Abstract
Marfan syndrome (MFS, OMIM #154700) is a hereditary connective tissue disorder, clinically presenting with cardinal features of skeletal, ocular, and cardiovascular systems. In classical MFS, changes in connective tissue integrity can be explained by defects in fibrillin-1, a major component of extracellular microfibrils. However, some of the clinical manifestations of MFS cannot be explained by mechanical properties alone. Recent studies manipulating mouse Fbn1 have provided new insights into the molecular pathogenesis of MFS. Dysregulation of transforming growth factor beta (TGFβ) signaling in lung, mitral valve and aortic tissues has been implicated in mouse models of MFS. TGFBR2 and TGFBR1 mutations were identified in a subset of patients with MFS (MFS2, OMIM #154705) and other MFS-related disorders, including Loeys–Dietz syndrome (LDS, #OMIM 609192) and familial thoracic aortic aneurysms and dissections (TAAD2, #OMIM 608987). These data indicate that genetic heterogeneity exists in MFS and its related conditions and that regulation of TGFβ signaling plays a significant role in these disorders.
Similar content being viewed by others
Introduction
Marfan syndrome (MFS, OMIM #154700) is a connective tissue disorder with autosomal dominant inheritance. MFS is clinically diagnosed according to the Ghent criteria, which describe pleiotropic manifestations affecting multiple organs (De Paepe et al. 1996). Typical MFS can affect the skeletal system (marfanoid habitus including arachnodactyly, dolichostenomelia, pectus deformity and scoliosis), the ocular system (ectopia lentis) and the cardiovascular system (aortic aneurysm/dissection and mitral regurgitation), as well as other systems, including lung, skin, integument, and dura mater. Significant phenotypic variability of MFS is commonly observed between affected members of different families and even among affected members within a single family. Neonatal MFS (nMFS) is the most severe type of MFS and is characterized by severe atrioventricular valve dysfunction, arachnodactyly, joint contracture, crumpled ears and pectus deformity. In addition to classic MFS, incomplete forms of MFS are seen, in which symptoms overlap with those of MFS but the phenotypes do not satisfy the Ghent criteria.
This review focuses on the recent advances in the genetics of MFS and its associated conditions, including Loeys–Dietz syndrome, non-syndromic thoracic aortic aneurysms and dissections, and Shprintzen–Goldberg craniosynostosis syndrome. Abnormal transforming growth factor beta (TGFβ) signaling will be discussed as the core pathogenesis of MFS.
Genetics of Marfan syndrome and its related disorders
FBN1 mutation-related disorders
Marfan syndrome
Molecular diagnosis of MFS became possible after mutations had been identified in the FBN1 gene (Dietz et al. 1991; Lee et al. 1991). FBN1 is a 230 kb gene, containing 65 exons, which encodes the structural protein fibrillin-1 (Corson et al. 1993). More than 600 FBN1 mutations are registered in the UMD-FBN1 database for MFS and its associated disorders (http://www.umd.be:2030/) (Collod-Beroud et al. 2003). The mutation detection rate of FBN1 in MFS varies among studies, ranging from 9% to 91% (Katzke et al. 2002; Loeys et al. 2004; Tynan et al. 1993). This variability could be explained, in part, by the different techniques used, but the most significant influencing factor is likely to be sample bias. The frequencies are quite different between patients fulfilling the Ghent criteria and those not fulfilling them (Biggin et al. 2004; Halliday et al. 2002; Loeys et al. 2001; Rommel et al. 2002, 2005; Tynan et al. 1993).
Extensive mutational analyses failed to show FBN1 involvement in almost 10% or more of patients with MFS satisfying the Ghent criteria. Although one possible explanation could be due to so-called missing mutations in the promoter region or in other non-coding sequences, the existence of a second locus for MFS (MFS2) was hypothesized (Collod et al. 1994; Dietz et al. 1995; Gilchrist 1994). In 2004, patients with MFS2 were shown to have mutations in the TGFBR2 gene, which encodes the transmembrane receptor type II of TGFβ (Mizuguchi et al. 2004). TGFBR2 mutations were later linked to other clinically overlapping syndromes, described below (Kosaki et al. 2006; Loeys et al. 2005; Pannu et al. 2005a).
Other fibrillinopathies
FBN1 mutations were also found in incomplete forms of MFS as well as in several other MFS-related disorders such as nMFS, isolated ectopia lentis, Shprintzen–Goldberg craniosynostosis syndrome (SGS), familial thoracic aortic aneurysms and dissections (TAAD), and autosomal dominant Weill–Marchesani syndrome (Table 1) (Faivre et al. 2003; Francke et al. 1995; Kainulainen et al. 1994; Milewicz et al. 1996; Sood et al. 1996). This resulted in the recognition of “fibrillinopathies” caused by FBN1 aberrations (Charbonneau et al. 2004).
TGFBR mutation-related disorders
Marfan syndrome type 2
In 1993 Boileau et al. (1993) reported a large French family (MS1 family) with a Marfan-like phenotype that was not linked to the FBN1 locus. The syndrome was subsequently designated Marfan syndrome type 2 (MFS2, OMIM #154705). In this review we are defining MFS2 genetically (not clinically) as the classic MFS phenotype (based on the Ghent criteria) caused by mutation in the TGFBR2 locus.
Marfan-like symptoms observed in this family consisted of severe cardiovascular findings, including sudden death of affected members at young age, probably due to a thoracic aortic dissection, and typical MFS skeletal features, but no significant ocular findings were seen. However, one affected family member (IV-83 from the large French family, MS1) was recently reported to suffer from ectopia lentis, which is clinically compatible with classic MFS according to the Ghent criteria but surprising in light of the absence of the condition in other family members (Mizuguchi et al. 2004). Clinical re-evaluation of this individual as well as other affected members is warranted.
