Abstract
Studies on hematopoietic stem cells (HSCs) and leukemia stem cells (LSCs) have helped to establish the paradigms of normal and cancer stem cell concepts. For both HSCs and LSCs, specific gene expression programs endowed by their epigenome functionally distinguish them from their differentiated progenies. MicroRNAs (miRNAs), as a class of small non-coding RNAs, act to control post-transcriptional gene expression. Research in the past decade has yielded exciting findings elucidating the roles of miRNAs in control of multiple facets of HSC and LSC biology. Here, we review recent progresses on the functions of miRNAs in HSC emergence during development, HSC switch from a fetal/neonatal program to an adult program, HSC self-renewal and quiescence, HSC aging, HSC niche, and malignant stem cells. While multiple different miRNAs regulate a diverse array of targets, two common themes emerge in HSC and LSC biology: miRNA-mediated regulation of epigenetic machinery and cell signaling pathways. In addition, we propose that miRNAs themselves behave like epigenetic regulators, as they possess key biochemical and biological properties that can provide both stability and alterability to the epigenetic program. Overall, the studies of miRNAs in stem cells in the hematologic contexts not only provide key understandings to post-transcriptional gene regulation mechanisms in HSCs and LSCs but also will lend key insights for other stem cell fields.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The field of hematology has initiated the concepts of both tissue stem cells and cancer stem cells [1]. Hematopoietic stem cells (HSCs) have been historically defined as cells that can provide long-term repopulation capacity for the hematopoietic system when transplanted into a new host. Although recent studies in native hematopoiesis have modified views on the role of stem cell activity in an unperturbed healthy mouse [2, 3], the traditional HSC definition is intrinsically related to the success of the life-saving medical procedure, bone marrow transplantation, and will be used in this review.
HSCs are capable of maintaining the hematopoietic system for the lifetime duration of an organism. HSCs are rare hematopoietic cells sitting at the apex of the hematopoietic hierarchy and possess extensive ability to self-renew as well as the capacity to gradually differentiate into downstream hematopoietic lineages. During fetal development, HSCs emerge as part of the definitive hematopoiesis program, best characterized through budding from special populations of endothelial cells [1, 4, 5]. Newly emerged HSCs migrate to and colonize the fetal liver, a stage that is accompanied by HSCs undergoing active cell cycle [4]. In adults, however, HSCs are predominantly localized within bone marrow and are maintained by a complex set of microenvironment cues provided by the HSC niche [6]. Furthermore, under normal conditions, most HSCs exist in a quiescent state without active cycling [7, 8]. Why must HSCs remain largely quiescent in adults? Although the answer is not fully clear, it has been proposed that stem cell cycling is associated with cancer incidence [9], and quiescence is thus one possible mechanism to avoid the accumulation of DNA replication errors and consequent leukemic transformation. Similar to normal hematopoiesis, leukemia cells are hierarchically organized [10, 11], with relatively rare leukemia stem cells (LSCs) being the only cell population that can re-initiate leukemia in a new host. The bulk of leukemia cells does not have strong self-renewal capacity and is derived from LSCs.
The stem cell concepts of both HSCs and LSCs underscore the importance of the epigenome in defining the functional cellular heterogeneity in normal and malignant hematopoiesis. HSCs and most of their differentiated progenies share the same genetic content, yet they differ strongly on cellular function. Likewise, the bulk population of leukemia inherits genetic contents from LSCs, and the pivotal difference between LSCs and the rest of the leukemia cells on leukemia initiation has to be attributed to their different epigenome.
MicroRNAs (miRNAs) are small non-coding RNAs of ∼22 nucleotides in length. miRNAs are well known to post-transcriptionally regulate gene expression, by degrading target messenger RNA and/or inhibiting protein translation. miRNAs are transcribed as long primary miRNAs, mostly by RNA polymerase II. Primary miRNAs undergo step-wise processing by Drosha-and-DGCR8-containing microprocessor complex in the nucleus to generate precursor miRNAs, which are then transported into the cytoplasm by exportin 5 and further processed by Dicer and its cofactors to generate mature miRNA duplexes. Loading of mature miRNAs into Argonaute (AGO) proteins, with the assistance of HSP90, allows miRNA-guided targeting of AGO proteins to target mRNA and results in downregulation of their binding targets [12–15]. The roles of miRNA in the hematopoietic system, including stem cells, have been documented by a number of excellent reviews [16–27]. In this review, we summarize recent progresses of miRNAs that functionally control HSC and LSC biology. Two emerging themes from these studies are that epigenetic machineries and cell signaling pathways are frequently regulated by miRNAs. We particularly emphasize miRNAs’ direct roles on regulating epigenetic enzymes, many of which have been found as somatically mutated in healthy individuals and malignant hematopoiesis. We propose that miRNAs themselves should be considered as an important component of the epigenetic program given their properties. As the focus of this review is on stem cells, we will not be able to cover every key and seminal study on miRNAs in other hematologic contexts, many of which have been reviewed elsewhere.
miRNAs Intrinsically Possess Epigenetic Properties and Serve as Guardians of Cell States
The epigenome has two key properties, its maintenance and its amenability to change, both of which are essential for HSCs and LSCs. During symmetric cell division, the two daughter cells perform similar functions as the mother cells and retain a similar gene expression program, requiring the epigenome to be faithfully copied. The achievement of this feat is by no means simple, the mechanism of which is still far from completely understood. For example, during DNA replication in the S phase of the cell cycle, DNA needs to be unwound from core histones. After replication, histones, including their modifications, need to be re-established at the right place with a high level of precision on both copies of DNA. The best-known mechanism that can potentially maintain the epigenetic state during cell division is DNA methylation at 5 position of cytosine (5mC). DNMT1 is known as a maintenance DNA methyltransferase that is capable of recognizing hemi-methylated DNA and copy the methylation mark onto the newly synthesized strand [28, 29]. Other than active demethylation (to be discussed in the next section), the most effective way to remove DNA methylation marks is by cell division, in which DNA methylation maintenance mechanisms are suppressed to allow passive dilution of DNA methylation in daughter cells. However, DNA methylation cannot be the only mechanism in maintaining the epigenome stability. In model eukaryotes, such as yeast, worms, and fruit flies, DNA methylation is not an actively utilized mechanism, yet their epigenome can be faithfully maintained. In mouse embryonic stem cells, the pluripotency epigenome can be maintained in the absence of all DNMTs (DNMT1, DNMT3a, and DNMT3b) [30, 31]. These facts argue for the existence of additional epigenome maintenance programs that share similar properties as DNA methylation.
miRNAs possess some key properties that are reminiscent to DNA methylation and can be considered an underappreciated component of the epigenetic program. One of the striking features of miRNAs is their stability. Studies from multiple laboratories, including ours, have found that the vast majority of measurable mature miRNAs have half-lives much longer than those of messenger RNAs [32•, 33, 34], possibly due to the protection of mature miRNAs from cytosolic RNAses by AGO proteins. The half-lives of many miRNAs are too long to be determinable in cell culture experiments [32•], but a study on miR-208 in mouse heart tissue shows that the half-life of this miRNA is >12 days [35]. The stability of miRNAs suggests that one of the most effective ways to decrease miRNA expression in cells is by passive dilution, a property that is similar to DNA methylation marks. Consistent with this notion, we have previously observed that the Lin−Kit+Sca− myeloid progenitor cells, some of which cycle at <8 h per cell cycle [36], have decreased global miRNA expression in comparison to the slower cycling HSCs or more mature cells [37]. In addition to stability, miRNA expression is also amenable to changes during cell-state transitions. Differential miRNA expression, both upregulation and downregulation, has been observed between HSCs, LSCs, and their differentiated progenies [37–41]. Transcriptional upregulation of miRNAs can lead to increased mature miRNA expression. The mechanisms underlying downregulation of miRNA during differentiation are much less understood, which may involve both cell division-based passive dilution in the presence of transcriptional silencing or active destabilization of miRNA. For the aforementioned downregulation of miRNAs in fast cycling myeloid progenitors in comparison to HSCs or Lin−Kit+Sca+ hematopoietic stem and progenitor cell (HSPC) population [36, 37], it is currently unknown how many cell divisions are required to go from HSCs or HSPCs to myeloid progenitors, the elucidation of which may reveal the relative contribution of passive dilution to miRNA downregulation in this process. Active miRNA degradation, either through specific enzymes, downregulation of the protective AGO proteins, or secretion through exosomes have been reported in other cellular systems to contribute to active mature miRNA destabilization [33, 34]. However, mechanisms of miRNA downregulation have not been extensively studied within the hematopoietic system, the elucidation of which will add key insights into miRNAs’ function in stem cells.
