Abstract
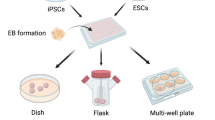
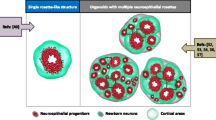
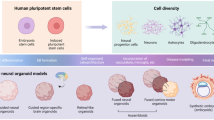
Autism is a neurodevelopmental disease caused by multiple mutations during development. However, a suitable disease model to study the molecular pathway of disease onset and progression is not available. Although many studies have used human stem cells such as induced pluripotent stem cells and embryonic stem cells to investigate the disease pathogenesis, these stem cell techniques are limited in their abilities to study the pathology and mechanism of pathogenesis of neurodevelopmental diseases such as autism. Therefore, researchers are focusing on the strengths of three-dimensional (3D) structures mimicking organs, organoids, for modeling autism. In this review, we highlight the advantages of 3D organoid systems to investigate the mechanisms of the pathogenesis of autism. Further, because the onset of autism is determined by genetic background, we suggest the application of the clustered regularly interspersed short palindromic repeat-associated protein 9 (CRISPR/Cas9) technique for genome editing in 3D organoid systems to study mutations that cause autism. We propose that 3D organoid systems combined with the CRISPR/Cas9 technique may advance autism research.

Similar content being viewed by others
References
Orlacchio A et al (2010) Stem cells: an overview of the current status of therapies for central and peripheral nervous system diseases. Curr Med Chem 17(7):595–608
Martelli MF et al (2014) HLA-haploidentical transplantation with regulatory and conventional T-cell adoptive immunotherapy prevents acute leukemia relapse. Blood 124(4):638–644
Bethge WA et al (2008) Haploidentical allogeneic hematopoietic cell transplantation in adults using CD3/CD19 depletion and reduced intensity conditioning: an update. Blood Cells Mol Dis 40(1):13–19
Simsek, S., et al. 2016, Modeling cystic fibrosis using pluripotent stem cell-derived human pancreatic ductal epithelial cells. Stem Cells Transl Med
Qin M et al (2016) Direct reprogramming of human amniotic fluid stem cells by OCT4 and application in repairing of cerebral ischemia damage. Int J Biol Sci 12(5):558–568
Dutta RC, Dutta AK (2009) Cell-interactive 3D-scaffold; advances and applications. Biotechnol Adv 27(4):334–339
Subramanian A, Krishnan UM, Sethuraman S (2009) Development of biomaterial scaffold for nerve tissue engineering: biomaterial mediated neural regeneration. J Biomed Sci 16:108
Tibbitt MW, Anseth KS (2009) Hydrogels as extracellular matrix mimics for 3D cell culture. Biotechnol Bioeng 103(4):655–663
Brito C et al (2012) 3D cultures of human neural progenitor cells: dopaminergic differentiation and genetic modification. [corrected]. Methods 56(3):452–460
Lancaster MA, Knoblich JA (2014) Organogenesis in a dish: modeling development and disease using organoid technologies. Science 345(6194):1247125
Yin X et al (2014) Niche-independent high-purity cultures of Lgr5+ intestinal stem cells and their progeny. Nat Methods 11(1):106–112
Takasato M et al (2014) Directing human embryonic stem cell differentiation towards a renal lineage generates a self-organizing kidney. Nat Cell Biol 16(1):118–126
Lancaster MA et al (2013) Cerebral organoids model human brain development and microcephaly. Nature 501(7467):373–379
Mondrinos MJ et al (2014) Engineering de novo assembly of fetal pulmonary organoids. Tissue Eng Part A 20(21–22):2892–2907
Mariani J et al (2015) FOXG1-dependent dysregulation of GABA/glutamate neuron differentiation in autism spectrum disorders. Cell 162(2):375–390
Lai MC, Lombardo MV, Baron-Cohen S (2014) Autism. Lancet 383(9920):896–910