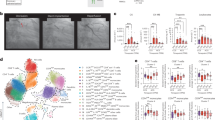
Genetic analysis of the French family enabled a successful mapping of the second locus for MFS (MFS2) to 3p24.2-p25 (Collod et al. 1994). A Japanese boy with MFS was later shown to have de novo complex chromosomal rearrangements involving 3p24.1, which is close to the MFS2 locus (Mizuguchi et al. 2004). Detailed genomic analysis revealed that the 3p24.1 breakpoint disrupted the TGFBR2 gene. Subsequent TGFBR2 sequence analysis in the MS1 family identified in all affected members a nucleotide substitution c.1524G > A (p.Q508Q) of TGFBR2, which is hypothesized to result in protein truncation due to abnormal splicing. Further analysis of 19 unrelated probands unlinked to FBN1 identified c.923T > C (p.L308P), c.1346C > T (p.S449F) and c.1609C > T (p.R537C; recurrent in two independent families) (Table 2 and Fig. 1). These missense mutations affect an evolutionarily conserved amino acid in the serine/threonine kinase domain of TGFβ receptor type II. This finding confirmed that the MFS phenotype can be caused not only by mutations in FBN1 but also by mutations in TGFBR2.
Mutation spectrum of TGFBR2 and TGFBR1 identified in Marfan syndrome and its related disorders. Numbered boxes indicate exons of each gene. Open and filled circles represent missense and splicing abnormality mutations, respectively. Each mutation was found in a different family. Functional domains are shown as patterned boxes. MFS Marfan syndrome, LDS Loeys-Dietz syndrome, TAAD Familial thoracic aortic aneurysms and dissections, SGS Shprintzen-Goldberg craniosynostosis syndrome, FS Furlong syndrome
Loeys–Dietz syndrome
Loeys et al. (2005) reported a new aortic aneurysm syndrome presenting with cardiovascular and skeletal manifestations consistent with those seen in MFS, along with other features not present in MFS. Loeys–Dietz syndrome (LDS, OMIM #609192) is characterized by hypertelorism, bifid uvula, cleft palate, generalized arterial tortuosity, and ascending aortic aneurysm and dissection. Hypothesizing that abnormal TGFβ signaling might cause vascular and craniofacial phenotypes, Loeys et al. (2005) investigated TGFBR2 as a candidate gene for LDS. Heterozygous TGFBR2 mutations were found in six of ten LDS patients: five missense mutations in the serine/threonine kinase domain [c.1006T > A (p.Y336N), c.1063G > C (p.A355P), c.1069G > T (p.G357W), c.1582C > T (p.R528C), c.1583G > A (p.R528H)], as seen in MFS2, and a single mutation in a splice-acceptor site (IVS1-2A > G) (Table 2, Fig. 1). The remaining four patients were shown to have TGFBR1 mutations: three missense mutations in the serine/threonine kinase domain [c.953T > G (p.M318R), c.1199A > G (p.D400G) and c.1460G > C (p.R487P)] and a missense mutation in the glycine/serine-rich (GS) domain [c.599C > T (p.T200I)] (Table 2, Fig. 1). Owing to the clinical overlap of MFS and Shprintzen–Goldberg craniosynostosis syndrome (SGS) with LDS, Loeys et al. (2005) also screened seven MFS patients (unlinked to FBN1) and five SGS patients for mutations in TGFBR1 and TGFBR2, but no abnormalities were seen at these loci.
Two other TGFBR2 mutations [c.773T > G (p.V258G) and c.1067G > C (p.R356P)] and another TGFBR1 mutation [c.722C > T (p.S241L)] have since been reported in other patients meeting the clinical description of LDS by other groups (Table 2, Fig. 1) (Ki et al. 2005; Matyas et al. 2006). Finally, Loeys et al. (2006) collected 30 more probands of LDS and found TGFBR2 mutations in 21 and TGFBR1 mutations in nine (Table 2, Fig. 1) (Loeys et al. 2006). Furthermore, LDS type II (LDS2) without craniofacial features was also proposed. Eight TGFBR2 and four TGFBR1 mutations were found in LDS2 patients (Tables 2 and 3, Fig. 1) (Loeys et al. 2006).
Familial thoracic aortic aneurysms and dissections
Non-syndromic thoracic aortic aneurysms and dissections (TAAD) have complex and heterogeneous etiology, with some families inheriting TAAD in an autosomal dominant fashion, with decreased penetrance and variable expression. To date, at least three TAAD loci have been mapped by linkage studies of large single families (Pannu et al. 2005b): TAAD1 at 5q13–q14 (Guo et al. 2001), the familial aortic aneurysm 1 locus (FAA1) at 11q23–q24 (Vaughan et al. 2001), and TAAD2 at 3p24–p25 (Hasham et al. 2003).
The realization that TAAD2 (OMIM, #608987) and MFS2 are clinically overlapping diseases that both map to 3p24–p25 led Pannu et al. (2005a) to look for TGFBR2 mutations in 80 unrelated TAAD families, including the large family with disease linkage to 3p24–p25. Two TGFBR2 mutations [c.1378C > T (p.R460C) and c.1379G > A (p.R460H)], both affecting the same amino acid residue in the serine/threonine kinase domain, were identified in four families, or approximately 5% of the TAAD cases (Pannu et al. 2005a) (Table 2, Fig. 1). Each mutation occurred in the unique haplotype block, indicating an independent mutation event.
The mutational hotspot at the p.R460 residue in TAAD2 suggested a positive phenotype–genotype correlation, and this observation was supported by the discovery of another family with p.R460H that was initially diagnosed as having TAAD2 and, later, as having a distinctive condition with cardiovascular findings consistent with TAAD2, together with arterious tortuosity and aneurysm (Law et al. 2005, 2006). Three additional missense TGFBR2 mutations [c.1159G > A (p.V387G), c.1181G > A (p.C394Y), c.1657T > A (p.S553T)] and, more importantly, one TGFBR1 mutation [c.1460G > A (p.R487Q)] were found in four TAAD patients (Matyas et al. 2006).
Shprintzen–Goldberg craniosynostosis syndrome
Shprintzen–Goldberg craniosynostosis syndrome (SGS, OMIM, #182212) is characterized by craniosynostosis and other craniofacial features, marfanoid skeletal abnormalities, and developmental delay (Robinson et al. 2005). Furlong syndrome (FS) is a similar marfanoid disorder with craniosynostosis, which differs from SGS by the absence of mental retardation (Furlong et al. 1987).
