Abstract
The epidermis, the outer layer of the skin composed of keratinocytes, is a stratified epithelium that functions as a barrier to protect the organism from dehydration and external insults. The epidermis develops depending on the transcription factor p63, a member of the p53 family of transcription factors. p63 is strongly expressed in the innermost basal layer where epithelial cells with high clonogenic and proliferative capacity reside. Deletion of p63 in mice results in a dramatic loss of all keratinocytes and loss of stratified epithelia, probably due to a premature proliferative rundown of the stem and transient amplifying cells. Here we report that microRNA (miR)-203 is induced in vitro in primary keratinocytes in parallel with differentiation. We found that miR-203 specifically targets human and mouse p63 3′-UTRs and not SOCS-3, despite bioinformatics alignment between miR-203 and SOCS-3 3′-UTR. We also show that miR-203 overexpression in proliferating keratinocytes is not sufficient to induce full epidermal differentiation in vitro. In addition, we demonstrate that miR-203 is downregulated during the epithelial commitment of embryonic stem cells, and that overexpression of miR-203 in rapidly proliferating human primary keratinocytes significantly reduces their clonogenic capacity. The results suggest that miR-203, by regulating the ΔNp63 expression level, is a key molecule controlling the p63-dependent proliferative potential of epithelial precursor cells both during keratinocyte differentiation and in epithelial development. In addition, we have shown that miR-203 can regulate ΔNp63 levels upon genotoxic damage in head and neck squamous cell carcinoma cells, thus controlling cell survival.
Similar content being viewed by others
Main
The epidermis is continuously regenerated by mitotically active keratinocytes in the inner basal layer which, following detachment from the basement membrane, migrates to the outer cornified layer (terminally differentiated compartment): a process called cornification.1 The epidermis is continuously renewed by a regulated balance between proliferation and differentiation. The renewal capacity is due to the presence of epidermal stem cells and transient amplifying (TA) cells in the basal layer of the interfollicular epidermis and in the bulge (at least in mouse) of the hair follicle.2 The importance of the transcription factor p63 for the formation of the epidermis and other stratified epithelia arises from two lines of evidence. First, p63 is strongly expressed in the innermost basal layer where epithelial cells with high clonogenic and proliferative capacity reside.3 Second, mice lacking all p63 isoforms have no epidermis4, 5, 6 or squamous epithelia (prostate, urothelium), probably due to a premature proliferative rundown of the stem and TA cells. Mice knockout for p63 (p63−/−) also show lack of epithelial appendages, such as mammary, salivary and lachrymal glands, hair follicles, teeth, and have truncated limbs and abnormal craniofacial development. This is most likely due to failure to maintain or differentiate the apical ectodermal ridge, a structure that is important for coordination of epithelial–mesenchymal interactions,4, 5 and is required for limb outgrowth and palatal and facial structure formation.
MicroRNAs (miRs) are 19–22-nt non-coding RNAs that can suppress the expression of protein-coding genes by targeting the 3′-UTR region of mRNAs for translational repression, degradation or both.7, 8 miRs are generated by sequential processing of long RNA polymerase II transcripts by two key RNase III proteins, Drosha and Dicer.9 By associating with the 3′-UTR region of target mRNAs, miRs have important regulatory roles in diverse cellular pathways including: development, differentiation, organogenesis, stem cells and germ line proliferation, growth control and apoptosis.10, 11, 12, 13, 14 Indeed, mutations in Dicer, which globally disrupts miR processing, cause a range of developmental defects.15 Loss of function of mouse Dicer results in early embryonic lethality, and inducible deletion of Dicer in mouse embryonic stem (ES) cells causes defects in their proliferation, suggesting that, among other functions, Dicer may be essential for expansion of stem cells in gastrulating embryos.16, 17 When miRs are ablated in skin epithelium by conditionally targeting Dicer1, barrier function of the epithelium is compromised and hairs fail to invaginate,18, 19 indicating the functional importance of these small RNAs in skin development. miRs expressed in the developing mouse skin and hair follicles have been identified using microarray approaches.18, 19 Many skin miRs can be classified in families on the basis of their 5′ seed sequences, but others cannot, and represent orphan sequences. In particular, the miR-200 family (containing miR-200a, miR-200b, miR-200c, miR-141 and miR-149) and the miR-19/20 family (containing miR-19b, miR-20, miR-17-5p, miR-93) are preferentially expressed in the epidermis, whereas the miR-199 family (containing miR-199a and miR-199b) is exclusively expressed in hair follicles.18 Among the orphan miRs, miR-203 was identified to be highly and specifically expressed in the epidermis. miR-203 is also upregulated in psoriatic plaques,20 suggesting that miR dysregulation could be involved in epidermal pathogenesis.
In the present study, we explored the functional significance of miR-203 in primary keratinocyte differentiation and in the proliferative potential of human epidermal cells. In addition, we also investigated the role of miR-203 in regulating the ΔNp63α protein level following genotoxic stress in squamous cell carcinoma cells.
Results and Discussion
miR-203 and ΔNp63 are inversely expressed in keratinocytes
To investigate the role of miR-203 during keratinocyte differentiation, we evaluated miR-203 expression by northern blot in primary human and mouse keratinocytes. As shown in Figure 1a and b, miR-203 was rapidly upregulated (about 12-fold in mouse keratinocytes upon 24 h calcium treatment) when cells where induced by calcium to differentiate in vitro. Concurrently, with miR-203 upregulation, we observed ΔNp63 downregulation in differentiating cells (Figure 1a and b). Keratinocyte differentiation was confirmed by western blot staining of involucrin or K10. Although differentiation in mouse and human keratinocytes follows different kinetics, as shown in the figure, the inverse relationship between miR-203 and ΔNp63 expression remains valid for cells of both species. This observation suggested that miR-203 may directly repress suprabasal ΔNp63 and restrict cell proliferation in differentiating keratinocytes. Indeed, bioinformatics suggest that miRs and their targets are usually mutually exclusive in tissues.
