Abstract
Due to the essential role of peptide deformylase (PDF) at the bacterial growth cycle, it is a noteworthy target for developing a novel antibacterial agent. In the current study, the antibacterial activities of a set of 44 new structures of formyl hydroxyamino derivatives as PDF inhibitors were quantified using quantitative structure–activity relationship (QSAR). Artificial neural networks (ANN) were used as a chemometrics tool for QSAR modeling. Three quantitative models were suggested to relate the chemical structural features of the formyl hydroxyamino derivatives to their antibacterial activities (pIC50) against Staphylococcus aureus, methicillin-susceptible S. aureus (MSSA), and methicillin-resistant S. aureus (MRSA) peptide deformylase. The sufficiency of the model for prediction of the antibacterial activities of the desired PDF inhibitor compounds against S. aureus, MSSA, and MRSA was statistically demonstrated according to the validation parameters such as coefficient of determination (R2), mean square error (MSE) in training, validation, and prediction sets, and also using applicability domain (AD) and randomization test.





Similar content being viewed by others
References
Meinnel T, Mechulam Y, Blanquet S (1993) Methionine as translation start signal: a review of the enzymes of the pathway in Escherichia coli. Biochimie 75:1061–1075. https://doi.org/10.1016/0300-9084(93)90005-D
Yuan Z, Trias J, White RJ (2001) Deformylase as a novel antibacterial target. Drug Discov Today 6:954–961. https://doi.org/10.1016/S1359-6446(01)01925-0
Waller AS, Clements JM (2002) Novel approaches to antimicrobial therapy: peptide deformylase. Curr Opin Drug Discov Devel 5:785–792
Ni H, Wendoloski J (2006) Structure-based design of new antibacterial agents. Annu Rep Comput Chem 2:279–295
Lee JY, Doddareddy MR, Cho YS et al (2007) Comparative QSAR studies on peptide deformylase inhibitors. J Mol Model 13:543–558. https://doi.org/10.1007/s00894-007-0175-x
Rajagopalan PTR, Yu XC, Pei D (1997) Peptide deformylase: a new type of mononuclear iron protein. J Am Chem Soc 119:12418–12419. https://doi.org/10.1021/ja9734096
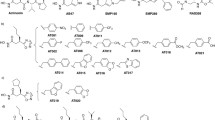
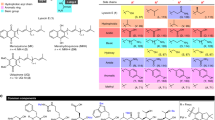
Yang S, Shi W, Xing D et al (2014) Synthesis, antibacterial activity, and biological evaluation of formyl hydroxyamino derivatives as novel potent peptide deformylase inhibitors against drug-resistant bacteria. Eur J Med Chem 86:133–152. https://doi.org/10.1016/j.ejmech.2014.07.106
Ekins S, Mestres J, Testa B (2007) In silico pharmacology for drug discovery: applications to targets and beyond. Br J Pharmacol 152:21–37. https://doi.org/10.1038/sj.bjp.0707306
Cai J, Han C, Hu T et al (2006) Peptide deformylase is a potential target for anti- Helicobacter pylori drugs: reverse docking, enzymatic assay, and X-ray crystallography validation. Protein Sci 15:2071–2081. https://doi.org/10.1110/ps.062238406
Yousefinejad S, Hemmateenejad B (2015) Chemometrics tools in QSAR/QSPR studies: a historical perspective. Chemom Intell Lab Syst 149:177–204. https://doi.org/10.1016/j.chemolab.2015.06.016
Pirhadi S, Shiri F, Ghasemi JB (2015) Multivariate statistical analysis methods in QSAR. RSC Adv 5:104635–104665. https://doi.org/10.1039/C5RA10729F
Veselinović JB, Đorđević V, Bogdanović M et al (2018) QSAR modeling of dihydrofolate reductase inhibitors as a therapeutic target for multiresistant bacteria. Struct Chem 29:541–551. https://doi.org/10.1007/s11224-017-1051-7
Kovesdi I, Dominguez-Rodriguez MF, Orfi L et al (1999) Application of neural networks in structure-activity relationships. Med Res Rev 19:249–269. https://doi.org/10.1002/(SICI)1098-1128(199905)19:3<249::AID-MED4>3.0.CO;2-0
Burbidge R, Trotter M, Buxton B, Holden S (2001) Drug design by machine learning: support vector machines for pharmaceutical data analysis. Comput Chem 26:5–14. https://doi.org/10.1016/S0097-8485(01)00094-8
Guha R, Jurs PC (2005) Interpreting computational neural network QSAR models: a measure of descriptor importance. J Chem Inf Model 45:800–806. https://doi.org/10.1021/ci050022a
Gupta MK, Mishra P, Prathipati P, Saxena AK (2002) 2D-QSAR in hydroxamic acid derivatives as peptide deformylase inhibitors and antibacterial agents. Bioorg Med Chem 10:3713–3716. https://doi.org/10.1016/S0968-0896(02)00421-2
Gao J, Cheng Y, Cui W et al (2012) 3D-QSAR and molecular docking studies of hydroxamic acids as peptide deformylase inhibitors. Med Chem Res 21:1597–1610. https://doi.org/10.1007/s00044-011-9672-7
Todeschini R, Consonni V (2009) Molecular descriptors for chemoinformatics, Second. WILEY-VCH, Weinheim
Mauri A, Consonni V, Pavan M, Todeschini R (2006) Dragon software: an easy approach to molecular descriptor calculations. MATCH Commun Math Comput Chem 56:237–248
Hawkins DM (2004) The problem of overfitting. J Chem Inf Comput Sci 44:1–12. https://doi.org/10.1021/ci0342472
Golbraikh A, Tropsha A (2000) Predictive QSAR modeling based on diversity sampling of experimental datasets for the training and test set selection. Mol Divers 5:231–243. https://doi.org/10.1023/A:1021372108686
Gramatica P (2014) External evaluation of QSAR models, in addition to cross-validation: verification of predictive capability on totally new chemicals. Mol Inform 33:311–314. https://doi.org/10.1002/minf.201400030
Gramatica P (2007) Principles of QSAR models validation: internal and external. QSAR Comb Sci 26:694–701. https://doi.org/10.1002/qsar.200610151
Netzeva TI, Worth AP, Aldenberg T et al (2005) Current status of methods for defining the applicability domain of ( quantitative ) structure – activity relationships. ATLA 2:1–19
Dimitrov S, Dimitrova G, Pavlov T et al (2005) A stepwise approach for defining the applicability domain of SAR and QSAR models. J Chem Inf Model 45:839–849. https://doi.org/10.1021/ci0500381
Yousefinejad S, Honarasa F, Montaseri H (2015) Linear solvent structure-polymer solubility and solvation energy relationships to study conductive polymer/carbon nanotube composite solutions. RSC Adv 5:42266–42275. https://doi.org/10.1039/C5RA05930E
Jagiello K, Makurat S, Pereć S et al (2018) Molecular features of thymidine analogues governing the activity of human thymidine kinase. Struct Chem 29:1367–1374. https://doi.org/10.1007/s11224-018-1124-2
Funding
This study was financially supported by Shiraz University of Medical Sciences (grant no. 97-01-42-17151).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
ESM 1
(DOCX 38 kb)
Rights and permissions
About this article
Cite this article
Yousefinejad, S., Mahboubifar, M. & Rasekh, S. Prediction of different antibacterial activity in a new set of formyl hydroxyamino derivatives with potent action on peptide deformylase using structural information. Struct Chem 30, 925–936 (2019). https://doi.org/10.1007/s11224-018-1242-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11224-018-1242-x