Abstract
Bladder cancer (BC) is a fatal malignancy with considerable mortality. BC urinary metabolomics has been extensively investigated for biomarker discovery, but few BC blood metabolomic studies have been performed. Hence, a plasma pseudotargeted metabolomic method based on gas chromatography–mass spectrometry with selected ion monitoring (GC-MS-SIM) was developed to study metabolic alterations in BC. The analytical performance of the developed method was compared with that of a nontargeted method. The relative standard deviation (RSD) values of 89 and 70.7 % of the peaks obtained using the pseudotargeted and nontargeted methods, respectively, were less than 20 %. The Pearson correlations of 90.7 and 78.3 % of the peaks obtained using the pseudotargeted and nontargeted methods, respectively, exceeded 0.90 in the linearity evaluation. Compared with the nontargeted method, the signal-to-noise ratios (S/N) of 97.9 and 69.3 % of the peaks increased two- and fivefold, respectively. The developed method was fully validated, with good precision, recovery, and stability of the trimethylsilyl (TMS) derivatives. The method was applied to investigate BC. Significant increases in the contents of metabolites involved in, for example, the pentose phosphate pathway (PPP) and nucleotide and fatty acid synthesis were found in the high-grade (HG) BC group compared to the healthy control (HC) group. These differences imply that the activated PPP may regulate BC cell proliferation by promoting lipid and nucleotide biosynthesis and the detoxification of reactive oxygen species (ROS). These results illustrate that the plasma pseudotargeted method is a powerful tool for metabolic profiling.

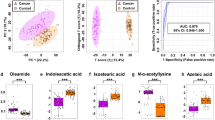
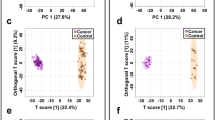
The plasma pseudotargeted metabolic profiling suggested the metabolic alterations in bladder cancer (BC) and the significantly differential metabolites for BC discrimination





Similar content being viewed by others
References
Ferlay J, Soerjomataram I, Dikshit R, Eser S, Mathers C, Rebelo M, et al. Cancer incidence and mortality worldwide: sources, methods and major patterns in GLOBOCAN 2012. Int J Cancer. 2015;136(5):E359–86. doi:10.1002/ijc.29210.
Burger M, Catto JW, Dalbagni G, Grossman HB, Herr H, Karakiewicz P, et al. Epidemiology and risk factors of urothelial bladder cancer. Eur Urol. 2013;63(2):234–41. doi:10.1016/j.eururo.2012.07.033.
Bansal N, Gupta A, Mitash N, Shakya PS, Mandhani A, Mahdi AA, et al. Low- and high-grade bladder cancer determination via human serum-based metabolomics approach. J Proteome Res. 2013;12(12):5839–50. doi:10.1021/pr400859w.
Conde VR, Oliveira PF, Nunes AR, Rocha CS, Ramalhosa E, Pereira JA, et al. The progression from a lower to a higher invasive stage of bladder cancer is associated with severe alterations in glucose and pyruvate metabolism. Exp Cell Res. 2015;335(1):91–8. doi:10.1016/j.yexcr.2015.04.007.
Chan EC, Pasikanti KK, Hong Y, Ho PC, Mahendran R, Raman Nee Mani L, et al. Metabonomic profiling of bladder cancer. J Proteome Res. 2015;14(2):587–602. doi:10.1021/pr500966h.
Hurle R, Losa A, Manzetti A, Lembo A. Upper urinary tract tumors developing after treatment of superficial bladder cancer: 7-year follow-up of 591 consecutive patients. Urology. 1999;53(6):1144–8.
Fiehn O. Metabolomics—the link between genotypes and phenotypes. Plant Mol Biol. 2002;48:155–71.
Cao M, Zhao L, Chen H, Xue W, Lin D. NMR-based metabolomic analysis of human bladder cancer. Anal Sci. 2012;28(5):451–6.
Huang Z, Lin L, Gao Y, Chen Y, Yan X, Xing J et al. Bladder cancer determination via two urinary metabolites: a biomarker pattern approach. Mol Cell Proteomics. 2011;10(10):M111 007922. doi:10.1074/mcp.M111.007922.