Visser JC et al (2016) Autism spectrum disorder and attention-deficit/hyperactivity disorder in early childhood: a review of unique and shared characteristics and developmental antecedents. Neurosci Biobehav Rev 65:229–263
Mazurek MO et al (2013) Anxiety, sensory over-responsivity, and gastrointestinal problems in children with autism spectrum disorders. J Abnorm Child Psychol 41(1):165–176
Christensen DL et al (2016) Prevalence and characteristics of autism spectrum disorder among children aged 8 years—autism and developmental disabilities monitoring network, 11 sites, United States, 2012. MMWR Surveill Summ 65(3):1–23
Gaugler T et al (2014) Most genetic risk for autism resides with common variation. Nat Genet 46(8):881–885
Klei L et al (2012) Common genetic variants, acting additively, are a major source of risk for autism. Mol Autism 3(1):9
Fatehullah A, Tan SH, Barker N (2016) Organoids as an in vitro model of human development and disease. Nat Cell Biol 18(3):246–254
Takahashi K et al (2007) Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell 131(5):861–872
Thomson JA et al (1998) Embryonic stem cell lines derived from human blastocysts. Science 282(5391):1145–1147
Sasai Y (2013) Next-generation regenerative medicine: organogenesis from stem cells in 3D culture. Cell Stem Cell 12(5):520–530
Sato T et al (2009) Single Lgr5 stem cells build crypt-villus structures in vitro without a mesenchymal niche. Nature 459(7244):262–265
Eiraku M et al (2011) Self-organizing optic-cup morphogenesis in three-dimensional culture. Nature 472(7341):51–56
Suga H et al (2011) Self-formation of functional adenohypophysis in three-dimensional culture. Nature 480(7375):57–62
Sato T et al (2011) Long-term expansion of epithelial organoids from human colon, adenoma, adenocarcinoma, and Barrett’s epithelium. Gastroenterology 141(5):1762–1772
Takebe T et al (2013) Vascularized and functional human liver from an iPSC-derived organ bud transplant. Nature 499(7459):481–484
Schlaermann P et al (2016) A novel human gastric primary cell culture system for modelling helicobacter pylori infection in vitro. Gut 65(2):202–213
Bartfeld S et al (2015) In vitro expansion of human gastric epithelial stem cells and their responses to bacterial infection. Gastroenterology 148(1):126–136 e6
Huch M et al (2015) Long-term culture of genome-stable bipotent stem cells from adult human liver. Cell 160(1–2):299–312
Sampaziotis F et al (2015) Cholangiocytes derived from human induced pluripotent stem cells for disease modeling and drug validation. Nat Biotechnol 33(8):845–852
Ogawa M et al (2015) Directed differentiation of cholangiocytes from human pluripotent stem cells. Nat Biotechnol 33(8):853–861
Takasato M et al (2015) Kidney organoids from human iPS cells contain multiple lineages and model human nephrogenesis. Nature 526(7574):564–568
Eiraku M et al (2008) Self-organized formation of polarized cortical tissues from ESCs and its active manipulation by extrinsic signals. Cell Stem Cell 3(5):519–532
Gaspard N et al (2008) An intrinsic mechanism of corticogenesis from embryonic stem cells. Nature 455(7211):351–357
Mariani J et al (2012) Modeling human cortical development in vitro using induced pluripotent stem cells. Proc Natl Acad Sci U S A 109(31):12770–12775
Evans M (2011) Discovering pluripotency: 30 years of mouse embryonic stem cells. Nat Rev Mol Cell Biol 12(10):680–686
Xia X, Zhang SC (2009) Differentiation of neuroepithelia from human embryonic stem cells. Methods Mol Biol 549:51–58
Weitzer G (2006) Embryonic stem cell-derived embryoid bodies: an in vitro model of eutherian pregastrulation development and early gastrulation. Handb Exp Pharmacol 174:21–51
