Abstract
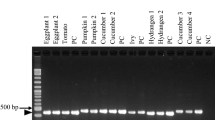
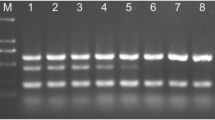
Phytophthora nicotianae Breda de Haan is one of the most important soil-borne plant pathogens. The identification of this pathogen based on morphological or physiological characters is time-consuming and labour-intensive and requires comprehensive knowledge of fungi. Molecular analysis of the internal transcribed spacer (ITS) regions of rDNA is a novel and very effective method of species determination. Based on this concept, conventional and single closed tube nested-PCRs were developed for the specific and sensitive detection of P. nicotianae. Two new specific primers, designed from the spacer regions ITS1 and ITS2, internal to the nucleotide sequence flanked by universal primers ITS4 and ITS6, were used. To evaluate the specificity of the method, 36 morphologically characterized isolates were tested. A positive reaction, characterized by an amplification product of 737 bp, was shown by all P. nicotianae isolates and two P. nicotianae/cactorum hybrids. No amplification product was observed when other Phytophthora species and genera were assayed. The sensitivity of this method was analysed by serial dilutions of a defined amount of fungal DNA in a healthy root extract. Nested-PCR was at least 1000 times more sensitive than conventional PCR. In addition, samples from different infection sites, origins and crops, samples from nutrient solution, water and the rockwool used in hydroponic cultures, were analysed to validate this method.
Similar content being viewed by others
References
Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W and Lipman DJ (1997) Gapped BLAST and PSI-BLAST: A new generation of protein databases search programs. Nucleic Acids Research 25: 3389–3402
Atherton JG and Rudich J (1986) The Tomato Crop. Chapman and Hall, London
Barr DJS (1992) Evolution and kingdoms of organisms from the perspective of a mycologist. Mycologia 84: 1–11
Bonants P, Hagenaar-de Weerdt M, Van Gent-Pelzer M, Lacourt I, Cooke D and Duncan J (1997) Detection and identification of Phytophthora fragariae Hickmann by the polymerase chain reaction. European Journal of Plant Pathology 103: 345–355
Bruce KD, Hioms WD, Hobmann JL, Osborn AM, Strike P and Ritschie DA (1992) Amplification of DNA from native population of soil bacteria by using the Polymerase Chain Reaction. Applied and Environmental Microbiology 58: 3413–3416
Cavalier-Smith T (1989) The kingdom Chromista. In: Green JC, Leadbeater BSC and Diver WL (eds) The Chromophyte Algae: Problems and Perspectives (Syst. Assoc. Spec.), Vol 38 (pp 381–407) Clarendon Press, Oxford, 429 pp
Coelho C, Cravador A, Bollen A, Ferraz JFP, Moreira AC, Fauconnier D and Godfroid E (1997) Highly specific and sensitive non-radioactive molecular identification of P. cinnamomi. Mycological Research 101: 1499–1507
Cooke DEL and Duncan JM (1997) Phylogenetics analysis of Phytophthora species based on ITS1 and ITS2 sequences of the ribosomal RNA gene repeat. Mycological Research 101: 667–677
Cooke DEL, Drenth A, Duncan JM, Wagels G and Brasier CM (2000) A molecular phylogeny of Phytophthora and related Oomycetes. Fungal Genetics and Biology 30: 17–32
Davies JML (1980) Disease in NFT. Acta Horticulturae 98: 299–305
Duniway JM (1977) Changes in resistance to water transport in safflower during the development of Phytophthora root rot. Phytopathology 67: 331–337
Dyer PS, Furneaux PA, Douhan G and Murray TD (2001) A multiplex PCR test for determination of mating type applied to the plant pathogens Tapesia yallundae and Tapesia acuformis. Fungal Genetics and Biology 33: 173–180
Ersek T, Schoelz JE and English JT (1994) PCR amplification of species-specific DNA sequence can distinguish among Phytophthora species. Applied and Environmental Microbiology 60: 2616–3621
Erwin DC and Ribeiro OK (1996) Phytophthora disease worldwide. American Phytopathological Society, St. Paul, 550pp