Qualifying miRNAs as an epigenetic component requires additional evidence that they confer functional stability of cell states, in addition to their biochemical properties of being both stable and alterable. Indeed, evidence accumulated in the past decade has identified not only specific miRNAs in control of cell fates but also the overall miRNA pathway as a guardian of cell states. One of the best examples for the latter is the overall miRNA regulation pathway serving as a tumor suppressor for safe-guarding normal cell states from cancer transformation, which we have recently reviewed in detail [42]. Briefly, global decrease of miRNA regulation, by global downregulation of miRNA expression, mutations in miRNA biogenesis pathway, inhibiting miRNA biogenesis through oncogene signaling, or evasion of miRNA regulation by shortening three prime untranslated region (3′UTR), can positively impact tumorigenesis in both mouse and human.
In view of miRNAs as an epigenetic component that helps to define cellular states, we review below both studies that link miRNA directly with the regulation of epigenetic machinery in the context of HSCs and LSCs and miRNA-mediated regulation of HSCs and LSCs through other direct mechanisms, including regulating cell signaling. Table 1 summarizes the function of specific miRNAs in HSC and LSC regulation.
miRNA-Mediated Control of Epigenetic Program in HSCs and LSCs
A complex epigenetic program safeguards HSCs by maintaining the self-renewal potential and limiting overproliferation, the disruption of which leads to HSC exhaustion. Studies in the past several years have revealed striking levels of somatic mutations in the form of clonal hematopoiesis, particularly in healthy elderly individuals [96–100]. Among the most frequently mutated genes in clonal hematopoiesis are DNMT3A and TET2 [101–103], both of which are recognized as the founding genetic mutations in hematopoietic malignancies such as acute myeloid leukemia [100, 104, 105]. In healthy populations aged 65 or older, ∼3 % contains somatic DNMT3A mutations and ∼0.5–1 % with TET2 mutations, with mutant alleles detectable in ∼20 % or more of mononuclear hematopoietic cells in peripheral blood. Both DNMT3A and TET2 are involved in the DNA methylation pathway. DNMT3A is a “de novo” methyltransferase that is not dependent on existing methylation marks. In contrast, TET2 is a member of the ten-eleven translocation (TET) family that catalyzes the oxidation of 5mC into 5-hydroxymethylcytosine (5hmC). TET2 proteins can further oxidize 5hmC into 5 formylcytosine (5fC) and then 5-carboxyl cytosine (5caC) that can be subjected to excision repair-mediated active demethylation [106–108]. In addition, due to DNMT1 not recognizing 5hmC, oxidation of 5mC into 5hmC effectively leads to site-specific suppression of maintenance methylation activity and hence proliferation-based passive demethylation [109]. Functionally, despite opposite biochemical activities, genetic ablation of DNMT3A and TET2 can both lead to HSC amplification that outcompete normal HSCs [110–115]. In particular, TET2 demonstrates haplo-insufficiency both in human malignancy samples and in mouse models, which is interesting given that miRNAs often only partially suppress target gene expression. In this section, we will review the roles of miRNAs that target epigenetic machinery, with a focus on discussing their regulation of DNA methylation/demethylation enzymes.
miRNAs Regulating TET2 and Other DNA Demethylation Genes
TET2 has been demonstrated to be under extensive regulation by miRNAs. The miR-125 family miRNAs are among these TET2-targeting miRNAs. miR-125 family consists of miR-125a and miR-125b, with the latter encoded by two separate genomic loci on human chromosomes 11 and 21. Members of the miR-125 family appear to play redundant roles in mouse hematopoiesis [116]. Both miR-125a and miR-125b show enriched expression in HSCs, and their expression decreases in more differentiated cell populations [37, 117, 118]. Interestingly, miR-125b is involved in rare cases of genomic rearrangements in both myeloid and lymphoid leukemia, which leads to increased miR-125b expression [48, 119]. Furthermore, increased expression of miR-125b from the miR-125b-2 locus has been observed in trisomy 21 cases, which demonstrate increased susceptibility toward acute megakaryocytic leukemia [49]. Even in the absence of direct genomic rearrangements involving miR-125 loci, overexpression of miR-125 family miRNAs has been observed frequently in acute myeloid leukemia and other hematopoietic malignancies and occasionally under the control of other oncogenes such as ETV6-RUNX1 fusion [120]. The functions of miR-125 family miRNA in the hematopoietic system have been reviewed previously [24]. In short, constitutive overexpression of either miR-125a or miR-125b leads to HSC expansion in vivo, and skewed myeloid and monocyte differentiation, with the latter phenotype dependent on continued miR-125 overexpression [50]. Among many targets of miR-125 family miRNAs that have been studied [24, 44••], miR-125-TET2-targeting relationships have been identified through a systematical screen for TET2-targeting miRNAs, and miR-125 is capable of altering the global 5hmC levels in hematopoietic cells [44••]. TET2 knockout displays similar phenotypes as miR-125 overexpression, and enforced expression of TET2 corrects nearly all miR-125 overexpression phenotypes including hematopoietic expansion and lineage differentiation skewing, supporting TET2 as an important target of miR-125 family miRNAs [44••]. Interestingly, other than TET2, other targets involved in attenuation of cytokine signaling have been reported [24, 50], placing this family of miRNAs as regulators of both epigenetic and signaling pathways. Germline knockout of miR-125a has recently been published. Curiously, heterozygous miR-125a knockout mice develop myeloproliferation similar to miR-125a overexpression, but this phenotype cannot be observed in homozygous miR-125a knockout mice [51]. Whether this is due to miR-125a function in the microenvironment, rather than within hematopoietic cells, remains to be determined.
Another miRNA family that target TET2 is the miR-29 family [44••, 121], which consists of miR-29a, miR-29b, and miR-29c. Unlike miR-125, miR-29 targets all three TET family members and regulates DNA methyltransferases and other components within DNA demethylation pathway. While the function of miR-29 family will be discussed in detail in the next section, overexpression of TET2 can also correct miR-29b-induced myeloproliferation [44••] supporting a role of TET2 as a downstream effector of miR-29 family miRNAs.
miR-22 overexpression, which is often seen in myelodysplastic syndrome (MDS), can also lead to direct targeting of TET2. Furthermore, in a genetic mouse model, overexpression of miR-22 leads to increased HSC activity and MDS-like phenotypes, whereas overexpression of TET2 suppressed miR-22 phenotypes in vitro and in vivo, including a partial rescue of mouse survival [122]. In contrast to these findings, miR-22 has recently been described as a tumor suppressor in acute myeloid leukemia (AML), where it negatively regulates the expression of a number of oncogenes including CRTC1, FLT3, and MYCBP. In this study, loss of miR-22 could occur either via genetic lesion or TET1-mediated silencing [123] suggesting that TET family of proteins may be negatively regulating miR-22. In addition to the abovementioned miRNAs, miR-26 family has been shown to downregulate multiple TET family genes, including TET2, in both hematological and pancreatic contexts [124], and overexpression of miR-26a in wild-type bone marrow cells leads to a transient myeloid expansion in vivo [44••]. Similarly, overexpression of miR-101, which can target both TET2 and EZH2, leads to myeloid skewing in differentiation [44••, 125]. Other TET2-targeting miRNAs have been identified through the 3′UTR screen of ∼460 mouse and human miRNAs, some of which selectively target human TET2 over mouse TET2 (for example, miR-7 had a much weaker effect on mouse TET2), indicating a species-specific mode of regulation. In a cohort of cytogenetically normal AML, overexpression of miR-125b, miR-29b, miR-29c, miR-101, and miR-7 is significantly associated with TET2 wild-type cases than TET2 mutant cases, suggesting that these miRNAs may also underlie human leukemia formation [44••]. Why is TET2 under the regulation of so many miRNAs? Although the answer is unclear, it is possible that there might be a pressure to attenuate TET2 expression in multiple tissue types during development so that overdemethylation of the epigenome does not occur. In addition to TET family-mediated demethylation, activation-induced cytidine deaminase (AID) and APOBEC can also lead to active DNA demethylation, although their roles may be more context-specific. Interestingly, AID can be regulated by miR-155 [126] and miR-181b [127] in the context of B cells, and both of these miRNAs are elevated in MDS and leukemia CD34+ cells [69, 70, 79]. Whether these targeting relationships regulate HSCs needs further investigation.
