Abstract
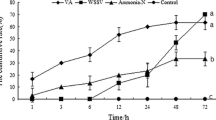
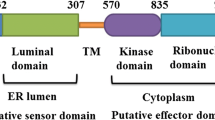
Reactive oxygen species (ROS) play key roles in many physiological processes. In particular, the sterilization mechanism of bacteria using ROS in macrophages is a very important function for biological defense. Xanthine dehydrogenase (XDH) and aldehyde oxidase (AOX), members of the molybdo-flavoenzyme subfamily, are known to generate ROS. Although these enzymes occur in many vertebrates, some insects, and plants, little research has been conducted on XDHs and AOXs in crustaceans. Here, we cloned the entire cDNA sequences of XDH (MjXDH: 4328 bp) and AOX (MjAOX: 4425 bp) from Marsupenaeus japonicus (kuruma shrimp) using reverse transcriptase-polymerase chain reaction (RT-PCR) and random amplification of cDNA ends (RACE). Quantitative real-time RT-PCR transcriptional analysis revealed that MjXDH mRNA is highly expressed in heart and stomach tissues, whereas MjAOX mRNA is highly expressed in the lymphoid organ and intestinal tissues. Furthermore, expression of MjAOX was determined to be up-regulated in the lymphoid organ in response to Vibrio penaeicida at 48 and 72 h after injection; in contrast, hydrogen peroxide (H2O2) concentrations increased significantly at 6, 12, 48, and 72 h after injection with white spot syndrome virus (WSSV) and at 72 h after injection with V. penaeicida. To the best of our knowledge, this study is the first to have identified and cloned XDH and AOX from a crustacean species.






Similar content being viewed by others
References
Valko M, Leibfritz D, Moncol J, Cronin MT, Mazur M, Telser J (2007) Free radicals and antioxidants in normal physiological functions and human disease. Int J Biochem Cell Biol 39(1):44–84. https://doi.org/10.1016/j.biocel.2006.07.001
Cohn CA (2005) Quantifying hydrogen peroxide in iron-containing solutions using leuco crystal violet. Geochem Trans 6(3):47. https://doi.org/10.1063/1.1935449
Graves JA, Metukuri M, Scott D, Rothermund K, Prochownik EV (2008) Regulation of reactive oxygen species homeostasis by peroxiredoxins and c-Myc. J Biol Chem 284(10):6520–6529. https://doi.org/10.1074/jbc.m807564200
Chelikani P, Fita I, Loewen PC (2004) Diversity of structures and properties among catalases. Cell Mol Life Sci 61(2):192–208. https://doi.org/10.1007/s00018-003-3206-5
Nordberg J, Arnér ESJ (2001) Reactive oxygen species, antioxidants, and the mammalian thioredoxin system. Free Radic Biol Med 31:1287–1312
Paglia DE, Valentine WN (1967) Studies on the quantitative and qualitative characterization of erythrocyte glutathione peroxidase. J Lab Clin Med 70:158–169
Juven BJ, Pierson MD (1996) Antibacterial effects of hydrogen peroxide and methods for its detection and quantitation. J Food Prot 59(11):1233–1241. https://doi.org/10.4315/0362-028x-59.11.1233
Skulachev VP (1998) Possible role of reactive oxygen species in antiviral defense. Biochem 63:1438–1440
Iwanaga S, Lee B (2005) Recent advances in the innate immunity of invertebrate animals. BMB Rep 38(2):128–150. https://doi.org/10.5483/bmbrep.2005.38.2.128
Lavine M, Strand M (2002) Insect hemocytes and their role in immunity. Insect Biochem Mol Biol 32(10):1295–1309. https://doi.org/10.1016/s0965-1748(02)00092-9
Manevski N, Balavenkatraman KK, Bertschi B, Swart P, Walles M, Camenisch G, Litherland K (2014) Aldehyde oxidase activity in fresh human skin. Drug Metab Dispos 42(12):2049–2057. https://doi.org/10.1124/dmd.114.060368
Pope SD, Chen L, Stewart V (2008) Purine utilization by Klebsiella oxytoca M5al: genes for ring-oxidizing and -opening enzymes. J Bacteriol 191(3):1006–1017. https://doi.org/10.1128/jb.01281-08
Garattini E, Mendel R, Romão MJ, Wright R, Terao M (2003) Mammalian molybdo-flavoenzymes, an expanding family of proteins: structure, genetics, regulation, function and pathophysiology. Biochem J 372(1):15–32. https://doi.org/10.1042/bj20030121
Llamas A, Chamizo-Ampudia A, Tejada-Jimenez M, Galvan A, Fernandez E (2017) The molybdenum cofactor enzyme mARC: moonlighting or promiscuous enzyme? BioFactors 43(4):486–494. https://doi.org/10.1002/biof.1362
Bray R (1975) 6 Molybdenum iron-sulfur flauin hydroxylases and related enzymes. Enzymes. https://doi.org/10.1016/s1874-6047(08)60229-2
