Abstract
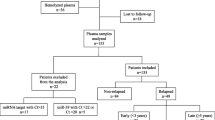
Circulating miR-21 is upregulated in breast cancer. However, correlation of miR-21 expression with clinic pathologic characteristics remains questionable. In this study, we investigate whether combination of circulation miR-21 with circulating tumor cells (CTCs) marker (EpCAM, MUS1, HER2) could improve diagnostic specificity of metastatic breast cancer. Total 223 breast cancer patients were included. 89 % patients were associated with upregulation of miR-21 compared with health control. 20 % patients were detected for CTCs marker positive. For higher specificity purpose, triple marker positive samples were selected as true CTCs positive, which only occupied 59.5 % of total metastatic breast cancer patients. Specificity of detection of CTCs was 96.7 %. Furthermore, 59.5 % metastatic breast cancer patients were shown both abnormal miR-21 and true CTCs positive according to distribution of true CTCs positive and abnormal miR-21; Combination of miR-21 and CTCs was increased specificity of metastatic detection to 100 %. Our findings suggested that combination of miR-21 with CTCs marker could be used for better diagnosis of metastatic breast cancer in the future.



Similar content being viewed by others
References
DeSantis, C. E., Lin, C. C., Mariotto, A. B., et al. (2014). Cancer treatment and survivorship statistics, 2014. CA: A Cancer Journal for Clinicians, 64(4), 252–271.
Cristofanilli, M., Ellis, M. J., Budd, G. T., et al. (2004). Circulating tumor cells, disease progression, and survival in metastatic breast cancer. New England Journal of Medicine, 351, 781–791.
Cristofanilli, M., Hayes, D. F., Budd, G. T., et al. (2005). Circulating tumor cells: a novel prognostic factor for newly diagnosed metastatic breast cancer. Journal of Clinical Oncology, 23, 1420–1430.
Borgen, E., Wiedswang, G., Karesen, R., et al. (2004). Isolatedtumor cells in bone marrow three years after diagnosis in disease-free breast cancer patients predict unfavorable clinicaloutcome. Clinical Cancer Research, 10, 5342–5348.
Gilbey, A. M., Burnett, D., Coleman, R. E., & Holen, I. (2004). The detection of circulating breast cancer cells in blood. Journal of Clinical Pathology, 57, 903–911.
Okumura, Y., Jotsuka, T., Nakano, S., et al. (2004). Persistent evidence of circulating tumor cells detected by means of RT-PCR for CEAmRNA predicts early relapse: a prospective study in node-negative breast cancer. Surgery, 135, 419–426.
Kwon, S., Kang, S. H., Ro, J., Jeon, C. H., Park, J. W., & Lee, E. S. (2005). The melanoma antigen gene as a surveillance marker for the detection of circulating tumor cells in patients with breast carcinoma. Cancer, 104, 251–256.
Miyashiro, I., Huynh, K., Kuo, C., et al. (2001). Molecular strategy for detecting metastatic cancers with use of multiple tumor-specific MAGE-A genes. Clinical Chemistry, 47, 505–512.
Reinholz, M. M., Nibbe, A., Jonart, L. M., et al. (2005). Evaluation of a panel of tumor markers for molecular detection of circulating cancer cells in women with suspected breast cancer. Clinical Cancer Research, 11, 3722–3732.
Ring, A. E., Zabaglo, L., Ormerod, M. G., Smith, I. E., & Dowsett, M. (2005). Detection of circulating epithelial cells in the blood of patients with breast cancer: comparison of three techniques. British Journal of Cancer, 92, 906–912.
Sabbatini, R., Morselli, M., Federico, M., et al. (2000). Detection of circulating tumor cells by reverse transcriptase polymerase chain reaction of maspin in patients with breast cancer undergoing conventional-dose chemotherapy. Journal of Clinical Oncology, 18, 1914–1920.
Taback, B., Chan, A. D., Kuo, C. T., et al. (2001). Detection of occult metastatic breast cancer cells in blood by a multimolecular marker assay: Correlation with clinical stage of disease. Cancer Research, 61, 8845–8850.
Weigelt, B., Bosma, A. J., Hart, A. A., Rodenhuis, S., & van’t Veer, L. J. (2003). Marker genes for circulating tumour cells predict survival in metastasized breast cancer patients. British Journal of Cancer, 88, 1091–1094.