miRNAs Regulating DNA Methylation
In addition to targeting TET family genes, the miR-29 family miRNAs have been demonstrated to regulate multiple DNA methyltransferase genes. miR-29b was initially identified as a potential tumor suppressor in established AML cells. When ectopic miR-29b was introduced into established human AML cell lines or patient blasts, induction of apoptosis and partial differentiation were observed [128, 129]. One of the studies found that miR-29b can target DNMT3A and DNMT3B directly and target DNMT1 indirectly through suppressing Sp1 [128] and miR-29b overexpression altered DNA methylation landscape in AML cells with re-expression of tumor suppressor genes. In contrast to these reports of miR-29b as a tumor suppressor in AML, miR-29a overexpression in wild-type hematopoietic cells leads to myeloproliferation and occasional AML progression, accompanied by aberrant self-renewal activity [90]. Recently, the effect of ablating miR-29a/miR-29b-1 cluster has been examined in the hematopoietic context. Their deletion leads reduced HSCs with reduced self-renewal activity. Reintroduction with miR-29a, but less so with miR-29b, leads to rescue effects, suggesting that miR-29a deficiency underlies this phenotype. Crossing miR-29a/miR-29b-1 heterozygous knockout mice with DNMT3A heterozygous knockout mice convincingly establishes that DNMT3A is an important target of miR-29 in this context [91•]. Notably, in miR-29a/miR-29b-1 knockout HSCs, DNMT3A is only upregulated mildly, at 1.55-fold the level seen in wild-type HSCs, suggesting a quantitative sensitivity of HSCs to small changes in DNMT3A levels. Furthermore, DNMT3A knockout cannot fully rescue the HSC deficiencies in miR-29 knockout, suggesting that other targets, such as TET2, may be involved. In addition to miR-29a, overexpression of miR-29b in wild-type bone marrow cells leads to myeloid expansion [44••]. Why do some studies found miR-29 family miRNA to be potential tumor suppressors and others assign oncogenic functions? One possibility is the cellular context differences, in which oncogenic functions are seen when miR-29 is examined in wild-type HSCs, whereas tumor suppressor functions are seen in established AML cells. The second possibility is that other targets of miR-29, such as TET2, MCL-1, or other targets [44••, 129], may alter biological output. It is thus interesting to note that some subtypes of AML are dependent on the oncogenic function of TET1 [130], which can also be targeted by miR-29 family miRNAs.
In addition to miR-29 family miRNAs, miR-199a has been demonstrated to target DNMT3A in other tissue contexts [131]. When overexpressed in the hematopoietic system, miR-199a leads to an expansion of myeloid progenitors [58]. Whether DNMT3A is functionally involved in the miR-199a effect is unknown. DNMT1 is also regulated by other miRNAs. miR-152, which potentially regulates DNMT1, is often heavily methylated in leukemia [68]. miR-21 [83] and miR-126 [54••] both of which play roles in leukemia have been shown to regulate DNMT1 in lymphoid cells [132, 133].
miRNAs Regulating Cell Signaling and Other Targets in HSC and LSC Biology
miRNA Regulation of Hematopoietic Development
In mammals, hematopoiesis is initially observed in the developing yolk sac, termed “primitive hematopoiesis.” This early wave can produce red blood cells and populations of macrophages. HSCs emerge from the subsequent waves of definitive hematopoiesis, with HSC activity detected in the aorta-gonad mesonephros (AGM) region by budding off the hemogenic endothelium. These cells then migrate to the fetal liver and finally the bone marrow where they predominantly reside in the adult organism. A proliferation switch occurs in mouse roughly 2–3 weeks post-birth, with HSCs changing from actively cycling in the fetal and neonatal stage to largely quiescent in the adult stage [134].
miR-126 has been reported to regulate the primitive wave of erythroid differentiation through modeling embryoid body differentiation from murine embryonic stem cells [135•]. Gain of miR-126 expression increases the number of erythroid progenitors, whereas knocking out miR-126 reduces these progenitors. The mechanism is through non-cell-autonomous regulation of Vcam-1 in mesenchymal cell that, in turn, impacted erythroid progenitor development.
miRNAs also play important roles in the development of HSCs during definitive hematopoiesis. One important miRNA in this process is miR-142-3p. miR-142 is abundantly expressed and enriched in adult hematopoietic cells [80]. The precursor miR-142 gives rise to two mature miRNAs, miR-142-5p and miR-142-3p, from the 5′ and 3′ arm of the miRNA hairpin, respectively. Unlike many other miRNAs, both miR-142-5p and miR-142-3p are abundantly expressed. In xenopus, miR-142-3p is strongly upregulated in newly emerged HSPCs from hemogenic endothelium and is functionally required for the development of the hemangioblast through targeting TGF-beta receptor and its signaling [60•]. In zebrafish, miR-142-3p is abundantly expressed in RUNX1 and MYB-positive HSCs. Inhibition of miR-142-3p in zebrafish by morpholino leads to decreased HSCs in AGM, a defect that can be corrected by inhibiting Irf7 [61], suggesting that interferon signaling might be involved. Similarly in mouse, inhibition of miR-142-3p in AGM cells leads to reduced colony formation in vitro and reduced spleen colonies in vivo [61].
In fetal liver HSCs and adult HSCs, despite their cell cycle differences, a shared expression program regulates the expression of the maternally expressed imprinted Dlk1-Gtl2 locus [136]. This non-coding region harbors a large number of miRNAs as well as other non-coding transcripts. Inactivation of the expression of this region leads to defective fetal HSC maintenance, resulting in reduced long-term HSC function. Mechanistically, the miRNAs expressed within this region collectively target the phosphoinositide 3-kinase (PI3K)-mechanistic target of rapamycin (mTOR) signaling pathway. In the absence of this non-coding mechanism, overactivation of mitochondria metabolism occurs with excessive reactive oxygen species.
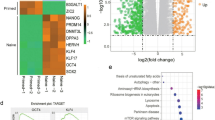
The post-natal switch between the fetal/neonatal hematopoiesis and the adult hematopoiesis program is controlled by a Lin28b-let-7-HMGA2 axis. let-7 is a family of miRNAs initially identified in Caenorhabditis elegans that controls the timing of larval development [137]. let-7 is highly conserved during evolution, and its processing from primary miRNA to precursor miRNA, and from precursor miRNA to mature miRNA, is inhibited by the RNA-binding protein Lin28A and Lin28B [138–140]. In most tissues, let-7 is a tumor suppressor miRNA that downregulates a plethora of oncogene targets, including HMGA2 [141], Ras [142], and Myc [143]. Both Lin28B and HMGA2 are expressed higher in fetal HSCs versus adult HSCs, whereas a few members of let-7 family miRNAs show opposite expression. Overexpression of Lin28B and HMGA2 leads to increased self-renewal of adult HSCs, whereas HMGA2 KO inhibits the heightened self-renewal of wild-type fetal HSCs [43•]. Consistent with this role, ectopic expression of Lin28 in adult HSPCs causes the adaptation of a fetal lymphopoiesis-like program [144], whereas overexpressing let-7 can drive fetal pro-B cells to become more adult-like, through repression of Arid3a [145].
miRNA Regulation of Adult HSC Self-Renewal and Quiescence
Balance of HSC self-renewal, quiescence, and differentiation is tightly controlled in order to maintain a relatively constant HSC pool size over the lifespan of an organism [146]. HSCs are largely quiescent, and the cell cycle status of the HSC reflects their functional output [7, 147, 148]. A number of miRNAs function to positively or negatively regulate HSC self-renewal and/or quiescence. In addition to the aforementioned miR-125 and miR-29 family miRNAs, miR-196b has enriched expression in HSPCs, although expression of this miRNA is higher in short-term HSCs than long-term HSCs. miR-196b is located within the HoxA cluster, and its expression is controlled by mixed-lineage leukemia (MLL). Overexpression of miR-196b in vitro leads to increased c-Kit+ myeloid progenitor cell expansion with hypersensitivity to cytokine stimulation, suggesting modulation of cell signaling [81]. However, the impact of miR-196b on stem cell activity and its targets was not characterized. In contrast to the above miRNAs, overexpression of miR-126 inhibited the self-renewal of HSCs in both human and mouse models, whereas lentiviral sequestering of miR-126 leads to increased HSC cycling and expansion without evidence of exhaustion. This miRNA suppresses multiple members of the PI3K-AKT signaling pathway, and a PI3K inhibitor can reduce cell proliferation upon miR-126 inhibition in vitro [55••]. miR-155 is also capable of expanding HSCs when overexpressed. However, sustained miR-155 overexpression leads to MDS [69] through regulation of PU.1 [71] and SHIP1 [72].
