Abstract
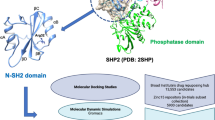
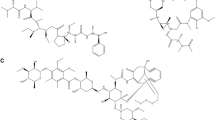
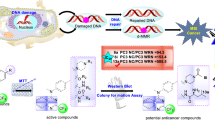
Human lemur tyrosine kinase-3 (LMTK3) is primarily involved in regulation of estrogen receptor–α (ERα) by phosphorylation activity. LMTK3 acts as key biomarker for ERα positive breast cancer and identified as novel drug target for breast cancer. Due to the absence of experimental reports, the computational approach has been followed to screen LMTK3 inhibitors from natural product curcumin derivatives based on rational inhibitor design. The initial virtual screening and re-docking resulted in identification of top three leads with favorable binding energy and strong interactions in critical residues of ATP-binding cavity. ADME prediction confirmed the pharmacological activity of the leads with various properties. The stability and binding affinity of leads were well refined in dynamic system from 25 ns MD simulations. The behavior of protein motion towards closure of ATP-binding cavity was evaluated based on eigenvectors by PCA. In addition, MM/PBSA calculations also confirmed the relative binding free energy of LMTK3–lead complexes in favor of the effective binding. From our study, novel LMTK3 inhibitors tetrahydrocurcumin, curcumin 4,4′-diacetate, and demethoxycurcumin have been proposed with inhibition mechanism. Further experimental evaluation on reported lead candidates might prove its role in breast cancer therapeutics.






Similar content being viewed by others
References
Aggarwal BB, Sundaram C, Malani N, Ichikawa H (2007) Curcumin: the Indian solid gold. Adv Exp Med Biol 595:1–75
Amadei A, Linssen AB, Berendsen HJ (1993) Essential dynamics of proteins. Proteins 17:412–425
Anbarasu K, Jayanthi S (2014) Structural modeling and molecular dynamics studies on the human LMTK3 domain and the mechanism of ATP binding. Mol BioSyst 10:1139–1145
Baker NA, Sept D, Joseph S, Holst MJ, McCammon JA (2001) Electrostatics of nanosystems: application to microtubules and the ribosome. Proc Natl Acad Sci USA 98:10037–10041
Berendsen HJC, van der Spoel D, van Drunen R (1995) Gromacs: a message-passing parallel molecular dynamics implementation. Comput Phys Commun 91:43–56
Creighton DJ, Zheng ZB, Holewinski R, Hamilton DS, Eiseman JL (2003) Glyoxalase I inhibitors in cancer chemotherapy. Biochem Soc Trans 31:1378–1382
Giamas G, Stebbing J, Vorgias CE, Knippschild U (2007) Protein kinases as targets for cancer treatment. Pharmacogenomics 8:1005–1016
Giamas G, Filipovic A, Jacob J, Messier W, Zhang H (2011) Kinome screening for regulators of the estrogen receptor identifies LMTK3 as a new therapeutic target in breast cancer. Nat Med 17(6):715–719
Guan SS, Han WW, Zhang H, Wang S, Shan YM (2016) Insight into the interactive residues between two domains of human somatic Angiotensin-converting enzyme and Angiotensin II by MM-PBSA calculation and steered molecular dynamics simulation. J Biomol Struct Dyn 34(1):15–28
Hess HB, Bekker H, Berendsen HJC, Fraaije JGEM (1996) LINCS: a linear constraint solver for molecular simulations. J Comput Chem 18:1463–1472
Johnson AB, O’Malley BW (2011) ERasing breast cancer resistance through the kinome. Nat Med 17(6):660–661
Jurenka JS (2009) Anti-inflammatory properties of curcumin, a major constituent of Curcuma longa: a review of preclinical and clinical research. Altern Med Rev 14:141–153
Kumari R, Kumar R, Lynn A, Open Source Drug Discovery Consortium (2014) g_mmpbsa-a GROMACS tool for high-throughput MM-PBSA calculations. J Chem Inf Model 54(7):1951–1962
Labrie F, Labrie C, Belanger A, Simard J, Gauthier S, Luu-The V, Merand Y et al (1999) EM-652 (SCH 57068), a third generation SERM acting as pure antiestrogen in the mammary gland and endometrium. J Steroid Biochem Mol Biol 69:51–84
Laskowski RA, Swindells MB (2011) LigPlot+: multiple ligand-protein interaction diagrams for drug discovery. J Chem Inf Model 51:2778–2786
Lin JK (2007) Molecular targets of curcumin. Adv Exp Med Biol 595:227–243
Lindahl E, Hess B, van Spoel D (2001) GROMACS 3.0: a package for molecular simulation and trajectory analysis. J Mol Model 7(8):306–317
Lipinski CA, Lombardo F, Dominy BW, Feeney PJ (1997) Toward minimalistic modeling of oral drug absorption. Adv Drug Deliv Rev 23:3–25
Miyamoto S, Kollman PA (1992) Settle: an analytical version of the SHAKE and RATTLE algorithm for rigid water models. J Comput Chem 13:952–962
Morris GM, Goodsell DS, Halliday RS, Huey R, Hart WE, Belew RK, Olson AJ (1998) Automated docking using a Lamarckian genetic algorithm and an empirical binding free energy function. J Comput Chem 19:1639–1662
Robinson DR, Wu YM, Lin SF (2000) The protein tyrosine kinase family of the human genome. Oncogene 19:5548–5557
Schuttelkopf AW, van Aalten DMF (2004) PRODRG: a tool for high-throughput crystallography of protein-ligand complexes. Acta Crystallogr D Biol Crystallogr 60(8):1355–1363
Seeliger D, de Groot BL (2010) Ligand docking and binding site analysis with PyMOL and Autodock/Vina. J Comput Aided Mol Des 24(5):417–422
Small-Molecule Drug Discovery Suite 2015-3: QikProp, version 4.5, Schrödinger, LLC, New York, NY
Stebbing J, Filipovic A, Giamas G (2011) Lemur tyrosine kinase-3 (LMTK3) in cancer and evolution. Oncotarget 2:428–429
Stebbing J, Filipovic A, Ellis IO, Green AR, D’Silva TR, Lenz HJ, Coombes RC, Wang T, Lee SC, Giamas G (2012) LMTK3 expression in breast cancer: association with tumor phenotype and clinical outcome. Breast Cancer Res Treat 132(2):537–544
Tetko IV, Gasteiger J, Todeschini R, Mauri A, Livingstone D et al (2005) Virtual computational chemistry laboratory–design and description. J Comput Aided Mol Des 19:453–463
Trott O, Olson AJ (2010) AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization and multithreading. J Comput Chem 31:455–461
Turner PJ (2005) XMGRACE, Version 5.1.19. Center for Coastal and Land-Margin Research, Oregon Graduate Institute of Science and Technology, Beaverton
Wilken R, Veena MS, Wang MB, Srivatsan ES (2011) Curcumin: a review of anti-cancer properties and therapeutic activity in head and neck squamous cell carcinoma. Mol Cancer 10:12
Acknowledgements
The authors thank the management of Vellore Insititute of Technology (VIT) for providing the facilities to carry out this work.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Anbarasu, K., Jayanthi, S. Identification of curcumin derivatives as human LMTK3 inhibitors for breast cancer: a docking, dynamics, and MM/PBSA approach. 3 Biotech 8, 228 (2018). https://doi.org/10.1007/s13205-018-1239-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13205-018-1239-6