Huang Z, Chen Y, Hang W, Gao Y, Lin L, Li DY, et al. Holistic metabonomic profiling of urine affords potential early diagnosis for bladder and kidney cancers. Metabolomics. 2012;9(1):119–29. doi:10.1007/s11306-012-0433-5.
Alberice JV, Amaral AF, Armitage EG, Lorente JA, Algaba F, Carrilho E, et al. Searching for urine biomarkers of bladder cancer recurrence using a liquid chromatography-mass spectrometry and capillary electrophoresis-mass spectrometry metabolomics approach. J Chromatogr A. 2013;1318:163–70. doi:10.1016/j.chroma.2013.10.002.
Jin X, Yun SJ, Jeong P, Kim IY, Kim W-J, Park S. Diagnosis of bladder cancer and prediction of survival by urinary metabolomics. Oncotarget. 2014;5(6):1635–45.
Peng J, Chen YT, Chen CL, Li L. Development of a universal metabolome-standard method for long-term LC-MS metabolome profiling and its application for bladder cancer urine-metabolite-biomarker discovery. Anal Chem. 2014;86(13):6540–7. doi:10.1021/ac5011684.
Putluri N, Shojaie A, Vasu VT, Vareed SK, Nalluri S, Putluri V, et al. Metabolomic profiling reveals potential markers and bioprocesses altered in bladder cancer progression. Cancer Res. 2011;71(24):7376–86. doi:10.1158/0008-5472.CAN-11-1154.
Walsh MC, Brennan L, Malthouse JPG, Roche HM, Gibney MJ. Effect of acute dietary standardization on the urinary, plasma, and salivary metabolomic profiles of healthy humans. Am J Clin Nutr. 2006;84:531–9.
Lin L, Huang Z, Gao Y, Chen Y, Hang W, Xing J, et al. LC-MS-based serum metabolic profiling for genitourinary cancer classification and cancer type-specific biomarker discovery. Proteomics. 2012;12(14):2238–46. doi:10.1002/pmic.201200016.
Pasikanti KK, Ho PC, Chan EC. Gas chromatography/mass spectrometry in metabolic profiling of biological fluids. J Chromatogr B, Anal Technol Biomed Life Sci. 2008;871(2):202–11. doi:10.1016/j.jchromb.2008.04.033.
Zeigler C, Macnamara K, Wang Z, Robbat Jr A. Total alkylated polycyclic aromatic hydrocarbon characterization and quantitative comparison of selected ion monitoring versus full scan gas chromatography/mass spectrometry based on spectral deconvolution. J Chromatogr A. 2008;1205(1-2):109–16. doi:10.1016/j.chroma.2008.07.086.
Robinson MD, De Souza DP, Keen WW, Saunders EC, McConville MJ, Speed TP, et al. A dynamic programming approach for the alignment of signal peaks in multiple gas chromatography-mass spectrometry experiments. BMC Bioinformatics. 2007;8:419. doi:10.1186/1471-2105-8-419.
Li Y, Ruan Q, Ye G, Lu X, Lin X, Xu G. A novel approach to transforming a non-targeted metabolic profiling method to a pseudo-targeted method using the retention time locking gas chromatography/mass spectrometry-selected ions monitoring. J Chromatogr A. 2012;1255:228–36. doi:10.1016/j.chroma.2012.01.076.
Ye G, Liu Y, Yin P, Zeng Z, Huang Q, Kong H, et al. Study of induction chemotherapy efficacy in oral squamous cell carcinoma using pseudotargeted metabolomics. J Proteome Res. 2014;13(4):1994–2004. doi:10.1021/pr4011298.
Zhao Y, Zhao C, Lu X, Zhou H, Li Y, Zhou J, et al. Investigation of the relationship between the metabolic profile of tobacco leaves in different planting regions and climate factors using a pseudotargeted method based on gas chromatography/mass spectrometry. J Proteome Res. 2013;12(11):5072–83. doi:10.1021/pr400799a.