Although a question was raised on the involvement of FBN1 abnormality in SGS (Robinson et al. 2005), at least two FBN1 mutations were identified (Kosaki et al. 2006; Sood et al. 1996).
Furthermore, the initial study of five SGS patients failed to reveal mutations in either TGFBR1 or TGFBR2 (Loeys et al. 2005), but Kosaki et al. (2006) recently reported a TGFBR2 mutation [IVS5-2A > G] (Table 2, Fig. 1) in an SGS patient with craniofacial and skeletal abnormalities, mild developmental delay, and cardiovascular features, including aortic regurgitation, annuloaortic ectasia and sigmoid configuration of the brachiocephalic, left common carotid and left subclavian arteries. Robinson et al. (2006), however, suggested that this patient might be more appropriately diagnosed as having LDS, due to the presence of bifid uvula and arterial manifestations.
An identical TGFBR1 mutation [c.722C > T (p.S241L)] was reported in two unrelated patients described as having probable FS (Table 2, Fig. 1) (Ades et al. 2006). One of these patients had learning difficulties, and the other had normal intelligence (Ades et al. 2006). Systemic arterial tortuosity was not evaluated in either of them, but one showed bifid uvula, consistent with LDS. The same mutation was also reported in an LDS patient with hypertelorism, tortuous arteries and bifid uvula (Matyas et al. 2006).
Germline TGFBR mutations and connective tissue disorders
The various mutations, as well as gene disruption by chromosomal structural abnormality, suggest that loss-of-function mutations of TGFBR2 are responsible for a wide spectrum of connective tissue disorders, but no simple genotype–phenotype correlation has been observed. It is intriguing that domain-specific germline mutations of TGFB1 cause Camurati–Engelmann disease (CED, OMIM #131300) (Kinoshita et al. 2000) associated with marfanoid habitus, despite the absence of connective tissue fragility. TGFB1 mutations in CED were shown to cause increased TGFβ signaling (Janssens et al. 2003). These findings suggest that abnormal TGFβ signaling could be responsible for the skeletal features of MFS. This hypothesis is corroborated by earlier studies describing the roles of TGFβ signaling in skeletal development (Alvarez and Serra 2004; Serra et al. 1999).
TGFBR2 and TGFBR1 mutations found in MFS and its related disorders are generally associated with severe vascular consequences that differ somewhat from those found in patients with FBN1 mutations. The vascular consequences associated with TGFBR2 and TGFBR1 mutations include aortic dissections in young age and aneurysms at sites distant from the aortic root. Thus, careful screening of arterial tortuosity and aneurysms from head to leg is highly recommended in this patient population.
Whether an MFS2 phenotype exists as a separate disorder from LDS is debatable. TGFBR1 and TGFBR2 aberrations are highly prevalent in LDS, which is characterized by severe cardiovascular features and family histories of aortic dissection with sudden death. Following the first report of TGFBR2 mutations in MFS (Mizuguchi et al. 2004), LDS, TAAD2, SGS, and FS were recognized as TGFBR mutation-related disorders. Although the original MFS2 patients with TGFBR2 mutations (Mizuguchi et al. 2004) could not be reasonably re-examined for the presence of LDS features such as bifid uvula, hypertelorism, craniosynostosis, and arterial tortuosity, at least three reports have since described TGFBR1 or TGFBR2 mutations in classic MFS patients in whom LDS had been ruled out (Disabella et al. 2006; Matyas et al. 2006; Singh et al. 2006). Four missense mutations [c.1273A > G (p.M425V), c.1378C > T (p.R460C), c.1379G > A (p.R460H), and c.1561T > C (p.W521R)] and compound heterozygous mutations [c.1337G > A (p.D446N) and c.-334T > A] in TGFBR2 were identified in five patients with MFS that satisfied the Ghent criteria. Seven more patients with MFS that did not satisfy the Ghent criteria were also reported to have mutations in TGFBR2 (Disabella et al. 2006; Matyas et al. 2006; Sakai et al. 2006; Singh et al. 2006). In addition, two TGFBR1 missense mutations [c.759G > A (p.M253I) and c.934G > A (p.G312S)] were found in two MFS patients and two TGFBR1 mutations [c.799A > C (p.N267H) and c.1135A > G (p.M379V)] were identified in two patients not fulfilling MFS. Thus, TGFBR2 and TGFBR1 aberrations were observed in typical cases of classic MFS with no signs of LDS (Singh et al. 2006).
However, it should be noted that arterial tortuosity, a cardinal feature of LDS, was not systematically evaluated in any of the four studies (Disabella et al. 2006; Matyas et al. 2006; Sakai et al. 2006; Singh et al. 2006). Moreover, two research groups were unable to identify TGFBR2 mutations in 29 MFS patients (FBN1 was normal in 24 and unknown in five) (Ki et al. 2005) and seven patients (FBN1 was normal) with MFS compatible with the Ghent criteria (Loeys et al. 2005). Thus, the question of whether an MFS2 phenotype and LDS should be classified as the same disorder remains unresolved.
At least six TGFBR2 mutations (R356P, N384S, R460H, R460C, S449F, and R537C) and two TGFBR1 mutations (S241L and R487Q) were recognized in two or three conditions (i.e., R460H found in MFS, LDS, and TAAD) (Ades et al. 2006; Disabella et al. 2006; Ki et al. 2005; Loeys et al. 2006; Matyas et al. 2006; Mizuguchi et al. 2004; Pannu et al. 2005a; Sakai et al. 2006; Singh et al. 2006) (Fig. 1, Table 2), implying that TGFBR mutations may cause various clinical consequences or that appropriate diagnosis is rather difficult for these disorders.
Pathogenesis of Marfan syndrome
Fibrillin-1 involvement in connective tissue disorders
Extracellular matrix (ECM) is formed by a number of macromolecules that are secreted and deposited into the space surrounding cells, where they are essential for tissue development and homeostasis. Fibrillin-1 is an ECM protein that is assembled into microfibrillar networks, where it interacts with other ECM proteins. Fibrillin-rich microfibrils form peripheral components of elastic fibers, which play a role as an architectural foundation and provide elasticity to tissues (Kielty et al. 2002).