Berry CE, Hare JM (2004) Xanthine oxidoreductase and cardiovascular disease: molecular mechanisms and pathophysiological implications. J Physiol 555(3):589–606. https://doi.org/10.1113/jphysiol.2003.055913
Nishino T, Okamoto K, Eger BT, Pai EF, Nishino T (2008) Mammalian xanthine oxidoreductase—mechanism of transition from xanthine dehydrogenase to xanthine oxidase. FEBS J 275(13):3278–3289. https://doi.org/10.1111/j.1742-4658.2008.06489.x
Ng’oma E, Groth M, Ripa R, Platzer M, Cellerino A (2014) Transcriptome profiling of natural dichromatism in the annual fishes Nothobranchius furzeri and Nothobranchius kadleci. BMC Genomics 15(1):754. https://doi.org/10.1186/1471-2164-15-754
Zhou X, Riddiford LM (2008) rosy function is required for juvenile hormone effects in Drosophila melanogaster. Genetics 178(1):273–281. https://doi.org/10.1534/genetics.107.080754
Terao M, Romão MJ, Leimkühler S, Bolis M, Fratelli M, Coelho C, Garattini E (2016) Structure and function of mammalian aldehyde oxidases. Arch Toxicol 90(4):753–780. https://doi.org/10.1007/s00204-016-1683-1
Kurosaki M, Bolis M, Fratelli M, Barzago MM, Pattini L, Perretta G, Garattini E (2012) Structure and evolution of vertebrate aldehyde oxidases: from gene duplication to gene suppression. Cell Mol Life Sci 70(10):1807–1830. https://doi.org/10.1007/s00018-012-1229-5
Marelja Z, Dambowsky M, Bolis M, Georgiou ML, Garattini E, Missirlis F, Leimkuhler S (2014) The four aldehyde oxidases of Drosophila melanogaster have different gene expression patterns and enzyme substrate specificities. J Exp Biol 217(12):2201–2211. https://doi.org/10.1242/jeb.102129
Gomezanduro G, Barillasmury C, Peregrinouriarte A, Gupta L, Gollasgalvan T, Hernandezlopez J, Yepizplascencia G (2006) The cytosolic manganese superoxide dismutase from the shrimp Litopenaeus vannamei: molecular cloning and expression. Dev Comp Immunol 30(10):893–900. https://doi.org/10.1016/j.dci.2006.01.002
Muñoz M, Cedeño R, Rodríguez J, Van der Knaap WP, Mialhe E, Bachère E (2000) Measurement of reactive oxygen intermediate production in haemocytes of the penaeid shrimp, Penaeus vannamei. Aquaculture 191(1–3):89–107. https://doi.org/10.1016/s0044-8486(00)00420-8
Tacon AG (2017) Biosecure shrimp feeds and feeding practices: guidelines for future development. J World Aquac Soc 48(3):381–392. https://doi.org/10.1111/jwas.12406
Bulbul MG, Kader MA, Asaduzzaman M, Ambak MA, Chowdhury AJ, Hossain MS, Koshio S (2016) Can canola meal and soybean meal be used as major dietary protein sources for kuruma shrimp Marsupenaeus japonicus? Aquaculture 452:194–199. https://doi.org/10.1016/j.aquaculture.2015.10.036
Thitamadee S, Prachumwat A, Srisala J, Jaroenlak P, Salachan PV, Sritunyalucksana K, Itsathitphaisarn O (2016) Review of current disease threats for cultivated penaeid shrimp in Asia. Aquaculture 452:69–87. https://doi.org/10.1016/j.aquaculture.2015.10.028
García-Triana A, Peregrino-Uriarte AB, Yepiz-Plascencia G (2016) Selenoprotein M gene expression, peroxidases activity and hydrogen peroxide concentration are differentially regulated in gill and hepatopancreas of the white shrimp Litopenaeus vannamei during hypoxia and reoxygenation. Comp Biochem Physiol Part A Mol Integr Physiol 199:14–20. https://doi.org/10.1016/j.cbpa.2016.04.019
Messner B, Boll M (1994) Cell suspension cultures of spruce (Picea abies): inactivation of extracellular enzymes by fungal elicitor-induced transient release of hydrogen peroxide (oxidative burst). Plant Cell Tissue Organ Cult 39(1):69–78. https://doi.org/10.1007/bf00037594
Letunic I, Doerks T, Bork P (2011) SMART 7: recent updates to the protein domain annotation resource. Nucleic Acids Res. https://doi.org/10.1093/nar/gkr931
Kobayashi K, Miki M, Okamoto K, Nishino T (1993) Electron transfer process in milk xanthine dehydrogenase as studied by pulse radiolysis. J Biol Chem 268:24642–24646
Olson JS, Ballou DP, Palmer G, Massey V (1974) The mechanism of action of xanthine oxidase. J Biol Chem 249:4363–4382
Kanda M, Brady FO, Rajagopalan KV, Handler P (1972) Studies on the dissociation of flavin adenine dinucleotide from metalloflavoproteins. J Biol Chem 247:765– 70