Expression of miR-203 during keratinocyte differentiation. (a) Primary mouse keratinocytes were isolated as described in Materials and Methods and plated on dishes. Before they reached confluence, cells were induced to differentiate by adding 1.2 mM CaCl2 to the culture medium. Cells were collected at the indicated time points to perform northern blot and western blot analysis. Hybridization to detect U2snRNA was used as a loading control. miR-203 expression was normalized to U2snRNA by densitometry and results are reported as fold change over zero time. Results obtained from the densitometry are shown as mean±S.D. from three independent experiments. Differentiation was evaluated by western blot for involucrin. ΔNp63 protein downregulation during differentiation is also shown; actin is used as loading control. The histograms show the quantitative laser densitometry of miR-203 (fold over control; black bars) and p63 expression (% of reduction; white bars) as mean±S.D. from three independent experiments. (b) Human epidermal keratinocytes (HEKs) were cultured in KBM medium with KGM-2 growth supplements. Cells were induced to differentiate by adding 1.2 mM CaCl2 to the culture medium. Cells were collected at the indicated time points to perform northern blot and western blot analysis. Hybridization to detect U2snRNA was used as a loading control. MiR-203 expression was normalized to U2snRNA by densitometry, and results are reported as fold change over zero time. Differentiation was evaluated by western blot for K10. ΔNp63 protein downregulation during differentiation is also shown; actin is used as loading control. The histograms show the quantitative laser densitometry of miR-203 (fold over control; black bars) and p63 expression (% of reduction; white bars) as mean±S.D. from three independent experiments. (c) Primary mouse keratinocytes, before reaching confluence, were infected with adenovirus expressing murine pre-miR-203 or a scrambled sequence as negative control and maintained in culture for 2 days (lanes 1–6). As a positive control, cells infected with scrambled sequence were induced to differentiate by adding 1.2 mM CaCl2 to the culture medium for 1 and 2 days (lanes 7–9). Cells were collected at the indicated time points to perform western blot analysis for SOCS-3, a putative miR-203 target (see Supplementary Figure S1). A representative result of three independent experiments is shown. The histogram shows the quantitative laser densitometry of SOCS-3 protein (fold over control) normalized over actin expression, as mean±S.D. from three independent experiments
First, we investigated the regulation of SOCS-3, a putative molecular target of miR-20320 (Supplementary Figure S1), during keratinocyte differentiation. As shown in Figure 1c (lanes 7–9), SOCS-3 protein expression showed a significant, consistent increase during in vitro-induced differentiation in primary mouse keratinocytes in parallel with miR-203, suggesting that SOCS-3 3′-UTR is not a miR-203 target site, in contrast with the bioinformatics data (Supplementary Figure S1), showing in silico alignment between SOCS-3 3′-UTR and miR-203.
To further evaluate directly if SOCS-3 is indeed a genuine target of miR-203, we exogenously expressed miR-203 in mouse keratinocytes by infection with adenovirus containing miR-203. Figure 1c (lanes 4–6) shows a slight upregulation of SOCS-3 as compared with keratinocytes infected with adenovirus containing scrambled sequence (lanes 1–3). These data indicate that SOSC-3 is not repressed by miR-203; conversely its upregulation is presumably independent of miR-203.
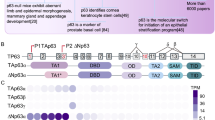
A predicted miR-203 target site in human p63 3′-UTR was identified by TargetScan 4.1 software (see Materials and Methods). More specifically, within the 3′-UTR of ΔNp63 mRNA, a heptamer matches perfectly with the miR-203 sequence (Figure 2a and b). The miR-203 recognition sequence is conserved and is present in the p63 3′-UTR of human, mouse, rat, dog and chicken (Figure 2a), suggesting that this mechanism for regulating p63 by miR-203 is common at least in these five species. An additional heptameric sequence that matches the miR-203 binding site was also found in the human p63 3′-UTR although this sequence is not conserved in other vertebrates, and we have not pursued it further here. When the entire 2772-bp p63 3′-UTR was introduced into the 3′-UTR of a luciferase reporter gene, miR-203 caused 45% reduction in activity, using both human and mouse miR-203 sequences. Deletion of the ΔNp63 3′-UTR heptamer abolished miR-203-mediated repression under the same conditions (Figure 2c). Mouse keratinocytes infected with adenovirus containing miR-203 also showed decreased ΔNp63 mRNA as compared with keratinocytes infected with adenovirus containing a scrambled sequence (Figure 2d). As control, Figure 2d also shows p63 mRNA levels in keratinocytes induced to differentiate by calcium treatment. p63 protein expression was also markedly diminished in mouse keratinocytes infected with adenovirus containing miR-203 (Figure 2e, lanes 3–4) compared with mouse keratinocytes infected with adenovirus containing a scrambled sequence (lanes 1–2). Together, these results indicate that effect of miR-203 on p63 is direct and is mediated by a specific 3′-UTR target site.