Fibrillin-1 is a multi-domain protein that contains three characteristic modules: an epidermal growth factor (EGF)-like motif, a latent TGFβ binding protein (LTBP) motif, and a fusion of the two (Fib motif). The majority of the EGF-like modules in the FBN1 gene have a conserved calcium-binding sequence (cbEGF-like module).
Mutations associated with MFS are distributed over the entire FBN1 gene. No genotype–phenotype correlation has been clearly established, except for neonatal MFS (nMFS). Most mutations causing nMFS seem to be clustered in exons 24–32, although other phenotypes are also associated with mutations in these exons.
The fibrillin-1 protein contains 47 EGF-like modules, characterized by six cysteine residues that form disulfide bonds with one another. Most MFS missense mutations occur in one of the 43 cbEGF-like modules (Boileau et al. 2005; Robinson and Godfrey 2000). These mutations are thought to influence the protein structure and calcium binding affinity of cbEGF-like modules, with various deleterious effects (Boileau et al. 2005; Downing et al. 1996).
In classical MFS the incidence of ectopia lentis was significantly higher in patients harboring cysteine substitutions in the cbEGF-like modules than in patients with premature termination codon (PTC) mutations (Arbustini et al. 2005; Biggin et al. 2004; Loeys et al. 2004; Rommel et al. 2005; Schrijver et al. 2002). Furthermore, isolated or predominant ectopia lentis is frequently associated with cysteine substitutions (Ades et al. 2004; Comeglio et al. 2002), suggesting that cysteine residues may have a critical function in suspensory ligaments of the eyes, as previously described (Rommel et al. 2005).
The dominant–negative mechanism of mutant fibrillin-1 in MFS pathology is an attractive hypothesis in the light of the polymerizing nature of fibrillin-1 molecules into microfibrils. Various studies have examined the correlation between the expression level of mutant mRNA produced by PTC mutations and the degree of clinical severity (Ades et al. 2004; Dietz et al. 1993; Schrijver et al. 2002). Nonsense-mediated mRNA decay could limit aberrant protein production from mutated alleles in heterozygous patients, but remnant mutant fibrillin-1 proteins may act in a dominant–negative fashion. However, the hypothesized association between expression level and clinical severity is controversial. It is also possible that different expression levels of normal FBN1 alleles leads to different phenotypes in MFS family members sharing the same heterozygous PTC mutation (Hutchinson et al. 2003).
Support for the haploinsufficiency hypothesis as a mechanism in MFS pathogenesis comes from the finding of an FBN1 deletion in a patient with a number of marfanoid characteristics (Hutchinson et al. 2003; Judge et al. 2004) as well as in mouse models (Judge et al. 2004). In transgenic mice carrying human mutant FBN1 (p.C1663R), no obvious cardiovascular and skeletal signs have been recognized, despite the co-assembly of the mutant protein into mouse microfibrillar networks. Other knock-in mice (in which the p.C1039G mutation Fbn1 C1039G/+ targeted the endogenous Fbn1) showed decreased microfibrillar deposition, skeletal abnormalities and changes in the architecture of the aortic wall. Notably, the aortic wall phenotype was rescued by the wildtype FBN1 transgene.
TGFβ signaling and connective tissue disorders
TGFβ is a secreted polypeptide that plays diverse roles in cell proliferation and differentiation, apoptosis, and extracellular matrix formation (Cohen 2003; Derynck et al. 2001; Ignotz and Massague 1986; Massague et al. 2000). TGFβ1 is abundant in the ECM. An inactive form of mature TGFβ1 stays in a complex with latency-associated polypeptide (LAP), which is an N-terminal peptide cleaved from proTGFβ1, and latent TGFβ binding protein (LTBP). Mature TGFβ1 and LAP are noncovalently associated in a small latency complex (SLC). The SLC binds to LTBP via disulfide bonds between LAP and LTBP, forming a large latency complex (LLC) (Annes et al. 2003). LTBP plays a role in folding and secreting TGFβ1, targeting it appropriately to the ECM, and modulating TGFβ activity (Charbonneau et al. 2004; Kaartinen and Warburton 2003; Ramirez et al. 2004; Rifkin 2005).
A recent study revealed that LTBP-1 (one of the LTBPs) and fibrillin interact in vitro and suggested that fibrillin-1 may stabilize the latent TGFβ complexes in the ECM (Isogai et al. 2003). Support for this hypothesis is seen in mouse models of MFS. Three strains of transgenic mice, each harboring a different type of Fbn1 mutation, displayed several MFS features with variable severity (Judge et al. 2004; Pereira et al. 1997, 1999). Increased TGFβ activity was observed in at least four organs (lung, mitral valve, aortic and dural tissues), possibly as a result of excess free LLC due to inadequate stabilization within the ECM, as previously hypothesized (Habashi et al. 2006; Jones et al. 2005; Neptune et al. 2003; Ng et al. 2004; Rifkin 2005). Administration of an anti-TGFβ neutralizing antibody rescued the lung, mitral valve, and aortic tissue phenotypes (Habashi et al. 2006; Neptune et al. 2003; Ng et al. 2004). Furthermore, aortic aneurysm was prevented by the administration of losartan, an angiotensin II type 1 receptor blocker that alleviates increased TGFβ activity (Habashi et al. 2006). Taken together, these findings support an association of abnormal TGFβ signaling with MFS pathogenesis. Interaction between the affected structural ECM components and aberrant TGFβ signaling may coordinately determine MFS phenotypes.
TGFβ transduces its signals via two distinct types of transmembrane receptors, type I (TβRI) and type II (TβRII), encoded by TGFBR1 and TGFBR2, respectively (ten Dijke et al. 1996; Wrana et al. 1994). Both types of receptors consist of an extracellular domain, a transmembrane domain and a serine/threonine kinase domain. A glycine/serine-rich domain (GS domain) is specific for TβRI. The ligand-bound TβRII phosphorylates the GS domain, which then acts in signal transduction (Wieser et al. 1995).
