number of genes have been identified as responsible for the cardiomyopathy,30 until now, the Itpa gene has not been reported as a candidate.
A deficiency in the Actc1 gene, one of the genes encoding sarcomeric protein, is a causative factor of human cardiomyopathy. This gene encodes cardiac α-actin, and the Actc1 knockout mouse showed a similar phenotype to that of the Itpa−/− mouse.31 As the Actc1 gene is located near the Itpa gene on mouse chromosome 2, we determined whether Itpa gene disruption affected Actc1 gene expression. Semiquantitative RT-PCR analysis showed that the amount of Actc1 mRNA was not affected by Itpa gene disruption in the heart (Figure 7). The nucleotide sequence of Actc1 cDNA cloned from Itpa−/− mouse heart showed no mutations (data not shown). These results support the concept that the presence of ITP in the ATP pool has a deleterious effect on sarcomeric proteins, such as actomyosin, and that ITPase activity may alter vulnerability to certain cardiac muscle diseases.
The expression of the Actc gene was not affected by Itpa gene disruption. The total RNA of each mouse heart was subjected to semiquantitative RT-PCR analysis. Amplification was performed for 25, 30 and 35 cycles. Results of the 25-cycle amplification are presented in the figure. The primer sequences used for RT-PCR were as follows: Actc, 5′-AATCCAGCCGCCCCTAGCACG-3′ and 5′-TGCAAGTCCTGGTCTGGTTTA-3′; Gapdh, 5′-CTGCCATTTGCAGTGGCAAAG-3′ and 5′-TGGTATTCAAGAGAGTAGGGA-3′; Itpa, 5′-AAGCTTGCCATGGCTGCGTCTTTGGTCG-3′ and 5′-AAGTCTTCTAGAATTTACATTTGC-3′
Discussion
To control the quality of the nucleotide pool, organisms possess a number of nucleoside triphosphatases, which degrade non-canonical nucleoside triphosphates.32 To clarify the biological significance of the damaged nucleotides and the enzymes that eliminate them, we produced and analyzed KO mice which lack sanitizing enzyme activity. Here, we have produced and analyzed ITPase-deficient (Itpa−/−) mice. In contrast to the Mth1−/− mouse, which lacks 8-oxo-dGTPase activity and survives normally, the Itpa−/− mouse dies before weaning with features of growth retardation. We found that the heart of the Itpa−/− mouse is structurally and functionally abnormal.
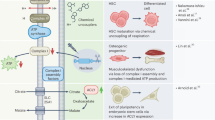
ATP, a multifunctional nucleotide and the most abundant in the nucleotide pool, plays a fundamental role in a wide variety of cellular processes, including RNA synthesis, energy translation, signal transduction, cytoskeleton remodeling and muscle contraction. In the case of cardiac function, a number of sarcomere proteins require ATP for their normal activities. Oxidative deamination of adenine at C-6 converts ATP to ITP. Because ITP retains a molecular structure similar to that of ATP, it can act as an aberrant substrate replacing ATP in some biological processes. For example, it has been shown that Mg•ITP-bound actomyosin has a greatly reduced shortening velocity and rate of force recovery as compared with the Mg•ATP-bound form, and shows disordered striations during activation in vitro.33 One likely possibility is that, during cardiac development of the Itpa−/− mouse, accumulated ITP competes with ATP (Figure 8a), which is required for actomyosin function in the sarcomere. Asynchronous sarcomere movements may cause a degradation of the striated pattern of the sarcomere, and destroy its well-organized structure (Figure 5j), which is essential for heart function. In addition, we observed that IMP is accumulated in the RNA of Itpa−/− mouse tissue (Figure 6c). It is also likely that the incorporated hypoxanthine nucleotides cause RNA information to be altered, resulting in the production of defective proteins and destruction of the structure of functional RNAs. In either case, it is necessary for cells to eliminate ITP from their ATP pool by ITPase to keep themselves healthy.