miR-203 targets p63 mRNA at 3′-UTR. (a) A predicted miR-203 target site on p63 3′-UTR was identified by TargetScan 4.1 software. The miR-203 recognition sequence exists in p63 3′-UTRs of human (Hs), mouse (Mm), rat (Rn), dog (Cf) and chicken (Gg). (b) Deletion of ΔNp63 3′-UTR heptamer. (c) Insertion of p63 3′-UTR in a luciferase reporter gene leads to diminished luciferase activity in the presence of miR-203, both using human and mouse p63 3′-UTR. Deletion of the ΔNp63 3′-UTR heptamer abolished miR-203-mediated repression. (d) Primary mouse keratinocytes were infected with adenovirus containing miR-203 versus that containing a scrambled sequence for 24 and 48 h. As an additional control, cells infected with the scrambled sequence were also treated with calcium to induce differentiation. Expression of p63 was evaluated by real-time PCR quantification of mRNAs. Results are presented as mean±S.D. from three independent experiments. (e) Western blot indicating that ΔNp63 protein is markedly diminished in mouse keratinocytes infected with adenovirus containing miR-203 versus mouse keratinocytes infected with adenovirus containing scrambled sequence. The histogram shows the quantitative laser densitometry of ΔNp63 protein over actin expression, as mean±S.D. from three independent experiments
miR-203 is not sufficient per se to trigger full expression of differentiating epithelial markers
To further test the hypothesis that miR-203 functions in the switch between proliferative and terminally differentiating keratinocytes, we overexpressed miR-203 in proliferating primary mouse keratinocytes. We found that miR-203, in contrast to calcium treatment, is not sufficient by itself to induce epidermal differentiation in vitro, as evaluated by western blot for involucrin expression (Figure 3a) and by real-time PCR of early (involucrin) and late (transglutaminase (TG)5, TG1) differentiation markers (Figure 3b). One and 2 days after infection of miR-203 into proliferating (low calcium) cultured mouse keratinocytes, epidermal differentiation markers were not induced as judged by expression of involucrin, TG1 and TG5 mRNAs (see scramble and miR-203-infected cells, Figure 3b). In contrast, 1 and 2 days after calcium-induced differentiation, all three differentiation markers were strongly induced.
miR-203 alone is not sufficient per se to induce the expression of epithelial differentiation markers. (a) Primary mouse keratinocytes were isolated as described in Materials and Methods and plated on dishes. Before they reached confluence, proliferating cells were infected with adenovirus containing murine pre-miR-203 or a scrambled sequence as negative control. As positive control, cells infected with the scrambled sequence were also treated with calcium to induce differentiation. Cells were collected at the indicated time points to perform western blot analysis for ΔNp63 and involucrin. ΔNp63 and involucrin expression were normalized to actin by densitometry, and results are reported as fold change over zero time. The histograms show the quantitative laser densitometry of involucrin (fold over control) and ΔNp63 expression (% of reduction). The densitometry is shown as mean±S.D. from three independent experiments. (b) In the same experimental conditions, involucrin and other epithelial differentiation marker mRNAs (TG1 and TG5) were quantified by real-time PCR. The data indicate that the induction of these differentiation markers is independent of miR-203, whereas with calcium, all three markers were abundantly induced. Results are shown as mean±S.D. from three independent experiments
Therefore, increased expression of miR-203 cannot be the only mechanism by which ΔNp63 protein levels are decreased during epidermal differentiation. Indeed, we have previously shown that the ubiquitin E3 ligase, Itch, contributes to the reduction in ΔNp63 protein levels.21 However, this post-translational regulation of ΔNp63 does not necessarily exclude a physiological role for miR-203, and potentially other miRs, in the regulation of ΔNp63. Nonetheless, our results confirm the role of ΔNp63 at the interface between the proliferative epithelial progenitor cells and the differentiation pathway3, 22, 23, 24, 25 and identify miR-203 as the molecular switch contributing to the downregulation of ΔNp63.
miR-203 inhibits the ΔNp63-dependent clonogenicity of epidermal cells
Epidermal stem cells display a higher proliferating potential than progenitor cells as detected in a clonogenicity assay.26 It has recently been demonstrated that inhibition of ΔNp63 drastically decreased the high proliferation rate of holoclones.3, 25 We thus compared the proliferating capacity of human primary keratinocytes transfected with pre-miR-203 with antago-miR-203 or with a scrambled sequence in a colony-formation assay. As expected, pre-miR-transfected cells showed reduced cell growth when compared to cells transfected with scrambled oligonucleotides. In contrast, overexpression of antago-miR-203 resulted in an increase in cell growth (Figure 4). The number of rapidly proliferating colonies significantly decreased from 1.7% in control to 0.7% with miR-203 and increased to 2.9% when an antago-miR was transfected into HEK. This suggests that overexpression of miR-203 triggers cells toward differentiation by interfering with the proliferative potential of proliferating keratinocytes through ΔNp63 downregulation. This result is consistent with a requirement for ΔNp63 in maintaining proliferative potential of epidermal stem cells.
miR-203 specifically inhibits the proliferative potential of human epidermal cells. Colony-forming assay obtained from normal human keratinocytes transfected with scrambled control, miR-203 and antago-miR. Micrographs shown are representative images obtained from three independent experiments. Numbers in parentheses represent the relative percentage of clones (*P<0.01).
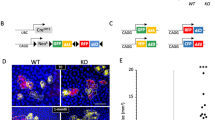
Regulation of miR-203 and ΔNp63 in ES cells
Embryonic stem cells can recapitulate in vitro the main steps of embryonic epidermal differentiation upon exogenous addition of bone morphogenic protein (BMP)-4. This causes sox-1+ neural precursors to undergo spontaneous cell death while inducing ectodermal commitment (K8/K18+ cells) and epidermal differentiation27, 28 (K5/K14+ cells; Figure 5a and b). Soon after BMP-4 treatment, the ΔNp63 gene is activated and remains strongly expressed until the cells undergo terminal epidermal differentiation.28 Using these experimental conditions, and concomitant with ΔNp63 mRNA upregulation, we observed that miR-203 expression is downregulated (Figure 5c). This further supports the direct link between p63 and miR-203 and suggests that miR-203 may be absent or only expressed at low levels in multipotent proliferating cells to allow p63-dependent epidermal commitment, whereas its expression increases as the multipotent epidermal cells exit the proliferating basal layer and terminally differentiate.