It is likely that abnormal TGFβ signaling is involved in the human MFS phenotype. Identification of TGFBR2 mutations in MFS2 supports the hypothesis and provided the first direct link between a human connective tissue disorder and abnormal TGFβ signaling. Loss-of-function TGFBR2 mutations are hypothesized to cause MFS2, LDS and TAAD2. In our previous study mammalian cells were transfected with MFS2-related mutant TGFBR2 constructs, and a luciferase assay clearly showed decreased TGFβ signaling activity (Mizuguchi et al. 2004). If the highly conserved p.R460 residue is essential for the structural integrity of the catalytic loop of TβRII, the p.R460 missense mutations found in TAAD2 could dramatically perturb TGFβ signaling (Pannu et al. 2005a).
By contrast, aortic tissues from LDS individuals showed increased TGFβ signaling activity. The heterozygous state (with one normal allele and the other mutant) of TGFBR2 abnormality in affected individuals might not simply reflect a loss-of-function nature of the mutation, probably because of the complex regulation of the TGFβ signaling pathway (Loeys et al. 2005). Paradoxically, increased TGFβ signaling was also shown in the kinase-deficient TβRII transgenic mice (Denton et al. 2003).
Conclusion
Recent genetic studies of MFS, both in mice and humans, revealed that TGFβ signaling is involved in the pathogenesis of MFS and related disorders. FBN1 abnormalities appear to be the major genetic cause of MFS, but not the only cause.
MFS-related disorders share many features with MFS. Clinical data should be carefully evaluated, with the recognition that incomplete data might lead to different diagnoses. Further studies are needed in order to establish frames of nosology for MFS and MFS-related disorders, by collection and analysis of extensive genetic and clinical data from many affected patients. An appropriate diagnostic system(s) is needed to differentiate MFS and MFS-related disorders.
References
Ades LC, Holman KJ, Brett MS, Edwards MJ, Bennetts B (2004) Ectopia lentis phenotypes and the FBN1 gene. Am J Med Genet A 126:284–289
Ades LC, Sullivan K, Biggin A, Haan EA, Brett M, Holman KJ, Dixon J, Robertson S, Holmes AD, Rogers J, Bennetts B (2006) FBN1, TGFBR1, and the Marfan-craniosynostosis/mental retardation disorders revisited. Am J Med Genet A 140:1047–1058
Alvarez J, Serra R (2004) Unique and redundant roles of Smad3 in TGF-beta-mediated regulation of long bone development in organ culture. Dev Dyn 230:685–699
Annes JP, Munger JS, Rifkin DB (2003) Making sense of latent TGFbeta activation. J Cell Sci 116:217–224
Arbustini E, Grasso M, Ansaldi S, Malattia C, Pilotto A, Porcu E, Disabella E, Marziliano N, Pisani A, Lanzarini L, Mannarino S, Larizza D, Mosconi M, Antoniazzi E, Zoia MC, Meloni G, Magrassi L, Brega A, Bedeschi MF, Torrente I, Mari F, Tavazzi L (2005) Identification of sixty-two novel and twelve known FBN1 mutations in eighty-one unrelated probands with Marfan syndrome and other fibrillinopathies. Hum Mutat 26:494
Biggin A, Holman K, Brett M, Bennetts B, Ades L (2004) Detection of thirty novel FBN1 mutations in patients with Marfan syndrome or a related fibrillinopathy. Hum Mutat 23:99
Boileau C, Jondeau G, Babron MC, Coulon M, Alexandre JA, Sakai L, Melki J, Delorme G, Dubourg O, Bonaiti-Pellie C, et al (1993) Autosomal dominant Marfan-like connective-tissue disorder with aortic dilation and skeletal anomalies not linked to the fibrillin genes. Am J Hum Genet 53:46–54
Boileau C, Jondeau G, Mizuguchi T, Matsumoto N (2005) Molecular genetics of Marfan syndrome. Curr Opin Cardiol 20:194–200
Charbonneau NL, Ono RN, Corson GM, Keene DR, Sakai LY (2004) Fine tuning of growth factor signals depends on fibrillin microfibril networks. Birth Defects Res C Embryo Today 72:37–50
Cohen MM Jr (2003) TGF beta/Smad signaling system and its pathologic correlates. Am J Med Genet A 116:1–10
Collod G, Babron MC, Jondeau G, Coulon M, Weissenbach J, Dubourg O, Bourdarias JP, Bonaiti-Pellie C, Junien C, Boileau C (1994) A second locus for Marfan syndrome maps to chromosome 3p24.2-p25. Nat Genet 8:264–268
Collod-Beroud G, Le Bourdelles S, Ades L, Ala-Kokko L, Booms P, Boxer M, Child A, Comeglio P, De Paepe A, Hyland JC, Holman K, Kaitila I, Loeys B, Matyas G, Nuytinck L, Peltonen L, Rantamaki T, Robinson P, Steinmann B, Junien C, Beroud C, Boileau C (2003) Update of the UMD-FBN1 mutation database and creation of an FBN1 polymorphism database. Hum Mutat 22:199–208
Comeglio P, Evans AL, Brice G, Cooling RJ, Child AH (2002) Identification of FBN1 gene mutations in patients with ectopia lentis and marfanoid habitus. Br J Ophthalmol 86:1359–1362
Corson GM, Chalberg SC, Dietz HC, Charbonneau NL, Sakai LY (1993) Fibrillin binds calcium and is coded by cDNAs that reveal a multidomain structure and alternatively spliced exons at the 5′ end. Genomics 17:476–484
De Paepe A, Devereux RB, Dietz HC, Hennekam RC, Pyeritz RE (1996) Revised diagnostic criteria for the Marfan syndrome. Am J Med Genet 62:417–426