Quality control of the ATP pool in mouse by ITPase. (a) Sanitizing function of ITPase in the ATP pool. In the wild-type mouse, ITP, a deamination product of ATP, is hydrolyzed by ITPase to produce IMP, which is a normal intermediate metabolite of the purine biosynthesis pathway. ITP accumulates in cells when ITPase is absent. The accumulated ITP may act as a competitor of ATP in various kinds of bioreactions. (b) The function of ITPase in mouse cells. Quality control of the nucleotide pool is important not only because DNA (I) and RNA (II) are made up of the nucleotides which it supplies, but also because of the biological pathways that require nucleotides for activities (III), such as those involved in muscle contraction and heart development
It is well known that quality control of the nucleotide pool is important for DNA and RNA synthesis. By way of analogy to 8-oxo-dGTPase (MTH1) function, ITPase has been considered to prevent dITP-induced mutagenesis.7, 16 However, because the Km value of ITPase to its substrate is not particularly small (0.31 mM for dITP), 16 as compared with the size of the dNTP pool (0.013 mM for dATP),34 it is more likely that ITP, a deaminated product of ATP, which is the most abundant nucleotide in the nucleotide pool (2.8 mM),34 is a physiological substrate for ITPase in mammalian cells. Our findings with the Itpa−/− mouse are in good agreement with the hypothesis that ITPase functions to exclude ITP from the ATP pool and to protect cells from the harmful influence of deaminated purine nucleotides (Figure 8b).
In the case of human ITPase deficiencies, however, no major phenotype except for the abnormal accumulation of ITP in erythrocytes has been reported. Sumi et al.35, 36 found that the P32T mutation is responsible for the human ITPase deficiency, and proposed a relationship between this deficiency and an increased sensitivity to the toxicity of purine analog drugs. They mentioned that the ITP level in ITPase-deficient human erythrocytes may represent 10–25% of the ATP pool, which is similar to the amount of ITP observed in the Itpa−/− mouse. To explain the difference in the phenotypes of ITPase-deficient mice and humans, we considered two hypotheses. One is that there is another compensational mechanism in human cells used in excluding ITP from the ATP pool, other than that involving ITPase. The other is that an ITPase deficiency may be related to some form of heart disease in human populations, although this possibility has not yet been sufficiently explored. Clinical and biochemical analyses of ITPase-deficient patients may provide an answer.
As observed in the Itpa−/− mouse, a loss in ITP degradation leads to a cardiomyopathy-like phenotype, which is mainly caused by mutations of sarcomeric protein-encoding genes. It is clear that maintenance of the quality of the ATP pool is important for sarcomere organization in the heart. As ITP has structural similarity not only to ATP but also to GTP, ITPase activity would be important in keeping biochemical processes in cells functioning properly. Using ITPase-deficient cells, we are now able to evaluate the significance of deaminated purine nucleoside triphosphates originating in mammalian cells.
Materials and Methods
Materials
Oligonucleotides were obtained from Genenet Co. Ltd. (Fukuoka, Japan). Recombinant Taq DNA polymerase and 1 Kb Plus DNA Ladder were obtained from Takara Bio Inc. (Otsu, Japan) and Invitrogen Japan K.K. (Tokyo, Japan), respectively. Anti-β-actin antibody (A-5316), ITP and other nucleotides were purchased from Sigma-Aldrich Japan K.K. (Tokyo, Japan). Nuclease P1 was obtained from Seikagaku Corporation (Tokyo, Japan).