Hayes, D. F., Budd, G. T., Cristofanilli, M., et al. (2006). Circulating tumor cells at each follow-up time point during therapy of metastatic breast cancer patients predict progression-free and overall survival. Clinical Cancer Research, 12, 4218–4224.
Zabaglo, L., Ormerod, M. G., Parton, M., Ring, A., Smith, I. E., & Dowsett, M. (2003). Cell filtration-laser scanning cytometry for the characterisation of circulating breast cancer cells. Cytometry A, 55, 102–108.
Zehentner, B. K., Secrist, H., Hayes, D. C., et al. (2006). Detection of circulating tumor cells in peripheral blood of breast cancer patients during or after therapy using a multigene real-time RT-PCR assay. Molecular Diagnosis and Therapy, 10, 41–47.
Zhang, B., Pan, X., Cobb, G. P., et al. (2007). MicroRNAs as oncogenes and tumor suppressors. Development Biology, 302, 1–12.
Zheng, D. L., Haddadin, S., Wang, Y., et al. (2011). Plasma microRNAs as novel biomarkers for early detection of lung cancer. International Journal of Clinical and Experimental Pathology, 4, 575–586.
Li, T., Leong, M. H., Harms, B., et al. (2013). MicroRNA-21 as a potential colon and rectal cancer biomarker. World Journal of Gastroenterology, 19(34), 5615–5621.
Yang, X. R., Du Gao, Y. N., et al. (2013). Serum microRNA-21 as a diagnostic marker for lung carcinoma: A systematic review and meta analysis. Plos One, 9(5), e97460.
Zeng, Z. Y., Wang, J. G., Zhao, L. Y., et al. (2013). Potential role of microRNA-21 in the diagnosis of gastric cancer: a meta-analysis. Plos One, 8(9), e73278.
Kroh, E. M., Parkin, R. K., Mitchell, P. S., & Tewari, M. (2010). Analysis of circulating microRNA biomarkers in plasma and serum using quantitative reverse transcription-PCR (qRT-PCR). Methods, 50(4), 298–301.
Phua, Y. L., & Ho, J. (2014). MicroRNAs in the pathogenesis of cystic kidney disease. Current Opinion in Pediatrics [Epub ahead of print].
Alečković, M., & Kang, Y. (2014). Regulation of cancer metastasis by cell-free miRNAs. Biochimica et Biophysica Acta, 1855(1), 24–42.
Gallach, S., Calabuig-Fariñas, S., Jantus-Lewintre, E., & Camps, C. (2014). MicroRNAs:promising new antiangiogenic targets in cancer. BioMed Research International, 2014, 878450.
Suzuki, H. I., Katsura, A., Matsuyama, H., & Miyazono, K. (2014). MicroRNA regulons in tumor microenvironment. Oncogene. doi:10.1038/onc.2014.254.
Croset, M., Santini, D., Iuliani, M., Fioramonti, M., Zoccoli, A., Vincenzi, B., et al. (2014). MicroRNAs and bone metastasis: a new challenge. Molecules, 19(7), 10115–10128.
Sekar, D., Hairul Islam, V. I., Thirugnanasambantham, K., & Saravanan, S. (2014). Relevance of miR-21 in HIV and non-HIV-related lymphomas. Tumor Biology, 35(9), 8387–8393.
Kang, C., Song, J. J., Lee, J., & Kim, M. Y. (2014). Epigenetics: An emerging player in gastric cancer. World Journal Gastroenterol., 20(21), 6433–6447.
Iorio, M. V., Ferracin, M., Liu, C. G., Veronese, A., Spizzo, R., Sabbioni, S., et al. (2005). MicroRNA gene expression deregulation in human breast cancer. Cancer Research, 65(16), 7065–7070.
Author information
Authors and Affiliations
Corresponding author
Additional information
Xingwang Yang and Xiaoming Wang have contributed equally to this work.
Rights and permissions
About this article
Cite this article
Yang, X., Wang, X., Shen, H. et al. Combination of miR-21 with Circulating Tumor Cells Markers Improve Diagnostic Specificity of Metastatic Breast Cancer. Cell Biochem Biophys 73, 87–91 (2015). https://doi.org/10.1007/s12013-015-0573-0
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12013-015-0573-0