Denton CP, Zheng B, Evans LA, Shi-wen X, Ong VH, Fisher I, Lazaridis K, Abraham DJ, Black CM, de Crombrugghe B (2003) Fibroblast-specific expression of a kinase-deficient type II transforming growth factor beta (TGFbeta) receptor leads to paradoxical activation of TGFbeta signaling pathways with fibrosis in transgenic mice. J Biol Chem 278:25109–25119
Derynck R, Akhurst RJ, Balmain A (2001) TGF-beta signaling in tumor suppression and cancer progression. Nat Genet 29:117–129
Dietz HC, Cutting GR, Pyeritz RE, Maslen CL, Sakai LY, Corson GM, Puffenberger EG, Hamosh A, Nanthakumar EJ, Curristin SM, et al (1991) Marfan syndrome caused by a recurrent de novo missense mutation in the fibrillin gene. Nature 352:337–339
Dietz HC, McIntosh I, Sakai LY, Corson GM, Chalberg SC, Pyeritz RE, Francomano CA (1993) Four novel FBN1 mutations: significance for mutant transcript level and EGF-like domain calcium binding in the pathogenesis of Marfan syndrome. Genomics 17:468–475
Dietz H, Francke U, Furthmayr H, Francomano C, De Paepe A, Devereux R, Ramirez F, Pyeritz R (1995) The question of heterogeneity in Marfan syndrome. Nat Genet 9:228–229
Disabella E, Grasso M, Marziliano N, Ansaldi S, Lucchelli C, Porcu E, Tagliani M, Pilotto A, Diegoli M, Lanzarini L, Malattia C, Pelliccia A, Ficcadenti A, Gabrielli O, Arbustini E (2006) Two novel and one known mutation of the TGFBR2 gene in Marfan syndrome not associated with FBN1 gene defects. Eur J Hum Genet 14:34–38
Downing AK, Knott V, Werner JM, Cardy CM, Campbell ID, Handford PA (1996) Solution structure of a pair of calcium-binding epidermal growth factor-like domains: implications for the Marfan syndrome and other genetic disorders. Cell 85:597–605
Faivre L, Gorlin RJ, Wirtz MK, Godfrey M, Dagoneau N, Samples JR, Le Merrer M, Collod-Beroud G, Boileau C, Munnich A, Cormier-Daire V (2003) In frame fibrillin-1 gene deletion in autosomal dominant Weill–Marchesani syndrome. J Med Genet 40:34–36
Francke U, Berg MA, Tynan K, Brenn T, Liu W, Aoyama T, Gasner C, Miller DC, Furthmayr H (1995) A Gly1127Ser mutation in an EGF-like domain of the fibrillin-1 gene is a risk factor for ascending aortic aneurysm and dissection. Am J Hum Genet 56:1287–1296
Furlong J, Kurczynski TW, Hennessy JR (1987) New Marfanoid syndrome with craniosynostosis. Am J Med Genet 26:599–604
Gilchrist DM (1994) Marfan syndrome or Marfan-like connective-tissue disorder. Am J Hum Genet 54:553–554
Guo D, Hasham S, Kuang SQ, Vaughan CJ, Boerwinkle E, Chen H, Abuelo D, Dietz HC, Basson CT, Shete SS, Milewicz DM (2001) Familial thoracic aortic aneurysms and dissections: genetic heterogeneity with a major locus mapping to 5q13–14. Circulation 103:2461–2468
Habashi JP, Judge DP, Holm TM, Cohn RD, Loeys BL, Cooper TK, Myers L, Klein EC, Liu G, Calvi C, Podowski M, Neptune ER, Halushka MK, Bedja D, Gabrielson K, Rifkin DB, Carta L, Ramirez F, Huso DL, Dietz HC (2006) Losartan, an AT1 antagonist, prevents aortic aneurysm in a mouse model of Marfan syndrome. Science 312:117–121
Halliday DJ, Hutchinson S, Lonie L, Hurst JA, Firth H, Handford PA, Wordsworth P (2002) Twelve novel FBN1 mutations in Marfan syndrome and Marfan related phenotypes test the feasibility of FBN1 mutation testing in clinical practice. J Med Genet 39:589–593
Hasham SN, Willing MC, Guo DC, Muilenburg A, He R, Tran VT, Scherer SE, Shete SS, Milewicz DM (2003) Mapping a locus for familial thoracic aortic aneurysms and dissections (TAAD2) to 3p24–25. Circulation 107:3184–3190
Hutchinson S, Furger A, Halliday D, Judge DP, Jefferson A, Dietz HC, Firth H, Handford PA (2003) Allelic variation in normal human FBN1 expression in a family with Marfan syndrome: a potential modifier of phenotype? Hum Mol Genet 12:2269–2276
Ignotz RA, Massague J (1986) Transforming growth factor-beta stimulates the expression of fibronectin and collagen and their incorporation into the extracellular matrix. J Biol Chem 261:4337–4345
Isogai Z, Ono RN, Ushiro S, Keene DR, Chen Y, Mazzieri R, Charbonneau NL, Reinhardt DP, Rifkin DB, Sakai LY (2003) Latent transforming growth factor beta-binding protein 1 interacts with fibrillin and is a microfibril-associated protein. J Biol Chem 278:2750–2757
Janssens K, ten Dijke P, Ralston SH, Bergmann C, Van Hul W (2003) Transforming growth factor-beta 1 mutations in Camurati–Engelmann disease lead to increased signaling by altering either activation or secretion of the mutant protein. J Biol Chem 278:7718–7724
Jones KB, Myers L, Judge DP, Kirby PA, Dietz HC, Sponseller PD (2005) Toward an understanding of dural ectasia: a light microscopy study in a murine model of Marfan syndrome. Spine 30:291–293
Judge DP, Biery NJ, Keene DR, Geubtner J, Myers L, Huso DL, Sakai LY, Dietz HC (2004) Evidence for a critical contribution of haploinsufficiency in the complex pathogenesis of Marfan syndrome. J Clin Invest 114:172–181
Kaartinen V, Warburton D (2003) Fibrillin controls TGF-beta activation. Nat Genet 33:331–332