Generation and analyses of the Itpa-gene disrupted mouse
To disrupt the Itpa gene, the λ-phage clones, λmITPA22 and λmITPA4,23 which contain exons 1–6 and 2–7 of the mouse Itpa gene, respectively, were used to construct a targeting vector. The targeting vector was constructed by replacing the 1.0 kbp XbaI–BamHI region of a 9.5 kb genomic sequence, which contains exons 2–4, with the polII-neo cassette (Figure 1a). The targeting vector was then introduced into CCE ES cells by electroporation using standard procedures.25 Two independently targeted ES clones were obtained and used to establish the knockout mouse strains. Disruption of the Itpa gene was confirmed by Southern blotting, PCR and RT-PCR. Southern blotting was performed using a 32P-labeled EcoRI-NheI fragment as a 5′-flanking probe. The null expression of the ITPase protein in the Itpa−/− mouse was confirmed by western blotting using a rabbit anti-ITPase antiserum. ITPase activity was determined using erythrocyte extracts of the mice.
To determine the birth ratio of the Itpa−/− mouse, Itpa+/− mice (N1), which were the offspring of chimera male mice and C57BL/6J females, were used for mating. Itpa−/− mice were obtained by the mating of Itpa+/− mice (N1–N5). C57BL/6J mice were purchased from CLEA Japan Inc. (Tokyo, Japan). Echocardiograms were obtained using a Vevo660 imaging system (Vevo660, VisualSonics Inc., Toronto, Canada). All studies were approved by the Animal Care and Use Committee, Medical Institute of Bioregulation, Kyushu University.
PCR and RT-PCR
Tail DNA was used for the genomic PCR to determine the genotype of each mouse. The position and direction of each primer are shown in Figure 1a. The primer sequences used to determine the genotype were as follows: WT, 5′-CAGACAACCCTGAGTCACTAGGCTG-3′ and 5′-GTCTGTCTTCCTGTGTCCTGACTC-3′; KO, 5′-GTACATGTACATATCTATGTGGCCG-3′ and 5′-AGAGCGAGGGAAGCGTCTACCTA-3′. For RT-PCR analyses, total RNAs of liver and heart were prepared using ISOGEN (Nippon Gene Co. Ltd, Tokyo, Japan), according to the manufacturer's instructions. First-strand cDNA was synthesized with a First-Strand cDNA Synthesis kit (GE Healthcare Bio-Sciences K.K., Tokyo, Japan) using NotI-d(T)18 primer. The primers 5′-AAGCTTGCCATGGCTGCGTCTTTGGTCG-3′ and 5′-AAGTCTTCTAGAATTTACATTTGC-3′ were used to amplify the Itpa cDNA.
Western blotting and antibodies
Western blotting analysis26 and preparation of the anti-ITPase antiserum23 were described earlier. The rabbit anti-ITPase antiserum (1/500 dilution), which was able to detect both human and mouse ITPase proteins with almost the same efficiency,23 was used for the western blotting analysis. Proteins recognized by the anti-ITPase antiserum were detected by horseradish peroxidase-conjugated protein A with an ECL-Plus kit (GE Healthcare Bio-Sciences K.K.) using an LAS1000 Plus (Fuji Film Co., Tokyo, Japan) as a detector. The same membrane was treated with WB stripping solution (Nacalai Tesque Inc., Kyoto, Japan), and reused to detect β-actin by mouse monoclonal anti-β-actin antibody (A5316, Sigma-Aldrich Japan K.K.).
ITPase assay
ITPase activity was assayed by measuring the hydrolysis of ITP to IMP.23 The red blood cells (RBCs) were washed three times with three volumes of 0.9% NaCl, and quickly frozen in LN2. The frozen RBCs were lysed by adding a 10-fold volume of H2O to obtain the erythrocyte extract. The protein concentration was determined with a Protein Assay system (Bio-Rad Laboratories Inc., Tokyo, Japan) using bovine serum albumin (Thermo Fisher Scientific K.K., Yokohama, Japan) as a standard. The reaction mixture (50 μl) contained 50 mM Tris•HCl (pH 8.5), 50 mM MgCl2, 1 mM DTT, 0.2 mM ITP and 2–20 μg of the erythrocyte extract to be examined. The reaction was run at 30°C for 15 min, and stopped by adding an equal volume (50 μl) of 10% trichloroacetic acid. The reaction mixture was neutralized with a 1/9 volume of 3 M K2HPO4, and applied to HPLC analysis.