Role of miRNAs in HSC Aging
In mammals, aging leads to an increase in the number of immunophenotypic HSCs but an overall decrease in HSC competitiveness and a skewing toward myeloid differentiation [149]. The miRNA-212/132 cluster shows enriched expression in HSCs and Lin− cells compared to more differentiated progenies. The expression of this cluster further increases during aging [84]. Knocking out this cluster does not strongly impact HSCs under normal conditions, but under conditions of inflammatory stress by LPS, knockout HSCs show reduced cycling [84]. LPS is a gram-negative bacteria endotoxin that can lead to both HSC cycling and a premature aging-like phenotype [150]. Furthermore, miRNA-212/132 knockout modestly improved competitive repopulation of aged HSCs but not young HSCs [84]. Related phenotypes have been observed in miR-146a knockout mice. miR-146a was initially discovered as an LPS-inducible miRNA in macrophages [64] and is thus involved in immune responses [151]. It is also located on chromosome 5q and, together with miR-145, has been demonstrated to induce MDS-like conditions in mice when both miRNAs are inhibited [63]. miR-145 has been reported to cooperate with another 5q gene, RPS14, in regulating erythroid and megakaryocyte differentiation [152]. Knockout of miR-146a does not affect HSCs at 6 weeks, but with increased age, miR-146a knockout mice develop myeloproliferation with a gradual decrease in HSC number and function, leading to HSC exhaustion and neoplasm [153••]. The function of miR-146a can be attributed to miRNA-mediated regulation of inflammatory signaling by targeting TRAF6 and resulting in IL-6 deregulation [153••].
Role of miRNAs in HSC Niche
A complex niche environment regulates HSC and HSPC behavior in vivo. Can miRNAs in niche cells regulate hematopoiesis? The first supporting evidence was revealed in a Dicer knockout model. Mice lacking Dicer1 in their osteoprogenitors, via Sbds expression reduction, develop an MDS-like condition, which progresses to leukemia [154••]. Transplanting wild-type bone marrow cells into mutant host can induce the MDS phenotypes, whereas transplanting MDS bone marrow cells from mutant mice into wild-type recipients corrects the phenotype, supporting a non-hematopoietic role of Dicer1 in modulating HSPC differentiation activity. Although Dicer1 processes additional RNA species other than miRNAs, a HITS-CLIP study performed in murine bone marrow endothelial cell and mesenchymal cells revealed extensive endogenous binding and regulation by miRNAs on potential niche-contributing signaling factors. These include miR-193a regulating JAG1, miR-200a regulating WNT5A, miR-9 regulating MMP2, and miR-185 regulating VEGFA [155]. Other candidate miRNAs that may regulate HSPCs within niche has been reported by a study comparing stromal cell lines that support HSC/HSPC activity in vitro versus those that do not. miRNAs that positively correlate with supportive activity include miR-143, miR-214*, miR-9*, miR-222, miR-342 3p, miR-193*, miR199b*, miR-214*, miR674*, mIR-351, miR-7a-1, miR-148b, miR-455, and miR-199a-3-p, whereas miR-155 and miR-411 are anti-correlated with HSC/HSPC supportive activity [156]. The function of miR-155 in the bone marrow microenvironment has been studied comprehensively in a myeloproliferative neoplasm (MPN) model induced by knocking out Notch-signaling component recombination signal binding protein for immunoglobulin kappa J region (RBPJ). RBPJ knockout leads to an elevation of miR-155 in bone marrow endothelial cells, which, in turn, upregulates pro-inflammatory signaling through repressing kB-Ras1 and hence NF-kB activation, resulting in myeloproliferation in a non-hematopoiesis-autonomous fashion [157•]. It is currently unclear whether the anti-correlation between miR-155 and non-supportive stromal lines is related to this functional activity of miR-155 regulating inflammatory signaling in the bone marrow microenvironment.
Another area with active miRNA research that relates to HSC niche is inter-cellular signaling/communication by secretory exosomes, which contain miRNAs. For example, exosomes from mesenchymal stromal cells contain a number of miRNAs, including miR-451, miR-1202, miR-630, miR1207-5p, miR-33b, miR-1268, miR-638, miR-575, and miR-1225-5p [158]. Whether exosomal miRNAs from bone marrow stromal cells regulate hematopoiesis in vivo is currently unknown. However, a recent study shows trans-cell delivery of cre in vivo [159], suggesting that this mode of epigenetic material exchange between cells may functionally impact hematopoiesis.
Role of miRNAs in LSCs
Comparisons between LSC-enriched populations versus their normal counterparts have revealed a long list of potential miRNAs that are differentially expressed in LSCs versus HSCs and have been discussed elsewhere [20]. Given the inability to isolate either LSCs or HSCs to perfect purity, whether these miRNAs are indeed differentially expressed between functional LSCs and HSCs is unknown. Additional studies sought to stratify phenotypic AML LSCs based on their ability to engraft in an immune compromised mouse. A comparison of miRNA expression and the engraftment ability identified miRNAs both positively (miR-99a, miR-125b, miR-155, miR-409, miR-100, miR-320, miR-126, miR-1, miR-542, and miR-15b) and negatively (miR-451, miR-103, miR-200c, miR-423-3p, let7g, miR-30e, miR-26b, miR-140, miR-22, and miR-21) correlate with AML engraftment and LSC activity [54••]. These candidate miRNAs provide a perfect list for functional validation as only engraftment and leukemia reconstitution is a reliable way of defining LSCs.
Some LSC-enriched miRNAs have been studied and positively influence leukemia development. Remarkably, most of these miRNAs can target enzymes that control DNA methylation. Overexpression of miR-125b-1 [24], miR-125b-2 [49], miR-155 [69], and miR-126 [54••] results in leukemia phenotype, and all of these miRNAs are capable of regulating enzymes which control DNA methylation as discussed previously. The best-studied miRNA in human AML LSCs is miR-126, whose high expression in leukemia specimens correlates with AML survival [54••]. However, opposite to its role in human HSCs discussed earlier, this same miRNA enhances LSC self-renewal and prevents differentiation in vivo. Intriguingly, miR-126 also seems to exert its effect in LSCs through the same PI3K-AKT-mTOR pathway as in HSCs, even though the biological outcome in HSCs and LSCs is different [54••]. These data suggest that the human LSCs behave similarly as fetal liver HSCs [136], in which unchecked PI3K-mTOR pathway damages stem cells, whereas adult HSCs may be opposite.
Among other miRNAs present on the LSC-enriched list, miR-99a and miR-100 are located in close proximity to the miR-125 family members [47] and often co-expressed with miR-125 miRNAs. Interestingly, elevated miR-1 expression is strongly correlated with IDH1/2 mutation status [160], and IDH1/2 mutation negatively regulates TET2 activity. miR-320 [161] and miR-15 [75] have been demonstrated to inhibit cell proliferation, which might be relevant to stem cell quiescence. Further work will be needed to assign functional roles for LSC-enriched miRNAs.
Most miRNAs whose expression is anti-correlated with engraftment have not been studied carefully in the context of HSCs of LSCs. Nevertheless, miR-22 has been discussed above which causes an MDS-like phenotype when overexpressed in wild-type cells and regulates TET2 gene expression.
Summary, Outlook, and Challenges
As evidenced above, miRNAs are functionally involved in most, if not all, aspects of stem cell behaviors in both normal and malignant hematopoiesis. Mechanistically, two common themes emerge from the diverse array of miRNAs and their targets. The downstream pathways of miRNAs that regulate HSC and LSC biology are overall enriched for factors that regulate other epigenetic machineries and/or cell signaling. Some miRNAs regulate both. For example, miR-125 family miRNAs target both TET2, an epigenetic enzyme [44••], and control the strength of cytokine signaling with inhibition of multiple protein phosphatases [50]. miRNA regulation of these two categories of targets makes much sense. miRNAs are known as quantitative regulators of target gene expression, and thus, the functional output of miRNAs must be mediated through factors or combinations of factors that are sensitive to quantitative alterations, rather than black and white binary distinctions. Many of the epigenetic machinery enzymes are intricately controlled in expression. A subtle change in their expression may cause widespread genome-level alterations in epigenetic landscape. For example, TET2 is a well-known haplo-insufficient tumor suppressor [162]. Loss of function of Ezh2, which occurs predominantly on a single allele, is observed in myeloid malignancies [163]. Similarly, signaling pathways are naturally balanced with both activation signaling and deactivation proteins, and quantitative alteration in signaling factors may lead to much amplified functional output than the level of miRNA-mediated target regulation.