Nakano T et al (2012) Self-formation of optic cups and storable stratified neural retina from human ESCs. Cell Stem Cell 10(6):771–785
Watanabe K et al (2005) Directed differentiation of telencephalic precursors from embryonic stem cells. Nat Neurosci 8(3):288–296
Watanabe K et al (2007) A ROCK inhibitor permits survival of dissociated human embryonic stem cells. Nat Biotechnol 25(6):681–686
Cyranoski D (2012) Tissue engineering: the brainmaker. Nature 488(7412):444–446
Danjo T et al (2011) Subregional specification of embryonic stem cell-derived ventral telencephalic tissues by timed and combinatory treatment with extrinsic signals. J Neurosci 31(5):1919–1933
Su HL et al (2006) Generation of cerebellar neuron precursors from embryonic stem cells. Dev Biol 290(2):287–296
Muguruma K et al (2010) Ontogeny-recapitulating generation and tissue integration of ES cell-derived Purkinje cells. Nat Neurosci 13(10):1171–1180
Lancaster MA, Knoblich JA (2014) Generation of cerebral organoids from human pluripotent stem cells. Nat Protoc 9(10):2329–2340
Fombonne E (2009) Epidemiology of pervasive developmental disorders. Pediatr Res 65(6):591–598
Elsabbagh M et al (2012) Global prevalence of autism and other pervasive developmental disorders. Autism Res 5(3):160–179
Geschwind DH, Levitt P (2007) Autism spectrum disorders: developmental disconnection syndromes. Curr Opin Neurobiol 17(1):103–111
Barger BD, Campbell JM, McDonough JD (2013) Prevalence and onset of regression within autism spectrum disorders: a meta-analytic review. J Autism Dev Disord 43(4):817–828
Robinson EB et al (2011) Evidence that autistic traits show the same etiology in the general population and at the quantitative extremes (5%, 2.5%, and 1%). Arch Gen Psychiatry 68(11):1113–1121
Lundstrom S et al (2012) Autism spectrum disorders and autistic like traits: similar etiology in the extreme end and the normal variation. Arch Gen Psychiatry 69(1):46–52
Ronald A et al (2006) Genetic heterogeneity between the three components of the autism spectrum: a twin study. J Am Acad Child Adolesc Psychiatry 45(6):691–699
Hallmayer J et al (2011) Genetic heritability and shared environmental factors among twin pairs with autism. Arch Gen Psychiatry 68(11):1095–1102
De Rubeis S, Buxbaum JD (2015) Genetics and genomics of autism spectrum disorder: embracing complexity. Hum Mol Genet 24(R1):R24–R31
Abrahams BS, Geschwind DH (2008) Advances in autism genetics: on the threshold of a new neurobiology. Nat Rev Genet 9(5):341–355
Jiang YH et al (2013) Detection of clinically relevant genetic variants in autism spectrum disorder by whole-genome sequencing. Am J Hum Genet 93(2):249–263
Iossifov I et al (2014) The contribution of de novo coding mutations to autism spectrum disorder. Nature 515(7526):216–221
Sutcliffe JS et al (2005) Allelic heterogeneity at the serotonin transporter locus (SLC6A4) confers susceptibility to autism and rigid-compulsive behaviors. Am J Hum Genet 77(2):265–279
Ma DQ et al (2010) Association and gene-gene interaction of SLC6A4 and ITGB3 in autism. Am J Med Genet B Neuropsychiatr Genet 153B(2):477–483
Rudie JD et al (2012) Autism-associated promoter variant in MET impacts functional and structural brain networks. Neuron 75(5):904–915
Yang SY et al (2010) Family-based association study of microsatellites in the 5’ flanking region of AVPR1A with autism spectrum disorder in the Korean population. Psychiatry Res 178(1):199–201
Yamasue H (2016) Promising evidence and remaining issues regarding the clinical application of oxytocin in autism spectrum disorders. Psychiatry Clin Neurosci 70(2):89–99
Persico AM, Napolioni V (2013) Autism genetics. Behav Brain Res 251:95–112
Wang L et al (2008) Association of the ENGRAILED 2 (EN2) gene with autism in Chinese Han population. Am J Med Genet B Neuropsychiatr Genet 147B(4):434–438