Detection of inosine nucleotides by HPLC
Separation and quantification of nucleotides were performed by HPLC using a Waters Alliance 2690 separation module equipped with a Model 996 photodiode array detector (Nihon Waters K.K., Tokyo, Japan). A buffer consisting of 0.1 M K2HPO4 (pH 6.0) was used as the mobile phase without any organic solvent. To detect ITP in the nucleotide pool, an erythrocyte extract corresponding to 90 μg of protein was treated with 5% trichloroacetic acid, and the soluble fraction was analyzed by HPLC (SunFire 5.0 μmC18 column, 4.6 × 250 mm, Nihon Waters K.K.). To quantify the inosine nucleotide in the heart RNA, the total RNA (2.5 μg) was digested with 0.5 U of nuclease P127 at 50°C for 1 h, and separated on a Wakopak Handy ODS column (4.6 × 250 mm, Wako Pure Chemical Industries Ltd, Osaka, Japan).
Pathological analysis
Hematoxylin and eosin, and anti-desmin immunohistochemical staining were performed as described.28 For the ultrastructural analysis, fixed heart sections were cut at a thickness of 70–80 nm, stained with 1% uranyl acetate followed by staining with a 1% lead citrate, 1% lead acetate and lead nitrate mixture, and were then examined with a JEM2000Ex (JEOL Ltd, Tokyo, Japan) electron microscope using an accelerating voltage of 80 KV.
Abbreviations
- ITPase:
-
inosine triphosphate pyrophosphatase
- RBCs:
-
red blood cells
- KO:
-
knockout
References
Lindahl T . Instability and decay of the primary structure of DNA. Nature 1993; 362: 709–715.
Mathews CK . DNA precursor metabolism and genomic stability. FASEB J 2006; 20: 1300–1314.
Taddei F, Hayakawa H, Bouton M, Cirinesi A, Matic I, Sekiguchi M et al Counteraction by MutT protein of transcriptional errors caused by oxidative damage. Science 1997; 278: 128–130.
Maki H, Sekiguchi M . Mut T protein specifically hydrolyses a potent mutagenic substrate for DNA synthesis. Nature 1992; 355: 273–275.
Chung JH, Back JH, Park YI, Han YS . Biochemical characterization of a novel hypoxanthine/xanthine dNTP pyrophosphatase from Methanococcus jannaschii. Nucleic Acids Res 2001; 29: 3099–3107.
Sakumi K, Tominaga Y, Furuichi M, Xu P, Tsuzuki T, Sekiguchi M et al Ogg1 knockout-associated lung tumorigenesis and its suppression by Mth1 gene disruption. Cancer Res 2003; 63: 902–905.
Nakabeppu Y, Sakumi K, Sakamoto K, Tsuchimoto D, Tsuzuki T, Nakatsu Y . Mutagenesis and carcinogenesis caused by the oxidation of nucleic acids. Biol Chem 2006; 387: 373–379.
Mo JY, Maki H, Sekiguchi M . Hydrolytic elimination of a mutagenic nucleotide oxodGTP by human 18-kilodalton protein: sanitization of nucleotide pool. Proc Natl Acad Sci USA 1992; 89: 11021–11025.
Sakumi K, Furuichi M, Tsuzuki T, Kakuma T, Kawabata S, Maki H et al. Cloning and expression of cDNA for a human enzyme that hydrolyzes 8-oxo-dGTP a mutagenic substrate for DNA synthesis. J Biol Chem 1993; 268: 23524–23530.
Rosenquist TA, Zharkov DO, Grollman AP . Cloning and characterization of a mammalian 8-oxoguanine DNA glycosylase. Proc Natl Acad Sci USA 1997; 94: 7429–7434.
Radicella JP, Dherin C, Desmaze C, Fox MS, Boiteux S . Cloning and characterization of hOGG1 a human homolog of the OGG1 gene of Saccharomyces cerevisiae. Proc Natl Acad Sci USA 1997; 94: 8010–8015.