Goodwin PH, English JT, Neher DA, Duniway JM and Kirkpatrick BC (1990) Detection of Phytophthora parasitica from soil and host tissue with species specific DNA probe. Phytopathology 80: 277–281
Grote D and Bucsi C (1992) Control of Phytophthora nicotianae var. nicotianae on tomatoes cultivated in soilless culture under glasshouse conditions. Gartenbauwissenschaft 57: 183–189
Grote D and Bucsi C (1998) Chemical control of Phytophthora nicotianae on tomato roots with Fosetyl-Aluminium in hydroponics. Gartenbauwissenschaft 63: 78–87
Grote D and Gabler J (1999) Quantification of Phytophthora nicotianae in tomato plants. Journal of Plant Disease and Protection 106: 445–454
Grote D and Claussen W (2001) Severity of root rot on tomato plants caused by Phytophthora nicotianae under nutrient and light stress conditions. Plant Pathology 50: 702–707
Hering O and Nirenberg HI (1995) Differentiation of Fusarium sambucinum Fuckel sensu lato and related species by RAPD PCR. Mycopathologia 129: 159–164
Hickmann CJ (1958) Phytophthora plant destroyer. Transactions of the British Mycological Society 41: 1–13
Jaeger EE, Carroll NM, Choudhury S, Dunlop AA, Towler HM, Matheson MM, Adamson P, Okhravi N and Lightman S (2000) Rapid detection and identification of Candida, Aspergillus, and Fusarium species in ocular samples using nested PCR. Journal of Clinical Microbiology 38: 2902–2908
Jones K and Shew HD (1988) Immunoassay procedure for the detection of Phytophthora nicotianae var. nicotianae in soil (abstr.). Phytopathology 78: 1577
Lacourt I and Duncan JM (1997) Specific detection of Phytophthora nicotianae using the polymerase chain reaction and primers based on the DNA sequences of its elicitin gene ParA1. European Journal of Plant Pathology 103: 73–83
Lee SB and Taylor JW (1990) Isolation of DNA from fungal mycelia and single spores. In: PCR Protocols: A Guide to Methods and Applications (pp 282–287) Academic Press, New York
Lee SB and Taylor JW (1992) Phylogeny of five fungus-like protocristan Phytophthora species, inferred from the internal transcribed spacers of ribosomal DNA. Journal of Molecular and Biological Evolution 9: 636–659
McDonald JD, Stines J and Kabashima J (1990) Comparison if serological and culture plate methods for detecting species of Phytophthora, Pythium and Rhizoctonia in ornamental plants. Plant Disease 78: 607–611
Olmos A, Cambra M, Esteban O, Gorris MT and Terrada E (1999) New device and method for capture, reverse transcription and nested PCR in a single closed-tube. Nucleic Acids Research 27: 1564–1565
Olmos A, Esteban O, Bertolini E and Cambra M (2002) Nested-PCR in a single closed tube. In: PCR Protocols: Methods and Applications. Humana Press (in press)
Picard C, Ponsennet C, Paget E, Nesmo X and Simonet P (1992) Detection and enumeration of bacteria in soil by direct DNA extraction and Polymerase Chain Reaction. Applied and Environmental Microbiology 58: 2717–2722
Tsao PH (1970) Selective media for isolation of pathogenic fungi. Annual Review of Phytopathology 8: 157–186
Tsushima S, Hasebe A, Komoto Y, Carter JP, Miyashita K, Yokoyama K and Pickup RW (1995) Detection of genetically engineered microorganism in paddy soil using a simple and rapid ‘nested’ polymerase chain reaction method. Soil Biology and Biochemistry 27: 219–227
Waterhouse GM (1963) Key to the species of Phytophthora de Bary. Mycological Paper 92: 22 pp. Commonwealth Mycological Institut Kew, UK
White TB, Bruns T, Lee S and Talor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: PCR Protocols: A Guide to Methods and Applications (pp 315–321) Academic Press, New York
Wilson IG (1997) Inhibition and facilitation of nucleic acid amplification. Applied and Environmental Microbiology 63: 3741–3751
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Grote, D., Olmos, A., Kofoet, A. et al. Specific and Sensitive Detection of Phytophthora nicotianae By Simple and Nested-PCR. European Journal of Plant Pathology 108, 197–207 (2002). https://doi.org/10.1023/A:1015139410793
Issue Date:
DOI: https://doi.org/10.1023/A:1015139410793