Expression profiling of ΔNp63 and miR-203 during epidermal differentiation of ES cells. (a) Schematic representation of the murine ES differentiation model. (i) Murine undifferentiated ES cells cultured on fixed NIH-3T3 cells under serum-free conditions undergo differentiation into neuronal cells. (ii) BMP-4 (0.5 nM) treatment at days 4–6 induces differentiation into K8/18-expressing ectodermal cells, and addition of serum at day 9 enhances efficient differentiation into K5/14-expressing keratinocytes. (b and c) RNA samples were collected at the indicated time points (at day 4, samples were collected 8 h after BMP-4 addition), and the relative expression levels of K-14 and CK-18 (b) or ΔNp63 (c) was determined by real-time PCR analysis as described in Materials and Methods. The relative levels of miR203 (c) were examined by TaqMan real-time PCR as detailed in Materials and Methods. Values were normalized to a control gene (36B4) or miR (Sno202), and results represent the calculated fold change in the amounts of mRNAs or miR-203 as compared with day 1 of differentiation (*P<0.05)
miR-203 induces cell cycle arrest and apoptosis in JHU-012 cells upon UVC irradiation
Besides the epidermis and other stratified squamous epithelia, ΔNp63 is commonly overexpressed in up to 80% of primary head and neck squamous cell carcinomas (HNSCCs).29 This is in part due to the fact that squamous cell carcinomas often exhibit genomic amplification of 3q, and p63 is located on chromosome 3q27–28.30, 31 HNSCCs are derived from cells within the basal epithelia and mucosa and are usually resistant to therapy.24 Recently, it has been shown that ΔNp63α is also overexpressed in HNSCC cell lines (including JHU-012) and in these cells, it acts as a crucial survival factor by suppressing p73-dependent apoptosis.29, 32, 33, 34 Knockdown of ΔNp63 induces the proapoptotic bcl-2-family members Puma and Noxa, and both their induction and cell death are p53-independent, but require transactivating isoforms of p73. The inhibition of p73-dependent transcription by ΔNp63α involves both direct promoter binding and physical sequestration of p73 by protein–protein interaction.29, 34, 35
JHU-012 cells were irradiated with UVC (0.005 J/cm2) for 30 s and collected after 12, 24 and 36 h of culture. As shown in Figure 6a, ΔNp63 expression was already decreased in JHU-012 cells 12 h after UVC treatment. Simultaneously, with ΔNp63 downregulation, miR-203 was significantly upregulated as evaluated by real-time PCR (Figure 6b). Significant effects on cell death, as compared to non-irradiated cells, were observed in JHU-012 cells at 12, 24 and 36 h after UVC irradiation, as evaluated by sub-G1 events (Figure 6d). Cell death was accompanied by increased cleavage of the poly(ADP-ribosylating) enzyme PARP-1, a specific hallmark of apoptotic cell death and, as a control of damage, c-jun accumulation (Figure 6e). In addition, UVC irradiation results in an increased number of cells in G1 with a reduction in the number of cells in G2 cell cycle phase (Figure 6c). To verify that the observed effect on apoptosis and cell cycle is, at least in part, mediated by miR-203, we infected JHU-012 cells with adenoviruses containing miR-203 or scrambled sequence. ΔNp63 protein expression was markedly diminished (about 50%) in JHU-012 cells infected with adenovirus containing miR-203 as compared with cells infected with adenovirus containing scrambled sequence (Figure 7a). In addition, we observed a significant increase of cell death (from 5 to 17%, Figure 7b), an increased number of cells in G1 and a reduction in the proportion of cells in G2 (Figure 7c).
Effects of UVC irradiation on miR-203 and ΔNp63α in JHU-012 cells. (a) Level of ΔNp63 after UVC exposure by western blot. ΔNp63 protein decreases by about 50% at 12 h after irradiation and by 70% at 36 h after irradiation. Actin staining was performed as loading control. The ΔNp63/actin ratio from the densitometry analysis of the western blot bands is shown in the lower panel. The densitometry is shown as mean±S.D. from three independent experiments. (b) Real-time PCR of miR-203 in UVC-irradiated cells (0.005 J/cm2 for 30 s). RNA was extracted 12, 24 and 36 h after UVC treatment. Error bars (S.D.) are calculated from three independent experiments. (c) Cell cycle profile of UVC-irradiated (0.005 J/cm2 for 30 s) cells, evaluated after 12, 24 and 36 h of culture. Cells were stained with PI and analyzed by flow cytometry. (d) Apoptosis was evaluated as sub-G1 events. Error bars (S.D.) are derived from three independent experiments. (e) Western blotting of PARP-1, a specific hallmark of apoptotic cell death, and as control of damage, c-jun accumulation is also shown. Actin staining is performed as loading control
miR-203 alone is sufficient to downregulate ΔNp63α in JHU-012 cells. (a) Western blot indicating that ΔNp63 protein is markedly diminished in cells infected with adenovirus containing miR-203 compared with cells infected with adenovirus containing a scrambled sequence. The ΔNp63/actin ratio from the densitometry analysis of the western blot bands is shown in the right panel. The densitometry is shown as mean±S.D. from three independent experiments. (b) Apoptosis is shown as sub-G1 events. Error bars (S.D.) are derived from three independent experiments. (c) Cell cycle profile of cells infected with adenovirus containing miR-203 and, as control, with adenovirus containing the scrambled sequence. Cells were stained with PI and analyzed by flow cytometry. All the experiments were performed in triplicate
These data indicate that the miR-203/ΔNp63 pathway is not solely related to normal keratinocytes, but it is retained in their tumor compartment, potentially opening novel ways for therapeutic intervention. Moreover, even though the reduction in proliferation is retained, it is of modest entity, possibly due to the simultaneous induction of cell death. Finally, the upstream stimulus for miR-203 induction is completely different, being related to UV-induced stress.