Lee EJ, Choi SY, Kim E (2015) NMDA receptor dysfunction in autism spectrum disorders. Curr Opin Pharmacol 20:8–13
Yoo HJ et al (2012) Family based association of GRIN2A and GRIN2B with Korean autism spectrum disorders. Neurosci Lett 512(2):89–93
Skaar DA et al (2005) Analysis of the RELN gene as a genetic risk factor for autism. Mol Psychiatry 10(6):563–571
Buxbaum JD et al (2002) Association between a GABRB3 polymorphism and autism. Mol Psychiatry 7(3):311–316
Kim SA et al (2006) Association of GABRB3 polymorphisms with autism spectrum disorders in Korean trios. Neuropsychobiology 54(3):160–165
Anney R et al (2010) A genome-wide scan for common alleles affecting risk for autism. Hum Mol Genet 19(20):4072–4082
Connolly JJ, Glessner JT, Hakonarson H (2013) A genome-wide association study of autism incorporating autism diagnostic interview-revised, autism diagnostic observation schedule, and social responsiveness scale. Child Dev 84(1):17–33
Stolerman ES et al (2016) CHD8 intragenic deletion associated with autism spectrum disorder. Eur J Med Genet 59(4):189–194
Chaste P et al (2015) A genome-wide association study of autism using the Simons simplex collection: does reducing phenotypic heterogeneity in autism increase genetic homogeneity? Biol Psychiatry 77(9):775–784
Liu X et al (2016) Genome-wide association study of autism spectrum disorder in the east Asian populations. Autism Res 9(3):340–349
Banerjee S, Riordan M, Bhat MA (2014) Genetic aspects of autism spectrum disorders: insights from animal models. Front Cell Neurosci 8:58
Chen J et al (2014) Synaptic proteins and receptors defects in autism spectrum disorders. Front Cell Neurosci 8:276
Zoghbi HY, Bear MF (2012) Synaptic dysfunction in neurodevelopmental disorders associated with autism and intellectual disabilities. Cold Spring Harb Perspect Biol 4(3). doi:10.1101/cshperspect.a009886
Ebert DH, Greenberg ME (2013) Activity-dependent neuronal signalling and autism spectrum disorder. Nature 493(7432):327–337
Neuhofer D et al (2015) Functional and structural deficits at accumbens synapses in a mouse model of fragile X. Front Cell Neurosci 9:100
Doll CA, Broadie K (2014) Impaired activity-dependent neural circuit assembly and refinement in autism spectrum disorder genetic models. Front Cell Neurosci 8:30
Port RG et al (2014) Convergence of circuit dysfunction in ASD: a common bridge between diverse genetic and environmental risk factors and common clinical electrophysiology. Front Cell Neurosci 8:414
Willsey AJ et al (2013) Coexpression networks implicate human midfetal deep cortical projection neurons in the pathogenesis of autism. Cell 155(5):997–1007
Amenduni M et al (2011) iPS cells to model CDKL5-related disorders. Eur J Hum Genet 19(12):1246–1255
Marchetto MC et al (2010) A model for neural development and treatment of Rett syndrome using human induced pluripotent stem cells. Cell 143(4):527–539
Urbach A et al (2010) Differential modeling of fragile X syndrome by human embryonic stem cells and induced pluripotent stem cells. Cell Stem Cell 6(5):407–411
Sheridan SD et al (2011) Epigenetic characterization of the FMR1 gene and aberrant neurodevelopment in human induced pluripotent stem cell models of fragile X syndrome. PLoS One 6(10):e26203
Kyttala A et al (2016) Genetic variability overrides the impact of parental cell type and determines iPSC differentiation potential. Stem Cell Reports 6(2):200–212
Mills JA et al (2013) Clonal genetic and hematopoietic heterogeneity among human-induced pluripotent stem cell lines. Blood 122(12):2047–2051
Butler MG et al (2005) Subset of individuals with autism spectrum disorders and extreme macrocephaly associated with germline PTEN tumour suppressor gene mutations. J Med Genet 42(4):318–321