Komai H, Massey V, Palmer G (1969) The preparation and properties of deflavo xanthine oxidase. J Biol Chem 244:1692–1700
Harrison R (2002) Structure and function of xanthine oxidoreductase: where are we now? Free Radic Biol Med 33(6):774–797. https://doi.org/10.1016/s0891-5849(02)00956-5
Wright RM, Vaitaitis GM, Wilson CM, Repine TB, Terada LS, Repine JE (1993) cDNA cloning, characterization, and tissue-specific expression of human xanthine dehydrogenase/xanthine oxidase. Proc Natl Acad Sci USA 90(22):10690–10694. https://doi.org/10.1073/pnas.90.22.10690
Kim YS, Nam HJ, Chung HY, Kim ND, Ryu JH, Lee WJ, Yoo MA (2001) Role of xanthine dehydrogenase and aging on the innate immune response of Drosophila. J Am Aging Assoc 24(4):187–193. https://doi.org/10.1007/s11357-001-0020-6
Seymour KJ, Roberts LE, Fini MA, Parmley LA, Oustitch TL, Wright RM (2006) Stress activation of mammary epithelial cell xanthine oxidoreductase is mediated by p38 MAPK and CCAAT/enhancer-binding protein-β. J Biol Chem 281(13):8545–8558. https://doi.org/10.1074/jbc.m507349200
Zhang Y, Wang W, Li M, Li S, Liu S (2017) Identification of putative carboxylesterase and aldehyde oxidase genes from the antennae of the rice leaffolder, Cnaphalocrocis medinalis (Lepidoptera: Pyralidae). J Asia Pac Entomol 20(3):907–913. https://doi.org/10.1016/j.aspen.2017.06.001
Pongsomboon S, Wongpanya R, Tang S, Chalorsrikul A, Tassanakajon A (2008) Abundantly expressed transcripts in the lymphoid organ of the black tiger shrimp, Penaeus monodon, and their implication in immune function. Fish Shellfish Immunol 25(5):485–493. https://doi.org/10.1016/j.fsi.2008.07.010
Kamli MR, Kim J, Pokharel S, Jan AT, Lee EJ, Choi I (2014) Expressional studies of the aldehyde oxidase (AOX1) gene during myogenic differentiation in C2C12 cells. Biochem Biophys Res Commun 450(4):1291–1296. https://doi.org/10.1016/j.bbrc.2014.06.126
Kundu TK, Hille R, Velayutham M, Zweier JL (2007) Characterization of superoxide production from aldehyde oxidase: an important source of oxidants in biological tissues. Arch Biochem Biophys 460(1):113–121. https://doi.org/10.1016/j.abb.2006.12.032
Yergaliyev TM, Nurbekova Z, Mukiyanova G, Akbassova A, Sutula M, Zhangazin S, Omarov RT (2016) The involvement of ROS producing aldehyde oxidase in plant response to Tombusvirus infection. Plant Physiol Biochem 109:36–44. https://doi.org/10.1016/j.plaphy.2016.09.001
Inada M, Kihara K, Kono T, Sudhakaran R, Mekata T, Sakai M, Itami T (2013) Deciphering of the dual oxidase (Nox family) gene from kuruma shrimp, Marsupenaeus japonicus: full-length cDNA cloning and characterization. Fish Shellfish Immunol 34(2):471–485. https://doi.org/10.1016/j.fsi.2012.11.026
Blokhina O (2003) Antioxidants, oxidative damage and oxygen deprivation stress: a review. Ann Bot 91(2):179–194. https://doi.org/10.1093/aob/mcf118
Arbi M, Pouliliou S, Lampropoulou M, Marmaras VJ, Tsakas S (2011) Hydrogen peroxide is produced by E. coli challenged haemocytes and regulates phagocytosis, in the medfly Ceratitis capitata. The active role of superoxide dismutase. Dev Comp Immunol 35(8):865–871. https://doi.org/10.1016/j.dci.2011.03.020
Acknowledgements
This work was supported by JSPS-KAKENHI a Grant-in-Aid for Scientific Research (C) 15K07555 and Grant- in-Aid for Young Scientists (B) 16K18747. We would like to thank Editage (http://www.editage.jp) for English language editing.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
No potential conflict of interest are disclosed.
Research involving human participants and/or animals
The kuruma shrimp that our target species in this study is edible shrimp in Japan. So, this study is not applicable study in this item. However, before dissecting, the kuruma shrimp was immersed into the iced water for around 5 mm into fall into suspended animation.
Informed consent
We agree that the information in this study is disclosed.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Okamura, Y., Inada, M., Elshopakey, G.E. et al. Characterization of xanthine dehydrogenase and aldehyde oxidase of Marsupenaeus japonicus and their response to microbial pathogen. Mol Biol Rep 45, 419–432 (2018). https://doi.org/10.1007/s11033-018-4177-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-018-4177-9