Concluding remarks
Together, the data presented here indicate that miR-203, by targeting ΔNp63 mRNA, acts as a switch between keratinocyte proliferation and differentiation in adult epidermis. Therefore, miR-203 is a key molecule controlling the passage of keratinocytes from the proliferative basal compartment to the differentiated suprabasal layers. In addition, the discovery that miR-203 levels are low during embryonic epidermal commitment suggests that miR-203 may have an important role during development by controlling the ‘stemness’ of cells. While this manuscript was being submitted, the group of Elaine Fuchs also reported the ability of miR-203 to promote epithelial differentiation by repressing ‘stemness’.36 The results presented in our paper are entirely in agreement with those reported by Fuchs.36 Both papers, while not excluding the contribution of additional miRs or other factors, indicate that miR-203 is a molecular switch at the boundary between proliferative basal progenitors and terminally differentiated suprabasal cells.
In addition, we found that miR-203 may also have a role in cancer. Indeed, miR-203 is upregulated upon UVC irradiation in the squamous cell carcinoma cells. Here, the data are compatible with the hypothesis that miR-203 could regulate p63mRNA and protein levels upon genotoxic damage in HNSCCs. This observation is important as recent evidence29, 35 demonstrates that, in addition to p53, both p63 (ΔNp63) and p73 are crucial mediators of cell death following chemotherapy in HNSCC. ΔNp63 is drastically downregulated following DNA damage, and ΔNp63 protein levels correlate with the patients response to cisplatin-based treatment.37 Therefore, our data indicate a possible new connection between miR-203 and activation of the apoptotic program in HNSCC.
Materials and Methods
Constructs
Human p63 3′-UTR was amplified by PCR from the first nucleotide after the stop codon to the last nucleotide before the polyadenylation signal from human genomic DNA using the following primers: p63UTR-SpeIF 5′-ggccactagtgcctcaccatgtgagctcttc-3′; p63UTR-SpeIR 5′-ggccactagtgcatgtcctggcaaacaaaaagag-3′. The 2770-bp fragment, after SpeI restriction, was ligated to a compatible XbaI-linearized pGL3Control vector (Promega, Madison, WI, USA). The miR-203-predicted target site (7 bp, AUUUCAA) was deleted by PCR using two overlapping primers: del203s 5′-ttaaagagaatgagtccttgagtttttgttgtacttaaatg-3′; del203e 5′-catttaagtacaacaaaactcaaggactcat-3′. To express human and mouse mir-203 in cell cultures by transfection, their genomic sequences were amplified from genomic DNA by PCR using the following primers: hsa203-BglIIF 5′-cgccagatctgtgcgcgaccagcggggatc-3′; hsa203-HindIIIR 5′-ggccaagcttccctggattggtcgccggcg-3′; mmu203-BamHIF 5′-ggccggatccccgccgcacagagtgcagcc-3′; mmu203-HindIIIR 5′-ggccaagcttctgccaggctgcgggactgc-3′. The resulting fragments (290 and 303 bp) were restricted and ligated to pSilencer 4.1-CMV puro (Ambion, Applied Biosystems, Foster City, CA, USA) linearized with BamHI/HindIII. To express human and mouse mir-203 in cell cultures by adenoviral infection, their genomic sequences were amplified from genomic DNA by PCR using the following primers: Ad-hsa203-XhoIF 5′-gcgcgcctcgaggtgcgcgaccagcggggatc-3′; Ad-hsa203-SpeIR 5′-gcgcgcactagtccctggattggtcgccggcg-3′; Ad-mmu203-XhoIF 5′-ggccctcgagccgccgcacagagtgcagcc-3′; Ad-mmu203-SpeIR 5′-ggccactagtctgccaggctgcgggactgc-3′. The resulting fragments (290 and 303 bp) were restricted and ligated to Ad Shuttle Vector 1.0-CMV (Ambion) XhoI/SpeI linearized. Viral production, growth, amplification and titration were performed in the human epidermal keratinocyte (HEK) 293E cell line according to the manufacturer's protocols.
Cell culture and transfection
HEK 293E cells were grown in DMEM High glucose, 10% FBS, 100 U penicillin, 100 μg streptomycin (GIBCO, Invitrogen, Carlsbad, CA, USA). HEK 293E cells were transfected by Lipofectamine 2000 according to the manufacturer's protocols (Invitrogen). JHU012 cells were grown in RPMI, 10% FBS, 100 U penicillin and 100 μg streptomycin (GIBCO, Invitrogen).
Primary mouse keratinocytes were isolated according to Yuspa et al.38 from CD1 strain newborn skin. The skin was floated overnight on trypsin/EDTA at 4°C. Trypsinization allows separation of dermis from epidermis. Primary keratinocytes were isolated from the latter and plated on collagen-coated dishes in medium supplemented with 0.05 mM CaCl2. Cells were induced to differentiate by adding 1.2 mM CaCl2 to the culture medium. Mouse primary keratinocytes were transfected with murine pre-mir 203, anti-mir 203 and Cy3-labeled negative control (Ambion) using the SiPORT neoFX transfection agent (Ambion) according to the manufacturer's protocols. Sixteen hours after transfection, the medium was removed, and calcium-induced differentiation was started by adding 1.2 mM CaCl2 to the culture medium. Normal Human Epidermal Keratinocytes (NHEK; Lonza, Basel, Switzerland) were cultured in KBM medium with KGM-2 growth supplements (Lonza).
Infection
Mouse primary keratinocytes were infected at MOI 50, for 1 h at 37°C in EMEM supplemented with 2% FBS. JHU012 cells were infected at MOI 50, for 1 h at 37°C in DMEM supplemented with 0.2% FBS.