In addition to their direct regulation of epigenetic enzymes, we propose that miRNAs should be considered as epigenetic regulators themselves, as they possess key biochemical and biological properties that may provide both stability and alterability to the epigenetic program. This is consistent with a proposed role of miRNAs as buffers for noise in biological systems overall [164].
While exciting progresses have been made, we also highlight three challenging areas that require more intensive research to further elucidate and capitalize on the roles of miRNAs in normal and malignant stem cells. First is on the regulation of miRNAs themselves. Overall, there lacks a clear understanding on what attributes to the specific expression patterns of miRNAs in HSCs and LSCs. The expression of miRNAs can be regulated on the level of primary miRNA transcription, miRNA processing, miRNA loading into AGO proteins, and miRNA stability. A few studies have examined the transcriptional control of miRNAs. For example, miR-146a can be regulated by NF-kB signaling [165], miR-125b can be regulated by HoxA10 [47], and MLL regulates the expression of multiple miRNAs, including miR-196b and miR-150 [81, 94]. However, what genetic elements and transcriptional programs that endow HSC-enriched expression patterns for most of the functional miRNAs are largely unknown. Even less is known about the post-transcriptional control of miRNAs in hematopoiesis. We have shown that human miR-125b-1 behaves differently from other miRNAs in that it has a much-decreased reliance on HSP90 activity when loading into AGO proteins [32•]. As another example, it is interesting to note that a human SNP in the XRN1 gene, which actively regulates miRNA let-7 degradation in C. elegans [166], is associated with mean corpuscular volume of red blood cells in human [167]. These data argue for the importance of examining the post-transcriptional regulation of miRNAs in hematologic systems, which can help reveal mechanisms that underlie stem cell-enriched expression of miRNAs and their downregulation during cell fate changes. Second, for the mechanisms of miRNAs, there is an overreliance on a single miRNA single target scheme, which unfortunately is a common problem across most disciplines of miRNA studies. As each miRNA is known to quantitatively regulate multiple targets, it is conceivable that the function of miRNAs may be contributed by small changes of multiple target genes. Of course, functional integration of quantitative regulation is challenging, and new methodologies are required to push this area forward. The fast development in the CRISPR-based gene editing and gene expression controls may provide a viable approach to control multiple downstream targets of a miRNA of interest. Third, while the functional analysis of miRNAs in HSCs and LSCs have provided mechanistic insight, multiple miRNAs could be interesting candidates for therapeutic modulation. For example, loss of miR-126 and gain of miR-125 family miRNAs both can enhance HSC self-renewal and could potentially be used in clinical amplification of stem cells if they or their inhibitors can be delivered into HSCs. While multiple methods of miRNA or miRNA inhibitor delivery in vitro has been possible [168], small RNA delivery in vivo is overall challenging. Recent studies, however, have showcased a few examples in which nanoparticles have been used to successfully deliver miRNAs or miRNA inhibitors in vivo (e.g., [169]), which may become a route toward successful translation of the basic biological insights on miRNAs.
In summary, exciting progresses in the past decade have provided ample evidence supporting the functions of miRNAs in multiple facets of HSC and LSC biology, including HSC emergence during embryogenesis, HSC switch from a fetal/neonatal program to an adult program, HSC self-renewal and quiescence, HSC aging, HSC niche, and malignant stem cells. The studies of miRNAs in stem cells in the hematologic contexts not only provides key insights into the mechanisms of HSC and LSC controls but also will actively influence the studies of other stem cell areas by establishing key principles.
References
Papers of particular interest, published recently, have been highlighted as: • Of importance; •• Of major importance
Orkin SH, Zon LI. Hematopoiesis: an evolving paradigm for stem cell biology. Cell. 2008;132(4):631–44.
Sun J et al. Clonal dynamics of native haematopoiesis. Nature. 2014;514(7522):322–7.
Busch K et al. Fundamental properties of unperturbed haematopoiesis from stem cells in vivo. Nature. 2015;518(7540):542–6.
Sanchez MJ et al. Characterization of the first definitive hematopoietic stem cells in the AGM and liver of the mouse embryo. Immunity. 1996;5(6):513–25.
Gritz E, Hirschi KK. Specification and function of hemogenic endothelium during embryogenesis. Cell Mol Life Sci. 2016;73(8):1547–67.
Morrison SJ, Scadden DT. The bone marrow niche for haematopoietic stem cells. Nature. 2014;505(7483):327–34.
Foudi A et al. Analysis of histone 2B-GFP retention reveals slowly cycling hematopoietic stem cells. Nat Biotechnol. 2009;27(1):84–90.
Takizawa H et al. Dynamic variation in cycling of hematopoietic stem cells in steady state and inflammation. J Exp Med. 2011;208(2):273–84.
Tomasetti C, Vogelstein B. Cancer etiology. Variation in cancer risk among tissues can be explained by the number of stem cell divisions. Science. 2015;347(6217):78–81.
Bonnet D, Dick JE. Human acute myeloid leukemia is organized as a hierarchy that originates from a primitive hematopoietic cell. Nat Med. 1997;3(7):730–7.
Lapidot T et al. A cell initiating human acute myeloid leukaemia after transplantation into SCID mice. Nature. 1994;367(6464):645–8.
Iwasaki S et al. Hsc70/Hsp90 chaperone machinery mediates ATP-dependent RISC loading of small RNA duplexes. Mol Cell. 2010;39(2):292–9.
Carthew RW, Sontheimer EJ. Origins and mechanisms of miRNAs and siRNAs. Cell. 2009;136(4):642–55.
Bartel DP. MicroRNAs: target recognition and regulatory functions. Cell. 2009;136(2):215–33.
Miyoshi T et al. A direct role for Hsp90 in pre-RISC formation in Drosophila. Nat Struct Mol Biol. 2010;17(8):1024–6.
Wei J et al. Epigenetic alterations of tumor marker microRNAs: towards new cancer therapies. Drug News Perspect. 2010;23(10):655–61.
Palma CA et al. MicroRNA control of myelopoiesis and the differentiation block in acute myeloid leukaemia. J Cell Mol Med. 2012;16(5):978–87.
Laine SK, Hentunen T, Laitala-Leinonen T. Do microRNAs regulate bone marrow stem cell niche physiology? Gene. 2012;497(1):1–9.
Tuna M, Machado AS, Calin GA. Genetic and epigenetic alterations of microRNAs and implications for human cancers and other diseases. Genes Chromosomes Cancer. 2016;55(3):193–214.
Chung SS, Hu W, Park CY. The role of microRNAs in hematopoietic stem cell and leukemic stem cell function. Ther Adv Hematol. 2011;2(5):317–34.
O’Connell RM, Baltimore D. MicroRNAs and hematopoietic cell development. Curr Top Dev Biol. 2012;99:145–74.
Ivey KN, Srivastava D. MicroRNAs as regulators of differentiation and cell fate decisions. Cell Stem Cell. 2010;7(1):36–41.
Arnold CP et al. MicroRNA programs in normal and aberrant stem and progenitor cells. Genome Res. 2011;21(5):798–810.
Shaham L et al. MiR-125 in normal and malignant hematopoiesis. Leukemia. 2012;26(9):2011–8.
Rhyasen GW, Starczynowski DT. Deregulation of microRNAs in myelodysplastic syndrome. Leukemia. 2012;26(1):13–22.
Pietras EM, Passegue E. Linking HSCs to their youth. Nat Cell Biol. 2013;15(8):885–7.
Gangaraju VK, Lin H. MicroRNAs: key regulators of stem cells. Nat Rev Mol Cell Biol. 2009;10(2):116–25.
Egger G et al. Identification of DNMT1 (DNA methyltransferase 1) hypomorphs in somatic knockouts suggests an essential role for DNMT1 in cell survival. Proc Natl Acad Sci U S A. 2006;103(38):14080–5.
Bernstein BE, Meissner A, Lander ES. The mammalian epigenome. Cell. 2007;128(4):669–81.
Liao J et al. Targeted disruption of DNMT1, DNMT3A and DNMT3B in human embryonic stem cells. Nat Genet. 2015;47(5):469–78.