Kainulainen K, Karttunen L, Puhakka L, Sakai L, Peltonen L (1994) Mutations in the fibrillin gene responsible for dominant ectopia lentis and neonatal Marfan syndrome. Nat Genet 6:64–69
Katzke S, Booms P, Tiecke F, Palz M, Pletschacher A, Turkmen S, Neumann LM, Pregla R, Leitner C, Schramm C, Lorenz P, Hagemeier C, Fuchs J, Skovby F, Rosenberg T, Robinson PN (2002) TGGE screening of the entire FBN1 coding sequence in 126 individuals with Marfan syndrome and related fibrillinopathies. Hum Mutat 20:197–208
Ki CS, Jin DK, Chang SH, Kim JE, Kim JW, Park BK, Choi JH, Park IS, Yoo HW (2005) Identification of a novel TGFBR2 gene mutation in a Korean patient with Loeys–Dietz aortic aneurysm syndrome; no mutation in TGFBR2 gene in 30 patients with classic Marfan’s syndrome. Clin Genet 68:561–563
Kielty CM, Sherratt MJ, Shuttleworth CA (2002) Elastic fibres. J Cell Sci 115:2817–2828
Kinoshita A, Saito T, Tomita H, Makita Y, Yoshida K, Ghadami M, Yamada K, Kondo S, Ikegawa S, Nishimura G, Fukushima Y, Nakagomi T, Saito H, Sugimoto T, Kamegaya M, Hisa K, Murray JC, Taniguchi N, Niikawa N, Yoshiura K (2000) Domain-specific mutations in TGFB1 result in Camurati–Engelmann disease. Nat Genet 26:19–20
Kosaki K, Takahashi D, Udaka T, Kosaki R, Matsumoto M, Ibe S, Isobe T, Tanaka Y, Takahashi T (2006) Molecular pathology of Shprintzen–Goldberg syndrome. Am J Med Genet A 140:104–108; author reply 109–110
Law C, Bunyan D, Castle B, Day L, Keeton B, Simpson I, Westwood G (2005) Clinical findings in a large family with a predisposition to aortic dilatation and dissection and an R460H mutation in TGFBR2. J Med Genet 42 [Suppl 1]:S31
Law CJ, Bunyan D, Castle B, Day L, Simpson I, Westwood G, Keeton B (2006) Clinical features in a family with a R460H mutation in TGFBR2. J Med Genet (in press)
Lee B, Godfrey M, Vitale E, Hori H, Mattei MG, Sarfarazi M, Tsipouras P, Ramirez F, Hollister DW (1991) Linkage of Marfan syndrome and a phenotypically related disorder to two different fibrillin genes. Nature 352:330–334
Loeys B, Nuytinck L, Delvaux I, De Bie S, De Paepe A (2001) Genotype and phenotype analysis of 171 patients referred for molecular study of the fibrillin-1 gene FBN1 because of suspected Marfan syndrome. Arch Intern Med 161:2447–2454
Loeys B, De Backer J, Van Acker P, Wettinck K, Pals G, Nuytinck L, Coucke P, De Paepe A (2004) Comprehensive molecular screening of the FBN1 gene favors locus homogeneity of classical Marfan syndrome. Hum Mutat 24:140–146
Loeys BL, Chen J, Neptune ER, Judge DP, Podowski M, Holm T, Meyers J, Leitch CC, Katsanis N, Sharifi N, Xu FL, Myers LA, Spevak PJ, Cameron DE, De Backer J, Hellemans J, Chen Y, Davis EC, Webb CL, Kress W, Coucke P, Rifkin DB, De Paepe AM, Dietz HC (2005) A syndrome of altered cardiovascular, craniofacial, neurocognitive and skeletal development caused by mutations in TGFBR1 or TGFBR2. Nat Genet 37:275–281
Loeys BL, Schwarze U, Holm T, Callewaert BL, Thomas GH, Pannu H, De Backer JF, Oswald GL, Symoens S, Manouvrier S, Roberts AE, Faravelli F, Greco MA, Pyeritz RE, Milewicz DM, Coucke PJ, Cameron DE, Braverman AC, Byers PH, De Paepe AM, Dietz HC (2006) Aneurysm syndromes caused by mutations in the TGF-beta receptor. N Engl J Med 355:788–798
Massague J, Blain SW, Lo RS (2000) TGFbeta signaling in growth control, cancer, and heritable disorders. Cell 103:295–309
Matyas G, Arnold E, Carrel T, Baumgartner D, Boileau C, Berer W, Steinmann B (2006) Identification and in silico analyses of novel TGFBR1 and TGFBR2 mutations in Marfan syndrome-related disorders. Hum Mutat 27:760–769
Milewicz DM, Michael K, Fisher N, Coselli JS, Markello T, Biddinger A (1996) Fibrillin-1 (FBN1) mutations in patients with thoracic aortic aneurysms. Circulation 94:2708–2711
Mizuguchi T, Collod-Beroud G, Akiyama T, Abifadel M, Harada N, Morisaki T, Allard D, Varret M, Claustres M, Morisaki H, Ihara M, Kinoshita A, Yoshiura K, Junien C, Kajii T, Jondeau G, Ohta T, Kishino T, Furukawa Y, Nakamura Y, Niikawa N, Boileau C, Matsumoto N (2004) Heterozygous TGFBR2 mutations in Marfan syndrome. Nat Genet 36:855–860
Neptune ER, Frischmeyer PA, Arking DE, Myers L, Bunton TE, Gayraud B, Ramirez F, Sakai LY, Dietz HC (2003) Dysregulation of TGF-beta activation contributes to pathogenesis in Marfan syndrome. Nat Genet 33:407–411
Ng CM, Cheng A, Myers LA, Martinez-Murillo F, Jie C, Bedja D, Gabrielson KL, Hausladen JM, Mecham RP, Judge DP, Dietz HC (2004) TGF-beta-dependent pathogenesis of mitral valve prolapse in a mouse model of Marfan syndrome. J Clin Invest 114:1586–1592
Pannu H, Fadulu VT, Chang J, Lafont A, Hasham SN, Sparks E, Giampietro PF, Zaleski C, Estrera AL, Safi HJ, Shete S, Willing MC, Raman CS, Milewicz DM (2005a) Mutations in transforming growth factor-beta receptor type II cause familial thoracic aortic aneurysms and dissections. Circulation 112:513–520
Pannu H, Tran-Fadulu V, Milewicz DM (2005b) Genetic basis of thoracic aortic aneurysms and aortic dissections. Am J Med Genet C Semin Med Genet 139:10–16