Luc assay
A total of 2 × 10e5 HEK 293E cells were seeded in 12-well dishes 24 h before transfection. Hundred nanograms of pGL3 vectors, 2 μg of pSilencer 4.1-CMV puro vectors and 10 ng of Renilla luciferase pRL-CMV vector were cotransfected using Lipofectamine 2000. Luciferase activities of cellular extracts were measured 24 h after transfection, by using a Dual Luciferase Reporter Assay System (Promega); light emission was measured over 10 s using an OPTOCOMP I luminometer. Efficiency of transfection was normalized using Renilla luciferase activity.
Western blotting
Total cell extracts were resolved on an SDS-10% polyacrylamide gel and blotted onto a Hybond P PVDF membrane (G&E Healthcare, UK). Membranes were blocked with PBST 5% non-fat dry milk, incubated with primary antibodies for 2 h at room temperature, washed and hybridized for 1 h at room temperature using the appropriate horseradish peroxidase-conjugated secondary antibody (rabbit and mouse; BioRad, Hercules, CA, USA). Detection was performed with the ECL chemiluminescence kit (Perkin Elmer, Waltham, MA, USA). The antibodies used were anti-p63 (Ab4; Neomarkers, Fremont, CA, USA; 1/500 dilution), polyclonal anti-involucrin (Covance, Princeton, NJ, USA; 1/1000 dilution), anti-β-actin (Sigma, St. Louis, MI, USA; 1/5000 dilution), anti-PARP1 (Cell Signalling, Danvers, MA, USA; 1/500 dilution), anti-cJun (BD, Franklin Lakes, NJ, USA; 1/1000 dilution), anti-SOCS3 (Alexis, Lausen, Switzerland; 1/300 dilution).
RNA extraction and northern blot analysis
RNA was extracted with the mirVana miRNA isolation kit using the microRNA enrichment protocol (Ambion) and quantified by spectrophotometric analysis.
A total of 1.5 μg RNA was resolved on a Ready Gel TBE-Urea 15% acrylamide:bisacrylamide (19 : 1; Biorad) in 1 × TBE buffer. Gels were stained briefly with ethidium bromide, and RNA was transferred by electroblotting onto a GeneScreen Plus membrane (Perkin Elmer Life Sciences) for 2 h at 20 V in 0.5 × TBE. The membrane was UV crosslinked (Stratalinker; Stratagene). Prehybridization was performed for 1 h at 42°C in ULTRAhyb-Oligo hybridization buffer (Ambion), and the membrane was hybridized overnight at 42°C in the same buffer with the labeled probe. Antisense RNA probes were 3′-end-labeled with 32P-γATP by T4 polynucleotide kinase, and miR203 antisense probe sequence CTAGTGGTCCTAAACATTTCAC and U2snRNA antisense probe sequence GGGTGCACCGTTCCTGGAGGTAC were used. The membranes were washed thrice for 30 min in 0.5% SDS, 2 × SSC and exposed on an Imaging Screen-K (Biorad) at room temperature. Image scanning and analysis was performed on a Personal Molecular Imager using the QuantityOne software (Biorad).
Quantitative real-time one-step RT-PCR
Murine ES cell growth and differentiation were carried as described in detail previously.27 RNA extractions of the indicated differentiation days were obtained using Trizol (Invitrogen). The determination of the relative expression levels of mRNA transcripts was obtained by one-step real-time RT-PCR. Each reaction contained 12.5 μl SYBR-Green PCR Master Mix (Applied Biosystems), 0.125 μl MultiScribe Reverse Transcriptase (Applied Biosystems), 100 ng RNA and the appropriate specific primers (0.5 μM, sequences available upon request) and was adjusted to 25 μl reaction volume. Amplification and fluorescence detection according to the manufacturer's instructions was performed using the ABI PRISM 7700 Sequence Detection System (Applied Biosystems, France). The relative expression levels of micro-RNA 203 were measured by a two-step TaqMan assay according to the manufacturer's instructions. Briefly, RNA extractions were the templates for reverse transcriptase PCR of miR-203 or the internal control Sno202 (Applied Biosystems) using the TaqMan micro-RNA reverse transcription kit (Applied Biosystems). Next, TaqMan real-time PCR was prepared by using TaqMan universal master mix (Applied Biosystems) and specific primers for miR-203 or Sno202. The relative amounts of each RNA or miR product to its internal control (36B4 for mRNAs and Sno202 for miRs; ΔCT=CTx−CT36B4/Sno202) were used for calculating the fold change at each time point as compared with day 1 of differentiation (fold change=2−ΔΔCT; ΔΔCTday a=ΔCTday a −ΔCTday 1). RNA was extracted from mouse keratinocytes by using the RNAeasy kit (Qiagen, Milan, Italy). A total of 500 ng of RNA was used for reverse transcription using the InPromII kit (Promega) and one-tenth of the reaction was used for PCR. Real-time PCR was performed by using the Platinum SYBR Green qPCR SuperMix UDG with Rox (Invitrogen Life Technologies), with an amplification program as follows: one cycle of 95° for 3 min and 40 cycles of 94° for 20 sec and 59° for 40 sec. The reaction was followed by a melting curve protocol according to the specification of the ABI 7500 instrument (Applied Biosystems). Primers used were as follows: mTG1F 5′-accaccacagtgctccgatg-3′ and mTG1R 5′-gtttggaacttccacgtgtgg-3′ (for mouse Transglutaminase 1); mTG5F 5′-cagcccaggagccagaag-3′ and mTG5R 5′-ggcctcggcggacaac-3′ (for mouse Transglutaminase 5); mInvF 5′-tctccctcctgtgagtttgtttgg-3′ and mInvR 5′-cagtgaagacctggcattgtgtagg-3′ (for mouse Involucrine); and mΔNp63F 5′-cctggaagcagaaaagaggagagc-3′ and mΔNp63R 5′-tgtgcgtggtctgtgttgtagg-3′ (for mouse ΔNp63). Mouse β-actin was used as housekeeping gene for quantity normalization, and primers used were mActF 5′-tgtccctgtatgcctctggtcg-3′ and mActR 5′-gaaccgctcgttgccaatagtg-3′. Relative quantification of gene expression was calculated according to the method of 2-ΔΔCT described in ABI User Bulletin no. 2 (updated on October 2001) and the RQ software version 1.3 of Applied Biosystems.