Tsumura A et al. Maintenance of self-renewal ability of mouse embryonic stem cells in the absence of DNA methyltransferases Dnmt1, Dnmt3a and Dnmt3b. Genes Cells. 2006;11(7):805–14.
Guo Y et al. Characterization of the mammalian miRNA turnover landscape. Nucleic Acids Res. 2015;43(4):2326–41. This study provides evidence that the majority of miRNAs within diverse mammalian cell types are extremely stable, suggesting that some miRNAs may be lost via replication mediated passive dilution similar to other epigenetic modifications.
Ji L, Chen X. Regulation of small RNA stability: methylation and beyond. Cell Res. 2012;22(4):624–36.
Kai ZS, Pasquinelli AE. MicroRNA assassins: factors that regulate the disappearance of miRNAs. Nat Struct Mol Biol. 2010;17(1):5–10.
van Rooij E et al. Control of stress-dependent cardiac growth and gene expression by a microRNA. Science. 2007;316(5824):575–9.
Guo S et al. Nonstochastic reprogramming from a privileged somatic cell state. Cell. 2014;156(4):649–62.
Guo S et al. MicroRNA miR-125a controls hematopoietic stem cell number. Proc Natl Acad Sci U S A. 2010;107(32):14229–34.
Georgantas 3rd RW et al. CD34+ hematopoietic stem-progenitor cell microRNA expression and function: a circuit diagram of differentiation control. Proc Natl Acad Sci U S A. 2007;104(8):2750–5.
Bissels U et al. Combined characterization of microRNA and mRNA profiles delineates early differentiation pathways of CD133+ and CD34+ hematopoietic stem and progenitor cells. Stem Cells. 2011;29(5):847–57.
Landgraf P et al. A mammalian microRNA expression atlas based on small RNA library sequencing. Cell. 2007;129(7):1401–14.
Petriv OI et al. Comprehensive microRNA expression profiling of the hematopoietic hierarchy. Proc Natl Acad Sci U S A. 2010;107(35):15443–8.
Roden C et al. microRNA expression profiling: technologies, insights, and prospects. Adv Exp Med Biol. 2015;888:409–21.
Copley MR et al. The Lin28b-let-7-Hmga2 axis determines the higher self-renewal potential of fetal haematopoietic stem cells. Nat Cell Biol. 2013;15(8):916–25. This study highlights a molecular pathway involving Lin28 and let-7 in controlling HSC cell cycle behavior during neonatal to adult program transition.
Cheng J et al. An extensive network of TET2-targeting MicroRNAs regulates malignant hematopoiesis. Cell Rep. 2013;5(2):471–81. This study identifies a number of miRNAs capable of negatively regulating mouse and human TET2.
Kamat V et al. MicroRNA screen of human embryonic stem cell differentiation reveals miR-105 as an enhancer of megakaryopoiesis from adult CD34+ cells. Stem Cells. 2014;32(5):1337–46.
Fontana L et al. MicroRNAs 17-5p-20a-106a control monocytopoiesis through AML1 targeting and M-CSF receptor upregulation. Nat Cell Biol. 2007;9(7):775–87.
Emmrich S et al. miR-99a/100 ∼ 125b tricistrons regulate hematopoietic stem and progenitor cell homeostasis by shifting the balance between TGFbeta and Wnt signaling. Genes Dev. 2014;28(8):858–74.
Bousquet M et al. Myeloid cell differentiation arrest by miR-125b-1 in myelodysplastic syndrome and acute myeloid leukemia with the t(2;11)(p21;q23) translocation. J Exp Med. 2008;205(11):2499–506.
Klusmann JH et al. miR-125b-2 is a potential oncomiR on human chromosome 21 in megakaryoblastic leukemia. Genes Dev. 2010;24(5):478–90.
Guo S et al. Complex oncogene dependence in microRNA-125a-induced myeloproliferative neoplasms. Proc Natl Acad Sci U S A. 2012;109(41):16636–41.
Tatsumi N, et al. Deficiency in WT1-targeting microRNA-125a leads to myeloid malignancies and urogenital abnormalities. Oncogene. 2015.
Surdziel E et al. Enforced expression of miR-125b affects myelopoiesis by targeting multiple signaling pathways. Blood. 2011;117(16):4338–48.
Bousquet M et al. MicroRNA miR-125b causes leukemia. Proc Natl Acad Sci U S A. 2010;107(50):21558–63.
Lechman ER et al. miR-126 Regulates Distinct Self-Renewal Outcomes in Normal and Malignant Hematopoietic Stem Cells. Cancer Cell. 2016;29(2):214–28. This study identifies a number of miRNAs, including miR-126, which are enriched in AML leukemia stem cells. In contrast to the early-defined role of miR-126 in the HSC, in the LSC elevated expression of miR-126 promotes quiescence and self-renewal.
Lechman ER et al. Attenuation of miR-126 activity expands HSC in vivo without exhaustion. Cell Stem Cell. 2012;11(6):799–811. This study identifies miR-126 as an important regulator of HSC numbers by negatively regulating PI3K/AKT/GSK3β pathway and preventing cell cycle entry.
de Leeuw DC et al. Attenuation of microRNA-126 expression that drives CD34 + 38- stem/progenitor cells in acute myeloid leukemia leads to tumor eradication. Cancer Res. 2014;74(7):2094–105.
Shen WF et al. MicroRNA-126 regulates HOXA9 by binding to the homeobox. Mol Cell Biol. 2008;28(14):4609–19.
Alemdehy MF et al. ICL-induced miR139-3p and miR199a-3p have opposite roles in hematopoietic cell expansion and leukemic transformation. Blood. 2015;125(25):3937–48.
Emmrich S, et al. miR-139-5p controls translation in myeloid leukemia through EIF4G2. Oncogene. 2016;35(14):1822–31.
Nimmo R et al. MiR-142-3p controls the specification of definitive hemangioblasts during ontogeny. Dev Cell. 2013;26(3):237–49. This study highlights the critical role of miR-142 in the specification of xenopus hemangioblast by inhibiting TGFB signaling.
Lu X et al. miR-142-3p regulates the formation and differentiation of hematopoietic stem cells in vertebrates. Cell Res. 2013;23(12):1356–68.
Vian L et al. Transcriptional fine-tuning of microRNA-223 levels directs lineage choice of human hematopoietic progenitors. Cell Death Differ. 2014;21(2):290–301.
Starczynowski DT et al. Identification of miR-145 and miR-146a as mediators of the 5q- syndrome phenotype. Nat Med. 2010;16(1):49–58.
Zhao JL, Starczynowski DT. Role of microRNA-146a in normal and malignant hematopoietic stem cell function. Front Genet. 2014;5:219.
Ghani S et al. Macrophage development from HSCs requires PU.1-coordinated microRNA expression. Blood. 2011;118(8):2275–84.
Starczynowski DT et al. MicroRNA-146a disrupts hematopoietic differentiation and survival. Exp Hematol. 2011;39(2):167–78. e4.
Adams BD et al. An in vivo functional screen uncovers miR-150-mediated regulation of hematopoietic injury response. Cell Rep. 2012;2(4):1048–60.
Stumpel DJ et al. Hypermethylation of specific microRNA genes in MLL-rearranged infant acute lymphoblastic leukemia: major matters at a micro scale. Leukemia. 2011;25(3):429–39.
O’Connell RM et al. Sustained expression of microRNA-155 in hematopoietic stem cells causes a myeloproliferative disorder. J Exp Med. 2008;205(3):585–94.
Zhang H et al. miR-155 is associated with the leukemogenic potential of the class IV granulocyte colony-stimulating factor receptor in CD34(+) progenitor cells. Mol Med. 2014;20:736–46.
Hu YL et al. HOXA9 regulates miR-155 in hematopoietic cells. Nucleic Acids Res. 2010;38(16):5472–8.
O’Connell RM et al. Inositol phosphatase SHIP1 is a primary target of miR-155. Proc Natl Acad Sci U S A. 2009;106(17):7113–8.
O’Connell RM et al. MicroRNA-155 promotes autoimmune inflammation by enhancing inflammatory T cell development. Immunity. 2010;33(4):607–19.
Zhao H et al. The c-myb proto-oncogene and microRNA-15a comprise an active autoregulatory feedback loop in human hematopoietic cells. Blood. 2009;113(3):505–16.
Klein U et al. The DLEU2/miR-15a/16-1 cluster controls B cell proliferation and its deletion leads to chronic lymphocytic leukemia. Cancer Cell. 2010;17(1):28–40.
