Abstract
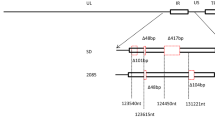
Here, we present the complete genomic sequence of the Chinese standard challenge strain (CSC) of duck enteritis virus (DEV), which was isolated in China in 1962. The DEV CSC genome is 162,131 bp long and contains 78 predicted open reading frames (ORFs). Comparison of the genomic sequences of DEV CSC and DEV live vaccine strain K at passage 63 (DEV K p63) revealed that the DEV CSC genome is 4,040 bp longer than the DEV K p63 genome, mainly because of 3,513-bp and 528-bp insertions at the 5′ and 3′ ends of the unique long segment, respectively. At the nucleotide level, 63 of the 76 ORFs in the DEV CSC genome were 100 % identical to the ORFs in the DEV K p63 genome. Two ORFs (UL56 and US10) had frameshift mutations in the C-terminal regions, while LORF5 was unique to the DEV K p63 genome. It is difficult to assign attenuated virulence to changes in specific genes. However, the complete DEV CSC genome will further advance our understanding of the genes involved in virulence and evolution. The DEV CSC genome sequence has been deposited in GenBank under accession number JQ673560.




Similar content being viewed by others
References
T.S. Sandhu, S.A. Metwally, Diseases of poultry, 12th edn. (Blackwell, Singapore, 2008)
A.E.R.F. Baudet, Tijdschr. Diergeneeskd. 50, 455–459 (1923)
Y.X. Huang, J. South Chin. Agric. Univ. 1, 1–12 (1959)
A.M.Q. King, M.J. Adams, E.B. Carstens, E.J. Lefkowitz, Virus taxonomy: classification and nomenclature of viruses: ninth report of the International Committee on Taxonomy of Viruses (Elsevier, San Diego, 2012)
Y. Li, B. Huang, X. Ma, J. Wu, F. Li, W. Ai, M. Song, H. Yang, Virology 391, 151–161 (2009)
J. Wang, D. Hoper, M. Beer, N. Osterrieder, Virus Res. 160, 316–325 (2011)
Y. Wu, A. Cheng, M. Wang, D. Zhu, R. Jia, S. Chen, Y. Zhou, X. Chen, J. Virol. 86, 13841–13842 (2012)
Y. Wu, A. Cheng, M. Wang, Q. Yang, D. Zhu, R. Jia, S. Chen, Y. Zhou, X. Wang, X. Chen, J. Virol. 86, 5965 (2012)
C. Sinzger, J. Knapp, K. Schmidt, M. Kahl, G. Jahn, J. Virol. Methods 81, 115–122 (1999)
M. Margulies, M. Egholm, W.E. Altman, S. Attiya, J.S. Bader, L.A. Bemben, J. Berka, M.S. Braverman, Y.J. Chen, Z. Chen, S.B. Dewell, L. Du, J.M. Fierro, X.V. Gomes, B.C. Godwin, W. He, S. Helgesen, C.H. Ho, G.P. Irzyk, S.C. Jando, M.L. Alenquer, T.P. Jarvie, K.B. Jirage, J.B. Kim, J.R. Knight, J.R. Lanza, J.H. Leamon, S.M. Lefkowitz, M. Lei, J. Li, K.L. Lohman, H. Lu, V.B. Makhijani, K.E. McDade, M.P. McKenna, E.W. Myers, E. Nickerson, J.R. Nobile, R. Plant, B.P. Puc, M.T. Ronan, G.T. Roth, G.J. Sarkis, J.F. Simons, J.W. Simpson, M. Srinivasan, K.R. Tartaro, A. Tomasz, K.A. Vogt, G.A. Volkmer, S.H. Wang, Y. Wang, M.P. Weiner, P. Yu, R.F. Begley, J.M. Rothberg, Nature 437, 376–380 (2005)
C. Yang, J. Li, Q. Li, H. Li, Y. Xia, X. Guo, K. Yu, H. Yang, Genome Announc 1(5), e00685 (2013)
L.F. Lee, P. Wu, D. Sui, D. Ren, J. Kamil, H.J. Kung, R.L. Witter, Proc. Natl. Acad. Sci. USA 97, 6091–6096 (2000)
A. Krogh, B. Larsson, G. von Heijne, E.L. Sonnhammer, J. Mol. Biol. 305, 567–580 (2001)
A.J. Davison, Vet. Microbiol. 143, 52–69 (2010)
R. Kehm, A. Rosen-Wolff, G. Darai, Virus Res. 40, 17–31 (1996)
I. Mori, B. Liu, F. Goshima, H. Ito, N. Koide, T. Yoshida, T. Yokochi, Y. Kimura, Y. Nishiyama, Microbes Infect. 7, 1492–1500 (2005)
A. Rosen-Wolff, W. Lamade, C. Berkowitz, Y. Becker, G. Darai, Virus Res. 20, 205–221 (1991)
R. Savva, K. McAuley-Hecht, T. Brown, L. Pearl, Nature 373, 487–493 (1995)
K. Arnold, L. Bordoli, J. Kopp, T. Schwede, Bioinformatics 22, 195–201 (2006)
L. Bordoli, F. Kiefer, K. Arnold, P. Benkert, J. Battey, T. Schwede, Nat. Protoc. 4, 1–13 (2009)
N. Guex, M.C. Peitsch, Electrophoresis 18, 2714–2723 (1997)
N. Guex, M.C. Peitsch, T. Schwede, Electrophoresis 30(Suppl 1), S162–S173 (2009)
T. Schwede, J. Kopp, N. Guex, M.C. Peitsch, Nucleic Acids Res. 31, 3381–3385 (2003)
R.B. Pyles, R.L. Thompson, J. Virol. 68, 4963–4972 (1994)
Acknowledgments
The authors would like to thank Dr. Wei Lu for double-checked the English manuscript. This work was supported by Important Animal Pathogens and Biological Reference Substance Research Program (#2008FY130100) from the Chinese Ministry of Science and Technology.
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Yang, C., Li, Q., Li, J. et al. Comparative genomic sequence analysis between a standard challenge strain and a vaccine strain of duck enteritis virus in China. Virus Genes 48, 296–303 (2014). https://doi.org/10.1007/s11262-013-1009-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11262-013-1009-9