Clonogenicity assay
HEKs were cultured on confluent feeder layers of lethally irradiated mouse J2-3T3 fibroblasts in keratinocyte medium (60% DMEM (Invitrogen), 30% Ham F12 (Invitrogen), 10% FetalcloneII (Hyclone), 5 μg/ml insulin (Sigma), 0.5 μg/ml hydrocortisone (Sigma), 0.2 nM Adenine (Sigma), 10 nM cholera toxin (Sigma) and 10 ng/ml EGF (Invitrogen). For clonogenicity assay, 1.5 × 105 HEK cells were cultured in six-well plates in keratinocyte-defined medium containing keratinocyte-SFM medium (Invitrogen), 30 μg/ml bovine pituitary extract (Invitrogen) and 10 ng/ml EGF (Invitrogen). On the next day, transfections of pre-miR203 or antago-miR-203 oligo-nucleotides (20–60 nM) were performed using Lipofectamine RNAiMAX (Invitrogen). After 48 h, cells were trypsinized and 100–500 cells were cultured on J2-3T3-coated 24-well plates in EGF-free medium A (to allow cell adherence) for 2 days. Complete keratinocyte medium that includes EGF was added at day 3 for an additional 12 days and colonies were visualized by Rhodamine staining followed by extensive washing.
Bioinformatics
mir203 target sites on p63 3′-UTR39 were predicted by TargetScan 4.1 software available at http://www.targetscan.org/.
Abbreviations
- ES cells:
-
embryonic stem cells
- HEK:
-
human epidermal keratinocytes
- HNSCC:
-
head and neck squamous cell carcinomas
- K:
-
keratin
- miR:
-
microRNA
- p63−/−:
-
mouse knockout for p63
- TA:
-
transactivation domain
- TA cells:
-
transient amplifying cells
- TG:
-
transglutaminase
- ΔN:
-
amino-terminal truncated protein
References
Candi E, Schmidt R, Melino G . The cornified envelope: a model of cell death in the skin. Nat Rev Mol Cell Biol 2005; 6: 328–340.
Blanpain C, Fuchs E . Epidermal stem cells of the skin. Ann Rev Cell Dev Biol 2006; 22: 339–373.
Senoo M, Pinto F, Crum CP, McKeon F . p63 Is essential for the proliferative potential of stem cells in stratified epithelia. Cell 2007; 129: 523–536.
Yang A, Kaghad M, Wang Y, Gillett E, Fleming MD, Dötsch V et al. p63, a p53 homolog at 3q27–29, encodes multiple products with transactivating, death-inducing, and dominant-negative activities. Mol Cell 1998; 2: 305–316.
Yang A, Schweitzer R, Sun D, Kaghad M, Walker N, Bronson RT et al. p63 is essential for regenerative proliferation in limb, craniofacial and epithelial development. Nature 1999; 398: 714–718.
Mills AA, Zheng B, Wang XJ, Vogel H, Roop DR, Bradley A . p63 is a p53 homologue required for limb and epidermal morphogenesis. Nature 1999; 398: 708–713.
Ambros V . The functions of animal microRNAs. Nature 2004; 431: 350–355.
Bartel DP . MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 2004; 16: 281–297.
Kim VN . MicroRNA biogenesis: coordinated cropping and dicing. Nat Rev Mol Cell Biol 2005; 6: 376–385.
Chen CZ, Li L, Lodish HF, Bartel DP . MicroRNAs modulate hematopoietic lineage differentiation. Science 2004; 303: 83–86.
Poy MN, Eliasson L, Krutzfeldt J, Kuwajima S, Ma X, Macdonald PE et al. A pancreatic islet-specific microRNA regulates insulin secretion. Nature 2004; 432: 226–230.
He L, Thomson JM, Hemann MT, Hernando-Monge E, Mu D, Goodson S et al. A microRNA polycistron as a potential human oncogene. Nature 2005; 435: 828–833.
O′Donnell KA, Wentzel EA, Zeller KI, Dang CV, Mendell JT . c-Myc-regulated microRNAs modulate E2F1 expression. Nature 2005; 435: 839–843.
Zhao Y, Samal E, Srivastava D . Serum response factor regulates a muscle-specific microRNA that targets Hand2 during cardiogenesis. Nature 2005; 436: 214–220.
Knight SW, Bass BL . A role for the RNase III enzyme DCR-1 in RNA interference and germ line development in Caenorhabditis elegans. Science 2001; 293: 2269–2271.
Bernstein E, Kim SY, Carmell MA, Murchison EP, Alcorn H, Li MZ et al. Dicer is essential for mouse development. Nat Genet 2003; 35: 215–217.
Murchison EP, Partridge JF, Tam OH, Cheloufi S, Hannon GJ . Characterization of Dicer-deficient murine embryonic stem cells. Proc Natl Acad Sci USA 2005; 102: 12135–12140.
Yi R, O′Carroll D, Pasolli HA, Zhang Z, Dietrich FS, Tarakhovsky A et al. Morphogenesis in skin is governed by discrete sets of differentially expressed microRNAs. Nat Genet 2006; 38: 356–362.
Andl T, Murchison EP, Liu F, Zhang Y, Yunta-Gonzalez M, Tobias JW et al. The miRNA-processing enzyme dicer is essential for the morphogenesis and maintenance of hair follicles. Curr Biol 2006; 16: 1041–1049.
Sonkoly E, Wei T, Janson PC, Sääf A, Lundeberg L, Tengvall-Linder M et al. A MicroRNAs: novel regulators involved in the pathogenesis of Psoriasis? PLoS ONE 2007; 2: e610.