Zhao Y, Zhang L, Zhao C, Hu C, Li Y, Zhao J, et al. Metabolic responses of rice leaves and seeds under transgenic backcross breeding and pesticide stress by pseudotargeted metabolomics. Metabolomics. 2015;11(6):1802–14. doi:10.1007/s11306-015-0834-3.
Zhao Y, Zhao J, Zhao C, Zhou H, Li Y, Zhang J, et al. A metabolomics study delineating geographical location-associated primary metabolic changes in the leaves of growing tobacco plants by GC-MS and CE-MS. Sci Rep. 2015;5:16346. doi:10.1038/srep16346.
Zhao J, Zhao Y, Hu C, Zhao C, Zhang J, Li L, et al. Metabolic profiling with gas chromatography-mass spectrometry and capillary electrophoresis-mass spectrometry reveals the carbon-nitrogen status of tobacco leaves across different planting areas. J Proteome Res. 2016;15(2):468–76. doi:10.1021/acs.jproteome.5b00807.
Storey JD. A direct approach to false discovery rates. J R Stat Soc B. 2002;64:479–98.
Currie E, Schulze A, Zechner R, Walther Tobias C, Farese RV. Cellular fatty acid metabolism and cancer. Cell Metab. 2013;18(2):153–61. doi:10.1016/j.cmet.2013.05.017.
Sugino T, Baba K, Hoshi N, Aikawa K, Yamaguchi O, Suzuki T. Overexpression of fatty acid synthase in human urinary bladder cancer and combined expression of the synthase and Ki-67 as a predictor of prognosis of cancer patients. Med Mol Morphol. 2011;44(3):146–50. doi:10.1007/s00795-010-0517-0.
Du W, Jiang P, Mancuso A, Stonestrom A, Brewer MD, Minn AJ, et al. TAp73 enhances the pentose phosphate pathway and supports cell proliferation. Nat Cell Biol. 2013;15(8):991–1000. doi:10.1038/ncb2789.
Jiang P, Du W, Wu M. Regulation of the pentose phosphate pathway in cancer. Protein Cell. 2014;5(8):592–602. doi:10.1007/s13238-014-0082-8.
Liu PF, Cao YW, Jiang HP, Wang YH, Yang XC, Wang XS, et al. Heterogeneity research in muscle-invasive bladder cancer based on differential protein expression analysis. Med Oncol. 2014;31:21. doi:10.1007/s12032-014-0021-9.
Langbein S, Zerilli M, zur Hausen A, Staiger W, Rensch-Boschert K, Lukan N, et al. Expression of transketolase TKTL1 predicts colon and urothelial cancer patient survival: Warburg effect reinterpreted. Br J Cancer. 2006;94(4):578–85. doi:10.1038/sj.bjc.6602962.
Semilia M, Hennenlotter J, Pavone C, Bischoff T, Kühs U, Gakis G, et al. Expression patterns and prognostic role of transketolase-like 1 in muscle-invasive bladder cancer. World J Urol. 2015;33(10):1403–9. doi:10.1007/s00345-014-1473-4.
Li HD, Xu QS, Liang YZ. Random frog: an efficient reversible jump Markov Chain Monte Carlo-like approach for variable selection with applications to gene selection and disease classification. Anal Chim Acta. 2012;740:20–6. doi:10.1016/j.aca.2012.06.031.
Picard RR, Cook RD. Cross-validation of regression models. J Am Stat Assoc. 1984;79(387):575–83. doi:10.1080/01621459.1984.10478083.
Xu Q-S, Liang Y-Z. Monte Carlo cross validation. Chemom Intell Lab. 2001;56:1–11.
Acknowledgments
The study has been supported by the foundations (No. 21435006, No. 21375127) and the creative research group project (No. 21321064) from the National Natural Science Foundation of China.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
This research was approved by the ethics committee of the Shanghai Changhai Hospital, China. All the participants signed informed consent forms.
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Yang Zhou and Ruixiang Song contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 773 kb)
Rights and permissions
About this article
Cite this article
Zhou, Y., Song, R., Zhang, Z. et al. The development of plasma pseudotargeted GC-MS metabolic profiling and its application in bladder cancer. Anal Bioanal Chem 408, 6741–6749 (2016). https://doi.org/10.1007/s00216-016-9797-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-016-9797-0