Bourguignon C, Li J, Papalopulu N (1998) XBF-1, a winged helix transcription factor with dual activity, has a role in positioning neurogenesis in Xenopus competent ectoderm. Development 125(24):4889–4900
Schwank G et al (2013) Functional repair of CFTR by CRISPR/Cas9 in intestinal stem cell organoids of cystic fibrosis patients. Cell Stem Cell 13(6):653–658
McCracken KW et al (2014) Modelling human development and disease in pluripotent stem-cell-derived gastric organoids. Nature 516(7531):400–404
Lugo JN et al (2014) Deletion of PTEN produces autism-like behavioral deficits and alterations in synaptic proteins. Front Mol Neurosci 7:27
Tilot AK, Frazier TW 2nd, Eng C (2015) Balancing proliferation and connectivity in PTEN-associated autism spectrum disorder. Neurotherapeutics 12(3):609–619
Amiri A et al (2012) Pten deletion in adult hippocampal neural stem/progenitor cells causes cellular abnormalities and alters neurogenesis. J Neurosci 32(17):5880–5890
Rogers JT et al (2011) Reelin supplementation enhances cognitive ability, synaptic plasticity, and dendritic spine density. Learn Mem 18(9):558–564
Wang Z et al (2014) Reelin gene variants and risk of autism spectrum disorders: an integrated meta-analysis. Am J Med Genet B Neuropsychiatr Genet 165B(2):192–200
Bailey A et al (1998) A clinicopathological study of autism. Brain 121(Pt 5):889–905
Nguyen A et al (2010) Global methylation profiling of lymphoblastoid cell lines reveals epigenetic contributions to autism spectrum disorders and a novel autism candidate gene, RORA, whose protein product is reduced in autistic brain. FASEB J 24(8):3036–3051
Zhu L et al (2014) Epigenetic dysregulation of SHANK3 in brain tissues from individuals with autism spectrum disorders. Hum Mol Genet 23(6):1563–1578
Qin J et al (2009) Association study of SHANK3 gene polymorphisms with autism in Chinese Han population. BMC Med Genet 10:61
Talebizadeh Z et al (2004) Do known mutations in neuroligin genes (NLGN3 and NLGN4) cause autism? J Autism Dev Disord 34(6):735–736
Jamain S et al (2003) Mutations of the X-linked genes encoding neuroligins NLGN3 and NLGN4 are associated with autism. Nat Genet 34(1):27–29
Arden KC (2004) FoxO: linking new signaling pathways. Mol Cell 14(4):416–418
Seoane J et al (2004) Integration of Smad and forkhead pathways in the control of neuroepithelial and glioblastoma cell proliferation. Cell 117(2):211–223
Yao J, Lai E, Stifani S (2001) The winged-helix protein brain factor 1 interacts with groucho and hes proteins to repress transcription. Mol Cell Biol 21(6):1962–1972
Martynoga B et al (2005) Foxg1 is required for specification of ventral telencephalon and region-specific regulation of dorsal telencephalic precursor proliferation and apoptosis. Dev Biol 283(1):113–127
Yang Y et al (2015) Impaired interneuron development after Foxg1 disruption. Cereb Cortex. doi:10.1093/cercor/bhv297
Roche-Martinez A et al (2011) FOXG1, a new gene responsible for the congenital form of Rett syndrome. Rev Neurol 52(10):597–602
Striano P et al (2011) West syndrome associated with 14q12 duplications harboring FOXG1. Neurology 76(18):1600–1602
Nageshappa S et al (2016) Altered neuronal network and rescue in a human MECP2 duplication model. Mol Psychiatry 21(2):178–188
Vogel G (2013) Neurodevelopment. Lab dishes up mini-brains. Science 341(6149):946–947
Jinek M et al (2012) A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337(6096):816–821
Mali P et al (2013) RNA-guided human genome engineering via Cas9. Science 339(6121):823–826
Ran FA et al (2013) Genome engineering using the CRISPR-Cas9 system. Nat Protoc 8(11):2281–2308