Roldan-Arjona T, Wei YF, Carter KC, Klungland A, Anselmino C, Wang RP et al. Molecular cloning and functional expression of a human cDNA encoding the antimutator enzyme 8-hydroxyguanine-DNA glycosylase. Proc Natl Acad Sci USA 1997; 94: 8016–8020.
Bjoras M, Luna L, Johnsen B, Hoff E, Haug T, Rognes T et al Opposite base-dependent reactions of a human base excision repair enzyme on DNA containing 7,8-dihydro-8-oxoguanine and abasic sites. EMBO J 1997; 16: 6314–6322.
Lu R, Nash HM, Verdine GL . A mammalian DNA repair enzyme that excises oxidatively damaged guanines maps to a locus frequently lost in lung cancer. Curr Biol 1997; 7: 397–407.
Aburatani H, Hippo Y, Ishida T, Takashima R, Matsuba C, Kodama T et al. Cloning and characterization of mammalian 8-hydroxyguanine-specific DNA glycosylase/apurinic, apyrimidinic lyase, a functional mutM homologue. Cancer Res 1997; 57: 2151–2156.
Lin S, McLennan AG, Ying K, Wang Z, Gu S, Jin H et al Cloning, expression, and characterization of a human inosine triphosphate pyrophosphatase encoded by the ITPA gene. J Biol Chem 2001; 276: 18695–18701.
Feng H, Dong L, Klutz AM, Aghaebrahim N, Cao W . Defining amino acid residues involved in DNA-protein interactions and revelation of 3′-exonuclease activity in endonuclease V. Biochemistry 2005; 44: 11486–11495.
Dianov G, Lindahl T . Preferential recognition of I T base-pairs in the initiation of excision-repair by hypoxanthine-DNA glycosylase. Nucleic Acids Res 1991; 19: 3829–3833.
Saparbaev M, Laval J . Excision of hypoxanthine from DNA containing dIMP residues by the Escherichia coli, yeast, rat, and human alkylpurine DNA glycosylases. Proc Natl Acad Sci USA 1994; 91: 5873–5877.
Elder RH, Jansen JG, Weeks RJ, Willington MA, Deans B, Watson AJ et al Alkylpurine-DNA-N-glycosylase knockout mice show increased susceptibility to induction of mutations by methyl methanesulfonate. Mol Cell Biol 1998; 18: 5828–5837.
Cardinal JW, Margison GP, Mynett KJ, Yates AP, Cameron DP, Elder RH . Increased susceptibility to streptozotocin-induced beta-cell apoptosis and delayed autoimmune diabetes in alkylpurine-DNA-N-glycosylase-deficient mice. Mol Cell Biol 2001; 21: 5605–5613.
Chern CJ, MacDonald AB, Morris AJ . Purification and properties of a nucleoside triphosphate pyrophosphohydrolase from red cells of the rabbit. J Biol Chem 1969; 244: 5489–5495.
Behmanesh M, Sakumi K, Tsuchimoto D, Torisu K, Ohnishi-Honda Y, Rancourt DE et al Characterization of the structure and expression of mouse Itpa gene and its related sequences in the mouse genome. DNA Res 2005; 12: 39–51.
Hwang KY, Chung JH, Kim SH, Han YS, Cho Y . Structure-based identification of a novel NTPase from Methanococcus jannaschii. Nat Struct Biol 1999; 6: 691–696.
Tsuzuki T, Sakumi K, Shiraishi A, Kawate H, Igarashi H, Iwakuma T et al Targeted disruption of the DNA repair methyltransferase gene renders mice hypersensitive to alkylating agent. Carcinogenesis 1996; 17: 1215–1220.
Tsuchimoto D, Sakai Y, Sakumi K, Nishioka K, Sasaki M, Fujiwara T et al Human APE2 protein is mostly localized in the nuclei and to some extent in the mitochondria, while nuclear APE2 is partly associated with proliferating cell nuclear antigen. Nucleic Acids Res 2001; 29: 2349–2360.