Diaz-Beya M et al. Acute myeloid leukemia with translocation (8;16)(p11;p13) and MYST3-CREBBP rearrangement harbors a distinctive microRNA signature targeting RET proto-oncogene. Leukemia. 2013;27(3):595–603.
Ventura A et al. Targeted deletion reveals essential and overlapping functions of the miR-17 through 92 family of miRNA clusters. Cell. 2008;132(5):875–86.
Wong P et al. The miR-17-92 microRNA polycistron regulates MLL leukemia stem cell potential by modulating p21 expression. Cancer Res. 2010;70(9):3833–42.
Su R et al. MiR-181 family: regulators of myeloid differentiation and acute myeloid leukemia as well as potential therapeutic targets. Oncogene. 2015;34(25):3226–39.
Chen CZ et al. MicroRNAs modulate hematopoietic lineage differentiation. Science. 2004;303(5654):83–6.
Popovic R et al. Regulation of mir-196b by MLL and its overexpression by MLL fusions contributes to immortalization. Blood. 2009;113(14):3314–22.
Garzon R et al. MicroRNA signatures associated with cytogenetics and prognosis in acute myeloid leukemia. Blood. 2008;111(6):3183–9.
Medina PP, Nolde M, Slack FJ. OncomiR addiction in an in vivo model of microRNA-21-induced pre-B-cell lymphoma. Nature. 2010;467(7311):86–90.
Mehta A et al. The microRNA-132 and microRNA-212 cluster regulates hematopoietic stem cell maintenance and survival with age by buffering FOXO3 expression. Immunity. 2015;42(6):1021–32.
Song SJ et al. The oncogenic microRNA miR-22 targets the TET2 tumor suppressor to promote hematopoietic stem cell self-renewal and transformation. Cell Stem Cell. 2013;13(1):87–101. This study identifies miR-22 as a negative regulator of TET2, and acts as an oncogne.
Fazi F et al. A minicircuitry comprised of microRNA-223 and transcription factors NFI-A and C/EBPalpha regulates human granulopoiesis. Cell. 2005;123(5):819–31.
Fazi F et al. Epigenetic silencing of the myelopoiesis regulator microRNA-223 by the AML1/ETO oncoprotein. Cancer Cell. 2007;12(5):457–66.
Johnnidis JB et al. Regulation of progenitor cell proliferation and granulocyte function by microRNA-223. Nature. 2008;451(7182):1125–9.
Ahn EE et al. SON protein regulates GATA-2 through transcriptional control of the microRNA 23a ∼ 27a ∼ 24-2 cluster. J Biol Chem. 2013;288(8):5381–8.
Han YC et al. microRNA-29a induces aberrant self-renewal capacity in hematopoietic progenitors, biased myeloid development, and acute myeloid leukemia. J Exp Med. 2010;207(3):475–89.
Hu W et al. miR-29a maintains mouse hematopoietic stem cell self-renewal by regulating Dnmt3a. Blood. 2015;125(14):2206–16. This study identifies the functional importance of miR-29a and its targeting of Dnmt3a in mouse HSCs.
Herrera-Merchan A et al. miR-33-mediated downregulation of p53 controls hematopoietic stem cell self-renewal. Cell Cycle. 2010;9(16):3277–85.
Dore LC et al. A GATA-1-regulated microRNA locus essential for erythropoiesis. Proc Natl Acad Sci U S A. 2008;105(9):3333–8.
Jiang X et al. Blockade of miR-150 maturation by MLL-fusion/MYC/LIN-28 is required for MLL-associated leukemia. Cancer Cell. 2012;22(4):524–35.
Chen P et al. miR-9 is an essential oncogenic microRNA specifically overexpressed in mixed lineage leukemia-rearranged leukemia. Proc Natl Acad Sci U S A. 2013;110(28):11511–6.
Jaiswal S et al. Age-related clonal hematopoiesis associated with adverse outcomes. N Engl J Med. 2014;371(26):2488–98.
Xie M et al. Age-related mutations associated with clonal hematopoietic expansion and malignancies. Nat Med. 2014;20(12):1472–8.
Genovese G et al. Clonal hematopoiesis and blood-cancer risk inferred from blood DNA sequence. N Engl J Med. 2014;371(26):2477–87.
Busque L et al. Recurrent somatic TET2 mutations in normal elderly individuals with clonal hematopoiesis. Nat Genet. 2012;44(11):1179–81.
Jan M et al. Clonal evolution of preleukemic hematopoietic stem cells precedes human acute myeloid leukemia. Sci Transl Med. 2012;4(149):149ra118.
Delhommeau F et al. Mutation in TET2 in myeloid cancers. N Engl J Med. 2009;360(22):2289–301.
Langemeijer SM et al. Acquired mutations in TET2 are common in myelodysplastic syndromes. Nat Genet. 2009;41(7):838–42.
Ley TJ et al. DNMT3A mutations in acute myeloid leukemia. N Engl J Med. 2010;363(25):2424–33.
Welch JS et al. The origin and evolution of mutations in acute myeloid leukemia. Cell. 2012;150(2):264–78.
Shlush LI et al. Identification of pre-leukaemic haematopoietic stem cells in acute leukaemia. Nature. 2014;506(7488):328–33.
Tahiliani M et al. Conversion of 5-methylcytosine to 5-hydroxymethylcytosine in mammalian DNA by MLL partner TET1. Science. 2009;324(5929):930–5.
Ito S et al. Tet proteins can convert 5-methylcytosine to 5-formylcytosine and 5-carboxylcytosine. Science. 2011;333(6047):1300–3.
He YF et al. Tet-mediated formation of 5-carboxylcytosine and its excision by TDG in mammalian DNA. Science. 2011;333(6047):1303–7.
Hashimoto H et al. Recognition and potential mechanisms for replication and erasure of cytosine hydroxymethylation. Nucleic Acids Res. 2012;40(11):4841–9.
Challen GA et al. Dnmt3a and Dnmt3b have overlapping and distinct functions in hematopoietic stem cells. Cell Stem Cell. 2014;15(3):350–64.
Challen GA et al. Dnmt3a is essential for hematopoietic stem cell differentiation. Nat Genet. 2012;44(1):23–31.
Moran-Crusio K et al. Tet2 loss leads to increased hematopoietic stem cell self-renewal and myeloid transformation. Cancer Cell. 2011;20(1):11–24.
Quivoron C et al. TET2 inactivation results in pleiotropic hematopoietic abnormalities in mouse and is a recurrent event during human lymphomagenesis. Cancer Cell. 2011;20(1):25–38.
Li Z et al. Deletion of Tet2 in mice leads to dysregulated hematopoietic stem cells and subsequent development of myeloid malignancies. Blood. 2011;118(17):4509–18.
Ko M et al. Ten-eleven-translocation 2 (TET2) negatively regulates homeostasis and differentiation of hematopoietic stem cells in mice. Proc Natl Acad Sci U S A. 2011;108(35):14566–71.
Wojtowicz EE et al. MicroRNA-125 family members exert a similar role in the regulation of murine hematopoiesis. Exp Hematol. 2014;42(10):909–18. e1.
O’Connell RM et al. MicroRNAs enriched in hematopoietic stem cells differentially regulate long-term hematopoietic output. Proc Natl Acad Sci U S A. 2010;107(32):14235–40.
Ooi AG et al. MicroRNA-125b expands hematopoietic stem cells and enriches for the lymphoid-balanced and lymphoid-biased subsets. Proc Natl Acad Sci U S A. 2010;107(50):21505–10.
Sonoki T et al. Insertion of microRNA-125b-1, a human homologue of lin-4, into a rearranged immunoglobulin heavy chain gene locus in a patient with precursor B-cell acute lymphoblastic leukemia. Leukemia. 2005;19(11):2009–10.
Gefen N et al. Hsa-mir-125b-2 is highly expressed in childhood ETV6/RUNX1 (TEL/AML1) leukemias and confers survival advantage to growth inhibitory signals independent of p53. Leukemia. 2010;24(1):89–96.
Zhang P et al. Ten-eleven translocation (Tet) and thymine DNA glycosylase (TDG), components of the demethylation pathway, are direct targets of miRNA-29a. Biochem Biophys Res Commun. 2013;437(3):368–73.
Kats LM et al. Proto-oncogenic role of mutant IDH2 in leukemia initiation and maintenance. Cell Stem Cell. 2014;14(3):329–41.
Jiang X et al. miR-22 has a potent anti-tumour role with therapeutic potential in acute myeloid leukaemia. Nat Commun. 2016;7:11452.