Barrangou R et al (2007) CRISPR provides acquired resistance against viruses in prokaryotes. Science 315(5819):1709–1712
Wang T et al (2014) Genetic screens in human cells using the CRISPR-Cas9 system. Science 343(6166):80–84
Shalem O, Sanjana NE, Zhang F (2015) High-throughput functional genomics using CRISPR-Cas9. Nat Rev Genet 16(5):299–311
Kan Y et al (2014) The mechanism of gene targeting in human somatic cells. PLoS Genet 10(4):e1004251
Elliott B et al (1998) Gene conversion tracts from double-strand break repair in mammalian cells. Mol Cell Biol 18(1):93–101
Baum C et al (2004) Chance or necessity? Insertional mutagenesis in gene therapy and its consequences. Mol Ther 9(1):5–13
Musunuru K (2013) Genome editing of human pluripotent stem cells to generate human cellular disease models. Dis Model Mech 6(4):896–904
Grobarczyk B et al (2015) Generation of isogenic human iPS cell line precisely corrected by genome editing using the CRISPR/Cas9 system. Stem Cell Rev 11(5):774–787
Cong L et al (2013) Multiplex genome engineering using CRISPR/Cas systems. Science 339(6121):819–823
Xie F et al (2014) Seamless gene correction of beta-thalassemia mutations in patient-specific iPSCs using CRISPR/Cas9 and piggyBac. Genome Res 24(9):1526–1533
Su S et al (2016) CRISPR-Cas9 mediated efficient PD-1 disruption on human primary T cells from cancer patients. Sci Rep 6:20070
Young CS et al (2016) A single CRISPR-Cas9 deletion strategy that targets the majority of DMD patients restores dystrophin function in hiPSC-derived muscle cells. Cell Stem Cell 18(4):533–540
Park CY et al (2015) Functional correction of large factor VIII gene chromosomal inversions in hemophilia a patient-derived iPSCs using CRISPR-Cas9. Cell Stem Cell 17(2):213–220
Mungenast AE, Siegert S, Tsai LH (2015) Modeling Alzheimer’s disease with human induced pluripotent stem (iPS) cells. Mol Cell Neurosci 73:13–31. doi:10.1016/j.mcn.2015.11.010
Matano M et al (2015) Modeling colorectal cancer using CRISPR-Cas9-mediated engineering of human intestinal organoids. Nat Med 21(3):256–262
Freedman BS et al (2015) Modelling kidney disease with CRISPR-mutant kidney organoids derived from human pluripotent epiblast spheroids. Nat Commun 6:8715
Drost J et al (2015) Sequential cancer mutations in cultured human intestinal stem cells. Nature 521(7550):43–47
Li X et al (2014) Oncogenic transformation of diverse gastrointestinal tissues in primary organoid culture. Nat Med 20(7):769–777
Boj SF et al (2015) Organoid models of human and mouse ductal pancreatic cancer. Cell 160(1–2):324–338
Iafrati J et al (2014) Reelin, an extracellular matrix protein linked to early onset psychiatric diseases, drives postnatal development of the prefrontal cortex via GluN2B-NMDARs and the mTOR pathway. Mol Psychiatry 19(4):417–426
Gonzalez F (2016) CRISPR/Cas9 genome editing in human pluripotent stem cells: harnessing human genetics in a dish. Dev Dyn 245(7):788–806
Acknowledgements
This work was supported by the Korea Health Technology R&D Project, Ministry of Health and Welfare (HI16C1176).
Author Contributions
Choi Hwan and Juhyun Song wrote the preliminary draft and revised details of the manuscript. Guiyeon Park drew the figure. Jongpil Kim revised all manuscript in detail.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of Interest
The authors declare that they have no conflicts of interest.
Additional information
Hwan Choi and Juhyun Song are equally contributed
Rights and permissions
About this article
Cite this article
Choi, H., Song, J., Park, G. et al. Modeling of Autism Using Organoid Technology. Mol Neurobiol 54, 7789–7795 (2017). https://doi.org/10.1007/s12035-016-0274-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-016-0274-8