Rossi M, Aqeilan RI, Neale M, Candi E, Salomoni P, Knight RA et al. The E3 ubiquitin ligase Itch controls the protein stability of p63. Proc Natl Acad Sci USA 2006; 103: 12753–12758.
Candi E, Rufini A, Terrinoni A, Giamboi-Miraglia A, Lena AM, Mantovani R et al. DeltaNp63 regulates thymic development through enhanced expression of FgfR2 and Jag2. Proc Natl Acad Sci USA 2007; 104: 11999–12004.
Candi E, Rufini A, Terrinoni A, Dinsdale D, Ranalli M, Paradisi A et al. Differential roles of p63 isoforms in epidermal development: selective genetic complementation in p63 null mice. Cell Death Differ 2006; 13: 1037–1047.
Lee H, Kimelman D . A dominant-negative form of p63 is required for epidermal proliferation in zebrafish. Dev Cell 2002; 2: 607–616.
Truong AB, Kretz M, Ridky TW, Kimmel R, Khavari PA . p63 regulates proliferation and differentiation of developmentally mature keratinocytes. Genes Dev 2006; 20: 3185–3197.
Barrandon Y, Green H . Three clonal types of keratinocyte with different capacities for multiplication. Proc Natl Acad Sci USA 1987; 84: 2302–2306.
Gambaro K, Aberdam E, Virolle T, Aberdam D, Rouleau M . BMP-4 induces a Smad-dependent apoptotic cell death of mouse embryonic stem cell-derived neural precursors. Cell Death Differ 2006; 13: 1075–1087.
Aberdam D, Gambaro K, Medawar A, Aberdam E, Rostagno P, de la Forest Divonne S et al. Embryonic stem cells as a cellular model for neuroectodermal commitment and skin formation. C R Biol 2007; 330: 479–484.
Rocco JW, Leong CO, Kuperwasser N, DeYoung MP, Ellisen LW . p63 mediates survival in squamous cell carcinoma by suppression of p73-dependent apoptosis. Cancer Cell 2006; 9: 45–56.
Tonon G, Wong KK, Maulik G, Brennan C, Feng B, Zhang Y et al. High-resolution genomic profiles of human lung cancer. Proc Natl Acad Sci USA 2005; 102: 9625–9630.
Hibi K, Trink B, Patturajan M, Westra WH, Caballero OL, Hill DE et al. AIS is an oncogene amplified in squamous cell carcinoma. Proc Natl Acad Sci USA 2000; 97: 5462–5467.
Forastiere A, Koch W, Trotti A, Sidransky D . Head and neck cancer. N Engl J Med 2001; 345: 1890–1900.
Rocco JW, Ellisen LW . p63 and p73: life and death in squamous cell carcinoma. Cell Cycle 2006; 5: 936–940.
Ratovitski E, Trink B, Sidransky D . p63 and p73: teammates or adversaries? Cancer Cell 2006; 9: 1–2.
DeYoung MP, Johannessen CM, Leong CO, Faquin W, Rocco JW, Ellisen LW . Tumor-specific p73 up-regulation mediates p63 dependence in squamous cell carcinoma. Cancer Res 2006; 66: 9362–9368.
Yi R, Poy MN, Stoffel M, Fuchs E . A skin microRNA promotes differentiation by repressing ‘stemness’. Nature 2008; 452: 225–229.
Zangen R, Ratovitski E, Sidransky D . DeltaNp63alpha levels correlate with clinical tumor response to cisplatin. Cell Cycle 2005; 4: 1313–1315.
Yuspa SH, Kilkenny AE, Steinert PM, Roop DR . Expression of murine epidermal differentiation markers is tightly regulated by restricted extracellular calcium concentrations in vitro. J Cell Biol 1989; 109: 1207–1217.
Lewis BP, Shih IH, Jones-Rhoades MW, Bartel DP, Burge CB . Prediction of mammalian microRNA targets. Cell 2003; 115: 787–798.
Acknowledgements
We thank Dr. Alessandro Giamboi Miraglia for technical support and Angelo Peschiaroli and Alessandro Terrinoni for scientific discussion. The work reported in this manuscript has been supported by the Medical Research Council (GM) and by grants from Telethon (GGP02251 to EC), AIRC (2743 to GM), EU (LSGBH-2005-019067-Epistem; LSHC-CT-2004-503576-Active p53) to GM, MIUR to GM, PRIN 06 to EC, PRIN 06 to GM, MinSan to GM, and by ANR (ANR-06-BLAN-0367) and EU (LSGBH-2005-019067-Epistem) to DA.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Edited by P Vandenabeele
Supplementary Information accompanies the paper on Cell Death and Differentiation website (http://www.nature.com/cdd)
Supplementary information
Rights and permissions
About this article
Cite this article
Lena, A., Shalom-Feuerstein, R., di Val Cervo, P. et al. miR-203 represses ‘stemness’ by repressing ΔNp63. Cell Death Differ 15, 1187–1195 (2008). https://doi.org/10.1038/cdd.2008.69
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cdd.2008.69
Keywords
This article is cited by
-
p63: a crucial player in epithelial stemness regulation
Oncogene (2023)
-
Identification and functional validation of SRC and RAPGEF1 as new direct targets of miR-203, involved in regulation of epidermal homeostasis
Scientific Reports (2023)
-
Differentially expressed miRNAs associated with generalized aggressive periodontitis
Clinical Oral Investigations (2023)
-
Reversal of dual epigenetic repression of non-canonical Wnt-5a normalises diabetic corneal epithelial wound healing and stem cells
Diabetologia (2023)
-
Tear miRNA expression analysis reveals miR-203 as a potential regulator of corneal epithelial cells
BMC Ophthalmology (2021)