Pereira L, Andrikopoulos K, Tian J, Lee SY, Keene DR, Ono R, Reinhardt DP, Sakai LY, Biery NJ, Bunton T, Dietz HC, Ramirez F (1997) Targeting of the gene encoding fibrillin-1 recapitulates the vascular aspect of Marfan syndrome. Nat Genet 17:218–222
Pereira L, Lee SY, Gayraud B, Andrikopoulos K, Shapiro SD, Bunton T, Biery NJ, Dietz HC, Sakai LY, Ramirez F (1999) Pathogenetic sequence for aneurysm revealed in mice underexpressing fibrillin-1. Proc Natl Acad Sci U S A 96:3819–3823
Ramirez F, Sakai LY, Dietz HC, Rifkin DB (2004) Fibrillin microfibrils: multipurpose extracellular networks in organismal physiology. Physiol Genomics 19:151–154
Rifkin DB (2005) Latent transforming growth factor-beta (TGF-beta) binding proteins: orchestrators of TGF-beta availability. J Biol Chem 280:7409–7412
Robinson PN, Godfrey M (2000) The molecular genetics of Marfan syndrome and related microfibrillopathies. J Med Genet 37:9–25
Robinson PN, Neumann LM, Demuth S, Enders H, Jung U, Konig R, Mitulla B, Muller D, Muschke P, Pfeiffer L, Prager B, Somer M, Tinschert S (2005) Shprintzen–Goldberg syndrome: fourteen new patients and a clinical analysis. Am J Med Genet A 135:251–262
Robinson P, Neumann L, Tinschert S (2006) Response to Kosaki et al. “Molecular pathology of Shprintzen–Goldberg syndrome”. Am J Med Genet 140A:109–110
Rommel K, Karck M, Haverich A, Schmidtke J, Arslan-Kirchner M (2002) Mutation screening of the fibrillin-1 (FBN1) gene in 76 unrelated patients with Marfan syndrome or Marfanoid features leads to the identification of 11 novel and three previously reported mutations. Hum Mutat 20:406–407
Rommel K, Karck M, Haverich A, von Kodolitsch Y, Rybczynski M, Muller G, Singh KK, Schmidtke J, Arslan-Kirchner M (2005) Identification of 29 novel and nine recurrent fibrillin-1 (FBN1) mutations and genotype–phenotype correlations in 76 patients with Marfan syndrome. Hum Mutat 26:529–539
Sakai H, Visser R, Ikegawa S, Ito E, Numabe H, Watanabe Y, Mikami H, Kondoh T, Kitoh H, Sugiyama R, Okamoto N, Ogata T, Fodde R, Mizuno S, Takamura K, Egashira M, Sasaki N, Watanabe S, Nishimaki S, Takada F, Nagai T, Okada Y, Aoka Y, Yasuda K, Iwasa M, Kogaki S, Harada N, Mizuguchi T, Matsumoto N (2006) Comprehensive genetic analysis of relevant four genes in 49 patients with Marfan syndrome or Marfan-related phenotypes. Am J Med Genet A 140:1719–1725
Schrijver I, Liu W, Odom R, Brenn T, Oefner P, Furthmayr H, Francke U (2002) Premature termination mutations in FBN1: distinct effects on differential allelic expression and on protein and clinical phenotypes. Am J Hum Genet 71:223–237
Serra R, Karaplis A, Sohn P (1999) Parathyroid hormone-related peptide (PTHrP)-dependent and -independent effects of transforming growth factor beta (TGF-beta) on endochondral bone formation. J Cell Biol 145:783–794
Singh KK, Rommel K, Mishra A, Karck M, Haverich A, Schmidtke J, Arslan-Kirchner M (2006) TGFBR1 and TGFBR2 mutations in patients with features of Marfan syndrome and Loeys–Dietz syndrome. Hum Mutat 27: 770–777
Sood S, Eldadah ZA, Krause WL, McIntosh I, Dietz HC (1996) Mutation in fibrillin-1 and the Marfanoid-craniosynostosis (Shprintzen–Goldberg) syndrome. Nat Genet 12:209–211
ten Dijke P, Miyazono K, Heldin CH (1996) Signaling via hetero-oligomeric complexes of type I and type II serine/threonine kinase receptors. Curr Opin Cell Biol 8:139–145
Tynan K, Comeau K, Pearson M, Wilgenbus P, Levitt D, Gasner C, Berg MA, Miller DC, Francke U (1993) Mutation screening of complete fibrillin-1 coding sequence: report of five new mutations, including two in 8-cysteine domains. Hum Mol Genet 2:1813–1821
Vaughan CJ, Casey M, He J, Veugelers M, Henderson K, Guo D, Campagna R, Roman MJ, Milewicz DM, Devereux RB, Basson CT (2001) Identification of a chromosome 11q23.2–q24 locus for familial aortic aneurysm disease, a genetically heterogeneous disorder. Circulation 103:2469–2475
Wieser R, Wrana JL, Massague J (1995) GS domain mutations that constitutively activate T beta R-I, the downstream signaling component in the TGF-beta receptor complex. EMBO J 14:2199–2208
Wrana JL, Attisano L, Wieser R, Ventura F, Massague J (1994) Mechanism of activation of the TGF-beta receptor. Nature 370:341–347
Acknowledgment
We thank Dr. Remco Visser at the Department of Pediatrics, Leiden University Medical Center, Leiden, The Netherlands, for his critical reading of this manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Mizuguchi, T., Matsumoto, N. Recent progress in genetics of Marfan syndrome and Marfan-associated disorders. J Hum Genet 52, 1–12 (2007). https://doi.org/10.1007/s10038-006-0078-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10038-006-0078-1
Keywords
This article is cited by
-
Identification of novel FBN1 variations implicated in congenital scoliosis
Journal of Human Genetics (2020)
-
Extracellular matrix, regional heterogeneity of the aorta, and aortic aneurysm
Experimental & Molecular Medicine (2019)
-
Novel FBN1 mutations are responsible for cardiovascular manifestations of Marfan syndrome
Molecular Biology Reports (2016)