Fujimoto M, Kuninaka A, Yoshino H . Purification of a nuclease from Penicillium citrinum. Agric Biol Chem 1974; 38: 777–783.
Dutta KK, Nishinaka Y, Masutani H, Akatsuka S, Aung TT, Shirase T et al Two distinct mechanisms for loss of thioredoxin-binding protein-2 in oxidative stress-induced renal carcinogenesis. Lab Invest 2005; 85: 798–807.
Seidman JG, Seidman C . The genetic basis for cardiomyopathy: from mutation identification to mechanistic paradigms. Cell 2001; 104: 557–567.
Towbin JA, Bowles NE . The failing heart. Nature 2002; 415: 227–233.
Kumar A, Crawford K, Close L, Madison M, Lorenz J, Doetschman T et al Rescue of cardiac alpha-actin-deficient mice by enteric smooth muscle gamma-actin. Proc Natl Acad Sci USA 1997; 94: 4406–4411.
Galperin MY, Moroz OV, Wilson KS, Murzin AG . House cleaning, a part of good housekeeping. Mol Microbiol 2006; 59: 5–19.
Burton K, White H, Sleep J . Kinetics of muscle contraction and actomyosin NTP hydrolysis from rabbit using a series of metal-nucleotide substrates. J Physiol 2005; 563: 689–711.
Kornberg A, Baker TA . The building blocks of DNA synthesis In: Kornberg A and Baker TA (eds) DNA Replication (2nd ed.) ( WH Freeman and Company, NY, USA. ) 2002 pp 53–54.
Sumi S, Marinaki AM, Arenas M, Fairbanks L, Shobowale-Bakre M, Rees DC et al Genetic basis of inosine triphosphate pyrophosphohydrolase deficiency. Hum Genet 2002; 111: 360–367.
Maeda T, Sumi S, Ueta A, Ohkubo Y, Ito T, Marinaki AM et al. Genetic basis of inosine triphosphate pyrophosphohydrolase deficiency in the Japanese population. Mol Genet Metab 2005; 85: 271–279.
Acknowledgements
We thank Dr. M Katsuki for CCE ES cells, Dr. Y Nakatsu for discussion, Y Yamada, M Sasaki, M Otsu, S Kitamura and A Matsuyama for technical assistance and Dr. W Campbell for comments on the manuscript. This study was supported by grants from Core Research for Evolutional Science and Technology (to YN), the Japan Science and Technology Agency, the Ministry of Education, Culture, Sports, Science, and Technology of Japan (18012035, 20012038 to KS), and the Japan Society for the Promotion of Science (19651097 to KS and 19390114 to DT).
Author information
Authors and Affiliations
Corresponding author
Additional information
Edited by M Piacentini
Supplementary Information accompanies the paper on Cell Death and Differentiation website (http://www.nature.com/cdd)
Rights and permissions
About this article
Cite this article
Behmanesh, M., Sakumi, K., Abolhassani, N. et al. ITPase-deficient mice show growth retardation and die before weaning. Cell Death Differ 16, 1315–1322 (2009). https://doi.org/10.1038/cdd.2009.53
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cdd.2009.53
Keywords
This article is cited by
-
An ITPA Enzyme with Improved Substrate Selectivity
The Protein Journal (2024)
-
Novel ITPA variants identified by whole genome sequencing and RNA sequencing
Journal of Human Genetics (2023)
-
Inosine triphosphate pyrophosphatase from Trypanosoma brucei cleanses cytosolic pools from deaminated nucleotides
Scientific Reports (2022)
-
A novel ITPA variant causes epileptic encephalopathy with multiple-organ dysfunction
Journal of Human Genetics (2020)
-
A disease spectrum for ITPA variation: advances in biochemical and clinical research
Journal of Biomedical Science (2016)