Fu X et al. MicroRNA-26a targets ten eleven translocation enzymes and is regulated during pancreatic cell differentiation. Proc Natl Acad Sci U S A. 2013;110(44):17892–7.
Varambally S et al. Genomic loss of microRNA-101 leads to overexpression of histone methyltransferase EZH2 in cancer. Science. 2008;322(5908):1695–9.
Teng G et al. MicroRNA-155 is a negative regulator of activation-induced cytidine deaminase. Immunity. 2008;28(5):621–9.
de Yebenes VG et al. miR-181b negatively regulates activation-induced cytidine deaminase in B cells. J Exp Med. 2008;205(10):2199–206.
Garzon R et al. MicroRNA-29b induces global DNA hypomethylation and tumor suppressor gene reexpression in acute myeloid leukemia by targeting directly DNMT3A and 3B and indirectly DNMT1. Blood. 2009;113(25):6411–8.
Garzon R et al. MicroRNA 29b functions in acute myeloid leukemia. Blood. 2009;114(26):5331–41.
Huang H et al. TET1 plays an essential oncogenic role in MLL-rearranged leukemia. Proc Natl Acad Sci U S A. 2013;110(29):11994–9.
Chen BF et al. microRNA-199a-3p, DNMT3A, and aberrant DNA methylation in testicular cancer. Epigenetics. 2014;9(1):119–28.
Zhao S et al. MicroRNA-126 regulates DNA methylation in CD4+ T cells and contributes to systemic lupus erythematosus by targeting DNA methyltransferase 1. Arthritis Rheum. 2011;63(5):1376–86.
Pan W et al. MicroRNA-21 and microRNA-148a contribute to DNA hypomethylation in lupus CD4+ T cells by directly and indirectly targeting DNA methyltransferase 1. J Immunol. 2010;184(12):6773–81.
Babovic S, Eaves CJ. Hierarchical organization of fetal and adult hematopoietic stem cells. Exp Cell Res. 2014;329(2):185–91.
Sturgeon CM et al. Primitive erythropoiesis is regulated by miR-126 via nonhematopoietic Vcam-1+ cells. Dev Cell. 2012;23(1):45–57. This study identifies miR-126 as a key negative regulator of primitive erythropoiesis.
Qian P et al. The Dlk1-Gtl2 Locus Preserves LT-HSC Function by Inhibiting the PI3K-mTOR Pathway to Restrict Mitochondrial Metabolism. Cell Stem Cell. 2016;18(2):214–28.
Reinhart BJ et al. The 21-nucleotide let-7 RNA regulates developmental timing in Caenorhabditis elegans. Nature. 2000;403(6772):901–6.
Piskounova E et al. Lin28A and Lin28B inhibit let-7 microRNA biogenesis by distinct mechanisms. Cell. 2011;147(5):1066–79.
Thornton JE et al. Lin28-mediated control of let-7 microRNA expression by alternative TUTases Zcchc11 (TUT4) and Zcchc6 (TUT7). RNA. 2012;18(10):1875–85.
Newman MA, Thomson JM, Hammond SM. Lin-28 interaction with the Let-7 precursor loop mediates regulated microRNA processing. RNA. 2008;14(8):1539–49.
Mayr C, Hemann MT, Bartel DP. Disrupting the pairing between let-7 and Hmga2 enhances oncogenic transformation. Science. 2007;315(5818):1576–9.
Johnson SM et al. RAS is regulated by the let-7 microRNA family. Cell. 2005;120(5):635–47.
Kumar MS et al. Impaired microRNA processing enhances cellular transformation and tumorigenesis. Nat Genet. 2007;39(5):673–7.
Yuan J et al. Lin28b reprograms adult bone marrow hematopoietic progenitors to mediate fetal-like lymphopoiesis. Science. 2012;335(6073):1195–200.
Zhou Y et al. Lin28b promotes fetal B lymphopoiesis through the transcription factor Arid3a. J Exp Med. 2015;212(4):569–80.
Seita J, Weissman IL. Hematopoietic stem cell: self-renewal versus differentiation. Wiley Interdiscip Rev Syst Biol Med. 2010;2(6):640–53.
Fleming WH et al. Functional heterogeneity is associated with the cell cycle status of murine hematopoietic stem cells. J Cell Biol. 1993;122(4):897–902.
Cheung TH, Rando TA. Molecular regulation of stem cell quiescence. Nat Rev Mol Cell Biol. 2013;14(6):329–40.
Geiger H, de Haan G, Florian MC. The ageing haematopoietic stem cell compartment. Nat Rev Immunol. 2013;13(5):376–89.
Esplin BL et al. Chronic exposure to a TLR ligand injures hematopoietic stem cells. J Immunol. 2011;186(9):5367–75.
Rusca N, Monticelli S. MiR-146a in immunity and disease. Mol Biol Int. 2011;2011:437301.
Kumar MS et al. Coordinate loss of a microRNA and protein-coding gene cooperate in the pathogenesis of 5q- syndrome. Blood. 2011;118(17):4666–73.
Zhao JL et al. MicroRNA-146a acts as a guardian of the quality and longevity of hematopoietic stem cells in mice. Elife. 2013;2:e00537. This study shows that loss of miR-146a, a critical negative regulator of inflammation-induced hematopoiesis in the HSC, leads to HSC exhaustion, bone marrow failure and oncogenic transformation.
Raaijmakers MH et al. Bone progenitor dysfunction induces myelodysplasia and secondary leukaemia. Nature. 2010;464(7290):852–7. This study shows loss of Dicer in bone progenitors induces alterations in the hematopoietic cells.
Balakrishnan I et al. Genome-wide analysis of miRNA-mRNA interactions in marrow stromal cells. Stem Cells. 2014;32(3):662–73.
Charbord P et al. A systems biology approach for defining the molecular framework of the hematopoietic stem cell niche. Cell Stem Cell. 2014;15(3):376–91.
Wang L et al. Notch-dependent repression of miR-155 in the bone marrow niche regulates hematopoiesis in an NF-kappaB-dependent manner. Cell Stem Cell. 2014;15(1):51–65. This study shows that deregulation of miR-155 in bone marrow niche leads to myeloid defects.
Phinney DG, Isakova IA. Mesenchymal stem cells as cellular vectors for pediatric neurological disorders. Brain Res. 2014;1573:92–107.
Zomer A et al. In vivo imaging reveals extracellular vesicle-mediated phenocopying of metastatic behavior. Cell. 2015;161(5):1046–57.
Marcucci G et al. IDH1 and IDH2 gene mutations identify novel molecular subsets within de novo cytogenetically normal acute myeloid leukemia: a Cancer and Leukemia Group B study. J Clin Oncol. 2010;28(14):2348–55.
Schaar DG et al. miR-320 targets transferrin receptor 1 (CD71) and inhibits cell proliferation. Exp Hematol. 2009;37(2):245–55.
Solary E et al. The ten-eleven translocation-2 (TET2) gene in hematopoiesis and hematopoietic diseases. Leukemia. 2014;28(3):485–96.
Lund K, Adams PD, Copland M. EZH2 in normal and malignant hematopoiesis. Leukemia. 2014;28(1):44–9.
Ebert MS, Sharp PA. Roles for microRNAs in conferring robustness to biological processes. Cell. 2012;149(3):515–24.
Ma X et al. MicroRNAs in NF-kappaB signaling. J Mol Cell Biol. 2011;3(3):159–66.
Chatterjee S et al. Target-mediated protection of endogenous microRNAs in C. elegans. Dev Cell. 2011;20(3):388–96.
van der Harst P et al. Seventy-five genetic loci influencing the human red blood cell. Nature. 2012;492(7429):369–75.
Stenvang J et al. Inhibition of microRNA function by antimiR oligonucleotides. Silence. 2012;3(1):1.
Cheng CJ et al. MicroRNA silencing for cancer therapy targeted to the tumour microenvironment. Nature. 2015;518(7537):107–10.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
Christine Roden and Jun Lu declare that they have no conflict of interest.
Human and Animal Rights and Informed Consent
This article does not contain any studies with human or animal subjects performed by any of the authors.
Additional information
This article is part of the Topical Collection on Stem Cell Switches and Regulators
Rights and permissions
About this article
Cite this article
Roden, C., Lu, J. MicroRNAs in Control of Stem Cells in Normal and Malignant Hematopoiesis. Curr Stem Cell Rep 2, 183–196 (2016). https://doi.org/10.1007/s40778-016-0057-1
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40778-016-0057-1